

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
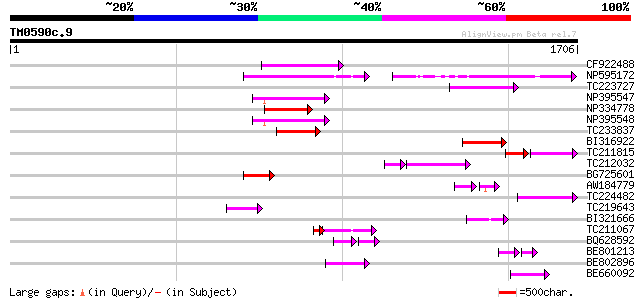
Query= TM0590c.9
(1706 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CF922488 203 6e-52
NP595172 polyprotein [Glycine max] 194 4e-49
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 183 5e-46
NP395547 reverse transcriptase [Glycine max] 156 6e-38
NP334778 reverse transcriptase [Glycine max] 153 5e-37
NP395548 reverse transcriptase [Glycine max] 144 3e-34
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 139 1e-32
BI316922 135 2e-31
TC211815 79 4e-28
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 92 3e-24
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 106 1e-22
AW184779 64 3e-19
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 86 1e-16
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 84 4e-16
BI321666 82 3e-15
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 58 3e-12
BQ628592 51 4e-12
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 52 8e-11
BE802896 65 2e-10
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 62 2e-09
>CF922488
Length = 741
Score = 203 bits (516), Expect = 6e-52
Identities = 109/245 (44%), Positives = 148/245 (59%)
Frame = +3
Query: 759 VKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHP 818
V K +GK MCVDY DLN A PKD +PLP I+ LVD + S MD +SGY+QI++ P
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 819 ADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVR 878
D +KT F+T +CY+ M FGLKN GATYQR M +F + + +EVY+DDMIVKS
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 879 GLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMK 938
+H +L + F +RK+ +RLNP KC F V+ K L F+ + RGIE++ K K I +M
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 939 SPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLS 998
P K+VQ GR+ + RF+ P F L KN +W +C AF R+K+ L
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 999 SPPIL 1003
+P +L
Sbjct: 723 NPHVL 737
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 194 bits (492), Expect = 4e-49
Identities = 118/379 (31%), Positives = 189/379 (49%)
Frame = +1
Query: 705 HRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 764
H + L PV R + +++ + ++L I+ P L +++VKK +G
Sbjct: 1759 HAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSPFSLP-ILLVKKKDG 1935
Query: 765 KWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKT 824
WR C DY LN KDS+P+P++D L+D G + S +D SGYHQI + P D +KT
Sbjct: 1936 SWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKT 2115
Query: 825 AFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQ 884
AF T +Y + MPFGL NA AT+Q LM+++F + + + V+ DD+++ S DH +
Sbjct: 2116 AFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLK 2295
Query: 885 DLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVK 944
LE +++H + KCSFG +LG ++ G+ + K +A+ +P+NVK
Sbjct: 2296 HLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVK 2475
Query: 945 EVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILS 1004
+++ G RF+ + + P L+K+ +F W E E AFV+LK+ ++ P+LS
Sbjct: 2476 QLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKD-SFLWNNEAEAAFVKLKKAMTEAPVLS 2652
Query: 1005 KPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLV 1064
P P L S + +V+ Q H I YF S L + + LA+
Sbjct: 2653 LPDFSQPFILETDASGIGVGAVLGQ---NGHPIAYF-SKKLAPRMQKQSAYTRELLAITE 2820
Query: 1065 TARRLRPYFQSFPVKVRTD 1083
+ R Y +RTD
Sbjct: 2821 ALSKFRHYLLGNKFIIRTD 2877
Score = 131 bits (330), Expect = 2e-30
Identities = 147/574 (25%), Positives = 243/574 (41%), Gaps = 20/574 (3%)
Frame = +1
Query: 1151 SNSSGSGAGVTLEGPGELVLEQSLKFEFKATNNQAEYEALIA---GLKLAREVKI-RSLL 1206
+++SG G G L G + S K + A L+A L R + +
Sbjct: 2686 TDASGIGVGAVLGQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFI 2865
Query: 1207 IRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLA 1266
IRTD + +++ + + Q P +L + L + +EY P +NQ ADAL+++
Sbjct: 2866 IRTDQRSLKSLMDQSLQT--PEQQAWLHKF-----LGYDFKIEYKPGKDNQAADALSRMF 3024
Query: 1267 STRKPGNNKSVIQETLAYPSIEGELMACVNRGRTWMDPIISILAGDPAEVEQCTKEQQRE 1326
+ ++E R R DP + L T +Q +
Sbjct: 3025 MLAWSEPHSIFLEEL---------------RARLISDPHLKQLME--------TYKQGAD 3135
Query: 1327 ASHYTLIDGHLYRRGFSTPLLKCVSPEKYEA---IMSEVHEGVCASHIG-GRSLACKVLR 1382
ASHYT+ +G LY + + V P + E I+ E H H G R+LA L+
Sbjct: 3136 ASHYTVREGLLYWKD------RVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLAR--LK 3291
Query: 1383 AGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW---GVDLVGPFP 1439
A FYWP +++D ++++C CQ + P L + P P +W +D + P
Sbjct: 3292 AQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL--PIPQQVWEDVAMDFITGLP 3465
Query: 1440 TARAQMKFILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQF 1498
+ + I+V +D TK+ PL ++K+V + IV GIPR+IVSD F
Sbjct: 3466 NSFG-LSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVF 3642
Query: 1499 SSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVL 1558
+S+ + K G + +S HPQ++GQ E N+ + LR E W+ LP
Sbjct: 3643 TSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAE 3822
Query: 1559 WSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETR 1618
+ YNT + TPFR YG + P + TR ++ A + +L
Sbjct: 3823 FWYNTAYHMSLGMTPFRALYGRE---PPTL------TRQACSIDDPAEVREQLTDRDALL 3975
Query: 1619 DEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL----KWRSGA----PGNKLTPNWEGP 1670
+ I T +Q + + + + Q+GD VL +R + KL+ + GP
Sbjct: 3976 AKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGP 4155
Query: 1671 YRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYF 1704
++++ +G+ AY LE R+ F+ L+ F
Sbjct: 4156 FKVLAKIGDVAYKLELPSAARIHPVFHVSQLKPF 4257
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 183 bits (465), Expect = 5e-46
Identities = 87/209 (41%), Positives = 125/209 (59%), Gaps = 1/209 (0%)
Frame = +1
Query: 1324 QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRA 1383
+R A+ + + LY+R L+CV + ++ EVHEG +H G ++A K+LRA
Sbjct: 217 RRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTHANGHAMARKILRA 396
Query: 1384 GFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPF-PTAR 1442
G+YW T+ DC V++C +CQ FAD APP L MS+PWPF+MWG+D++G P A
Sbjct: 397 GYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMWGIDVIGAIEPKAS 576
Query: 1443 AQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQ 1502
+FILVA+DYFTKW+EA + +V F K I+CR+G+PR I++DNGT ++
Sbjct: 577 NGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRKIITDNGTNLNNKM 756
Query: 1503 TREFCKEMGIQMRFASVEHPQTNGQVESA 1531
E C+E IQ + P+ N VE A
Sbjct: 757 MGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 156 bits (395), Expect = 6e-38
Identities = 90/248 (36%), Positives = 128/248 (51%), Gaps = 18/248 (7%)
Frame = +1
Query: 731 VQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG------------------KWRMCVDY 772
V++EV KLL A I + +W++ V +V K G +WRMC+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 773 TDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVN 832
LN+A KD YPLP +D ++ + +D YSGY+QI + P D++KTAF
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 833 YCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGE 892
+ YR MPFGL NA T+QR M +F V + +EV++DD + +LE+
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 893 IRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGR 952
K ++ LN EKC F VQ G LG I+ RGIE+ EK I ++ P NVK + G
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 953 IAALSRFL 960
+ RF+
Sbjct: 721 VGFYRRFI 744
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 153 bits (387), Expect = 5e-37
Identities = 72/143 (50%), Positives = 97/143 (67%)
Frame = +3
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
RMCVDY DLN+A PKD++PLP ID L+ + L S MD +SGY+QI+M P D +KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
+T +CY+ M FGLKN GATY R M +F + + +E YVD+MI KS +H +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 887 EEAFGEIRKHSMRLNPEKCSFGV 909
+ FG++RK+ +RLNP KC FG+
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 144 bits (363), Expect = 3e-34
Identities = 84/248 (33%), Positives = 132/248 (52%), Gaps = 18/248 (7%)
Frame = +1
Query: 731 VQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG------------------KWRMCVDY 772
V++EV KLL I + W++ V++V K G W++C+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 773 TDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVN 832
LN+A KD +PLP +D +++ +G+ +DAY GY+QI + P D++K AF
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 833 YCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGE 892
+ YR +PFGL NA T+Q M +FA V +++EV++DD V + LE
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 893 IRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGR 952
+ ++ LN EKC F V+ G LG I++RGIE++ K I+++ PSNVK ++ G+
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 953 IAALSRFL 960
RF+
Sbjct: 721 ARFYRRFI 744
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 139 bits (349), Expect = 1e-32
Identities = 67/132 (50%), Positives = 92/132 (68%)
Frame = +2
Query: 803 SLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVG 862
S MD +SGY+QI M D +KT F+T + YR M FGLKN GATYQR M +F +
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMMH 184
Query: 863 RNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSR 922
+ +EVYVDDMI KS +H +L + FG ++K+ ++LNP KC+FGV+ GK LGF+++ +
Sbjct: 185 KEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQK 364
Query: 923 GIEINPEKCKAI 934
GIEI+PEK KA+
Sbjct: 365 GIEIDPEKVKAL 400
>BI316922
Length = 405
Score = 135 bits (340), Expect = 2e-31
Identities = 61/134 (45%), Positives = 91/134 (67%)
Frame = +3
Query: 1361 EVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVT 1420
E+H G+C H + + +VLR G+Y T+RK C ++VK+C+EC F ++S +EL
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 1421 MSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRI 1480
+ APWPFA+ GVD++ PFP ++ Q+K++LV +D FTKWIE E +A I+ A + F + I
Sbjct: 183 IVAPWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNI 362
Query: 1481 VCRFGIPRAIVSDN 1494
VC FGIP ++SDN
Sbjct: 363 VC*FGIPNTLISDN 404
>TC211815
Length = 704
Score = 79.0 bits (193), Expect(2) = 4e-28
Identities = 50/144 (34%), Positives = 77/144 (52%), Gaps = 3/144 (2%)
Frame = +2
Query: 1566 QSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRE 1625
QSTT ETPF + YG+ MLP+E+ E N+ + ++LDL+ + R++ I
Sbjct: 230 QSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIMT 409
Query: 1626 TAMKQRVAAKFNSRVRVRDMQVG---DLVLKWRSGAPGNKLTPNWEGPYRIVKVLGNGAY 1682
A KQR+ FNS++ G + + + +K T N EGP++I NGAY
Sbjct: 410 *AFKQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVRSKFTTN*EGPFKIRHNSKNGAY 589
Query: 1683 HLEELDGRRLPRSFNGLSLRYFYS 1706
LEEL G+ + R +N + L+ +YS
Sbjct: 590 KLEELSGKVVLRIWNSMHLKVYYS 661
Score = 66.2 bits (160), Expect(2) = 4e-28
Identities = 30/67 (44%), Positives = 45/67 (66%)
Frame = +3
Query: 1493 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 1552
DNG QF++ + EF + I+ R SV+HPQTN + E+AN+VIL L++ L A G W++
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLLDGANGRWVE 182
Query: 1553 ELPAVLW 1559
+L +LW
Sbjct: 183 DLVEILW 203
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 91.7 bits (226), Expect(2) = 3e-24
Identities = 60/199 (30%), Positives = 100/199 (50%), Gaps = 5/199 (2%)
Frame = +2
Query: 1194 LKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPR 1253
++ A + ++ L + DS LV +Q++G + +DPNLI Y ++ L F E+ +V
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1254 AENQRADALAKLASTRK--PGNNKSVIQETLAYPSIEGELMACVNRGRTWMDPIISILAG 1311
ENQ ADALA L S + P + I+ L+ G+ W I +
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLVEEERDGKPWYFDIKRYVES 565
Query: 1312 DPAEVEQCTKEQQRE---ASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCA 1368
+E +++R+ A+ + + LY+R LL CV+ ++ E ++ EVHEG
Sbjct: 566 KEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFG 745
Query: 1369 SHIGGRSLACKVLRAGFYW 1387
+H G ++A K+LRAG+YW
Sbjct: 746 THSNGHAMARKILRAGYYW 802
Score = 40.4 bits (93), Expect(2) = 3e-24
Identities = 19/62 (30%), Positives = 31/62 (49%)
Frame = +1
Query: 1128 VVELTPDRFERVDTQWTLFVDGSSNSSGSGAGVTLEGPGELVLEQSLKFEFKATNNQAEY 1187
++ L ++ + +W ++ D +SN G G G L P + + + F TNN AEY
Sbjct: 7 IMALFEEKLDEDRDKWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEY 186
Query: 1188 EA 1189
EA
Sbjct: 187 EA 192
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 106 bits (264), Expect = 1e-22
Identities = 51/91 (56%), Positives = 67/91 (73%)
Frame = -3
Query: 705 HRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 764
H+LA+ VK V+Q +R++ ++ + V QEV KL A FIR++ Y T L +VVMVKK NG
Sbjct: 277 HKLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNG 98
Query: 765 KWRMCVDYTDLNKACPKDSYPLPSIDSLVDG 795
KWR+C DY DLN ACPKD+YPLP+ID + DG
Sbjct: 97 KWRICTDYIDLN*ACPKDAYPLPNIDHMTDG 5
>AW184779
Length = 432
Score = 64.3 bits (155), Expect(2) = 3e-19
Identities = 28/69 (40%), Positives = 41/69 (58%)
Frame = +1
Query: 1337 LYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMD 1396
LY+R LL+CV + E ++ EVHEG +H ++A K+LR G+YW T+ DC
Sbjct: 16 LYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDCCI 195
Query: 1397 FVKQCKECQ 1405
V +C +CQ
Sbjct: 196 HVWKCHKCQ 222
Score = 50.8 bits (120), Expect(2) = 3e-19
Identities = 26/70 (37%), Positives = 38/70 (54%), Gaps = 10/70 (14%)
Frame = +3
Query: 1414 PPKELVTMSAPWPFA---------MWGVDLVGPF-PTARAQMKFILVAVDYFTKWIEAEP 1463
P + + M P P+ MWG+D++G P A FILVA+DYFTKW+EA
Sbjct: 219 PVRPSLIMLTPHPYL*MSWQHLGHMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVS 398
Query: 1464 LAKITSAKIV 1473
A +T + ++
Sbjct: 399 YASVTRSVVI 428
Score = 30.0 bits (66), Expect = 9.1
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +2
Query: 1405 QVFADLSKAPPKELVTMSAPWPF 1427
Q FAD APP L ++APWP+
Sbjct: 224 QTFADNVNAPPIPLNVLAAPWPY 292
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 85.9 bits (211), Expect = 1e-16
Identities = 57/184 (30%), Positives = 94/184 (50%), Gaps = 6/184 (3%)
Frame = +1
Query: 1529 ESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLP--V 1586
E+AN+ I + +++ K W + LP L Y T+ +++T TPF + YG++A+LP V
Sbjct: 1 EAANKNIKKIIQKMTVSYKD-WHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 1587 EIDNFTWRTRPGFEEENQANMAV-ELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDM 1645
E+ + G +E A +L+L+ R A +QR+ + F+ +V +R
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 1646 QVGDLVLKWRSGAPGN---KLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLR 1702
GDLVLK S A + K PN+EGP+ + + GA L +DG LP N ++
Sbjct: 358 HEGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVVK 537
Query: 1703 YFYS 1706
+Y+
Sbjct: 538 RYYA 549
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 84.3 bits (207), Expect = 4e-16
Identities = 44/113 (38%), Positives = 64/113 (55%), Gaps = 4/113 (3%)
Frame = +3
Query: 651 EETKALKFGD----RTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHR 706
EET+ + G R +KIGT +T L LL + D+FAWS +DMPG+ + + HR
Sbjct: 978 EETELVDLGSGSGKREVKIGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQHR 1157
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMV 759
L LNP PV Q RR+ + +++EV K A F+ +YP W+AN+V +
Sbjct: 1158 LPLNPECSPVKQKLRRMKPETSLKIKEEVKK*FDAGFLAVARYPKWVANIVPI 1316
>BI321666
Length = 430
Score = 81.6 bits (200), Expect = 3e-15
Identities = 41/126 (32%), Positives = 69/126 (54%)
Frame = +2
Query: 1376 LACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLV 1435
++ VL++ FY P++ KD + C +CQ +SK L T+ F WG+D V
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFV 181
Query: 1436 GPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNG 1495
GPFP + + ++ILV VDY +KW+EA K + ++ F K+I R G+P ++ + G
Sbjct: 182 GPFPPSFSN-EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGG 358
Query: 1496 TQFSSS 1501
+ ++
Sbjct: 359 SHLCNA 376
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 58.2 bits (139), Expect(2) = 3e-12
Identities = 43/162 (26%), Positives = 73/162 (44%), Gaps = 1/162 (0%)
Frame = +1
Query: 942 NVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPP 1001
+V +++ G + RF+P + P + ++KN+AF W + E+AF LKE L+ P
Sbjct: 109 SVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKAP 288
Query: 1002 ILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALA 1061
+L+ P L S + +V+LQ G H I YF S L A + Y +K A
Sbjct: 289 VLALPDFSKTFELECDASGVGVRAVLLQ---GGHPIAYF-SEKLHSATLNYPTYDKELYA 456
Query: 1062 VLVTARRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAW 1102
++ + + + +D L+ + K L+ R W
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKW 582
Score = 33.5 bits (75), Expect(2) = 3e-12
Identities = 14/33 (42%), Positives = 22/33 (66%)
Frame = +2
Query: 914 FLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
F GF++ G++++PEK KAIQ+ P V E+
Sbjct: 23 FSGFVVGRNGVQMDPEKIKAIQEWPPP*KVWEI 121
>BQ628592
Length = 423
Score = 51.2 bits (121), Expect(2) = 4e-12
Identities = 23/71 (32%), Positives = 42/71 (58%), Gaps = 2/71 (2%)
Frame = -1
Query: 974 LRKNVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDG 1033
L KN A W + +EAF ++K+ L++P +L P+ G P LY + D ++ V++Q D
Sbjct: 423 LPKNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
Query: 1034 --EHRIVYFVS 1042
+ + +Y++S
Sbjct: 243 GKKEQAIYYLS 211
Score = 40.0 bits (92), Expect(2) = 4e-12
Identities = 18/64 (28%), Positives = 36/64 (56%), Gaps = 1/64 (1%)
Frame = -3
Query: 1049 EVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDL-PLRQVLQKPDLSGRLVAWSVELS 1107
++ Y +E+ ++ + RLR Y S + + + P++ + +KP L+GR+ W V LS
Sbjct: 193 KMNYSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLS 14
Query: 1108 EYGL 1111
E+ +
Sbjct: 13 EFNI 2
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 52.4 bits (124), Expect(2) = 8e-11
Identities = 23/61 (37%), Positives = 36/61 (58%)
Frame = +3
Query: 1472 IVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESA 1531
++ F K I RFG+PR ++SD G+ F SQ + K ++ + + HPQTNGQ + +
Sbjct: 18 VIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNGQAKVS 197
Query: 1532 N 1532
N
Sbjct: 198 N 200
Score = 34.3 bits (77), Expect(2) = 8e-11
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = +2
Query: 1541 RRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEI 1588
+ +A ++ W +L LW+ T +++ TPF+M Y LPVE+
Sbjct: 227 KNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVEL 370
>BE802896
Length = 416
Score = 65.5 bits (158), Expect = 2e-10
Identities = 40/133 (30%), Positives = 60/133 (45%)
Frame = -2
Query: 951 GRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKPIQGH 1010
G RF+ + P L+K V F++ +C+ AF LK L + PI+ P
Sbjct: 406 GHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPDWTA 227
Query: 1011 PLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLR 1070
P L S+ AL V+ Q+ID R++Y+ S TL A+ Y EK LA++ +
Sbjct: 226 PFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALEKFH 47
Query: 1071 PYFQSFPVKVRTD 1083
Y + V D
Sbjct: 46 SYLLGTRIIVYID 8
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 62.0 bits (149), Expect = 2e-09
Identities = 39/118 (33%), Positives = 62/118 (52%), Gaps = 3/118 (2%)
Frame = -3
Query: 1508 KEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQS 1567
K+ G+ R ++ HPQTNGQ E +NR I R L + + ++ W L LW++ T ++
Sbjct: 361 KKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAYKA 182
Query: 1568 TTRETPFRMTYGVDAMLPVEIDNFT-WRTRPGFEEENQANMAVELDL--LSETRDEAH 1622
+P+R+ +G LPVEI++ W + +QA +L L L E R EA+
Sbjct: 181 PIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRLEAY 8
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,274,296
Number of Sequences: 63676
Number of extensions: 872116
Number of successful extensions: 4227
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 4133
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4203
length of query: 1706
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1596
effective length of database: 5,635,272
effective search space: 8993894112
effective search space used: 8993894112
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0590c.9


