

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
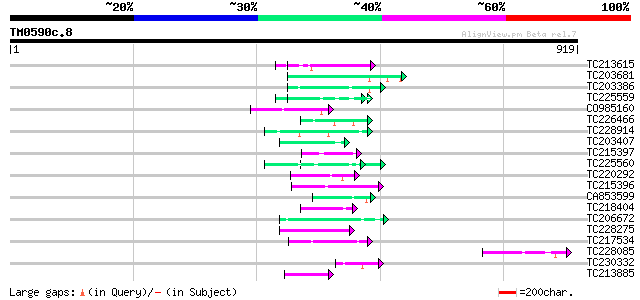
Query= TM0590c.8
(919 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase ... 54 3e-07
TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 51 3e-06
TC203386 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 50 5e-06
TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 48 2e-05
CO985160 47 3e-05
TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {... 47 4e-05
TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (... 47 5e-05
TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pr... 46 7e-05
TC215397 weakly similar to UP|Q8S9J6 (Q8S9J6) AT5g10770/T30N20_4... 45 1e-04
TC225560 similar to UP|Q9FUL8 (Q9FUL8) Arabinogalactan protein, ... 45 1e-04
TC220292 weakly similar to UP|O94096 (O94096) Kexin, partial (4%) 45 1e-04
TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast ... 45 2e-04
CA853599 weakly similar to PIR|F86379|F86 protein F21J9.28 [impo... 44 2e-04
TC218404 UP|Q9MAX7 (Q9MAX7) Epsilon2-COP (Fragment), partial (49%) 44 2e-04
TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyra... 44 2e-04
TC228275 weakly similar to UP|Q9M3D0 (Q9M3D0) Lipase-like protei... 44 2e-04
TC217534 similar to GB|AAB62826.1|2252827|IG005I10 coded for by ... 44 4e-04
TC228085 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (17%) 44 4e-04
TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/... 43 6e-04
TC213885 weakly similar to UP|Q9VUB7 (Q9VUB7) CG32133-PA, partia... 43 7e-04
>TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase
(CBL-interacting protein kinase 4), partial (45%)
Length = 674
Score = 53.9 bits (128), Expect = 3e-07
Identities = 37/144 (25%), Positives = 67/144 (45%)
Frame = +3
Query: 450 AGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATA 509
A T+ P + +T ++ P+ + S +++T P A ASS S+ A AA + S +
Sbjct: 258 ASTTTPTSSKSTRSSPPRPR-----STSSSTSPVA--ASSSPSSPAAAASRSPSPAATSR 416
Query: 510 ASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSC 569
+S+ ++A+ A +S T + TP + R SPPS + + + S P +
Sbjct: 417 SSSPLSASATAMASPTETSSHRTSSSTPPATSRSPTSASPPSRNTSTTVSSTPPAARRPS 596
Query: 570 PGAATTSEAAQIEQAPAPEVGSSS 593
P +++ +A P P +SS
Sbjct: 597 PRQRSSAASATTAXKPTPGPAASS 668
Score = 43.9 bits (102), Expect = 3e-04
Identities = 49/168 (29%), Positives = 75/168 (44%), Gaps = 6/168 (3%)
Frame = +3
Query: 432 AESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPA---- 487
A ST P + T++S P + T+++ +P + + +S AAA + PA
Sbjct: 255 AASTTTPTSSKSTRSSP------PRPRSTSSSTSPVAASSSPSSPAAAASRSPSPAATSR 416
Query: 488 -SSPASAGATA-AFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDA 545
SSP SA ATA A E+S T++S A+S +P T SPP ++
Sbjct: 417 SSSPLSASATAMASPTETSSHRTSSST------PPATSRSP---------TSASPPSRNT 551
Query: 546 PPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
+ ST A PSP ++S +ATT+ + P P SS+
Sbjct: 552 STTVSSTPPAARRPSP---RQRSSAASATTA----XKPTPGPAASSST 674
>TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 913
Score = 50.8 bits (120), Expect = 3e-06
Identities = 50/210 (23%), Positives = 83/210 (38%), Gaps = 17/210 (8%)
Frame = +3
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAA 510
G P A PTT+ A A + A + + P +SP S+ A+ ++ A+
Sbjct: 69 GGQSPAASPTTSPPAATTTPFASPATAPSKPKSPAPVASPTSSSPPASSPNAATATPPAS 248
Query: 511 SAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMP--SPPHQGEKS 568
S V + + A++ P+ +PP P P + P+P SPP S
Sbjct: 249 SPTVASPPSKAAAPAPVATPPAATPPAATPPAATPPAVTPVSSPPAPVPVSSPPAPVPVS 428
Query: 569 CPGAATTSEAAQI------EQAPAPEVGSSSYYNMLPNAIEPSEFLLA----GLNRDVIE 618
P A + A + APAP+ + + A PS LL +
Sbjct: 429 SPPALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHTAPAPSPSLLGPPAPPVGAPGSS 608
Query: 619 KEVLSQGLNVTKE----ETLACLLRA-GCI 643
++ +S G V+++ ET+ CL + GC+
Sbjct: 609 QDSMSPGPAVSEDESGAETIRCLKKVIGCL 698
Score = 39.3 bits (90), Expect = 0.008
Identities = 41/152 (26%), Positives = 60/152 (38%), Gaps = 15/152 (9%)
Frame = +3
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVA--------AATEPTAVP------ASSPA 491
+S +A T+ P A T A+ P K A A VA AAT P A P +S PA
Sbjct: 210 SSPNAATATPPASSPTVASPPS-KAAAPAPVATPPAATPPAATPPAATPPAVTPVSSPPA 386
Query: 492 SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
++ A A + A A S++ P + + +T KS PSP
Sbjct: 387 PVPVSSPPAPVPVSSPPALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHTAPAPSPSL 566
Query: 552 T-HDAGPMPSPPHQGEKSCPGAATTSEAAQIE 582
A P+ +P + PG A + + + E
Sbjct: 567 LGPPAPPVGAPGSSQDSMSPGPAVSEDESGAE 662
>TC203386 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 649
Score = 50.1 bits (118), Expect = 5e-06
Identities = 45/165 (27%), Positives = 62/165 (37%), Gaps = 6/165 (3%)
Frame = +1
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAA 510
G P A PTT++A+P + P A P SSP ++ AA A T A
Sbjct: 64 GGQSPAAAPTTSSASPA--TAPSKPKPKSPAPVASPTSSPPASSPNAATATPPVSSPTVA 237
Query: 511 SAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCP 570
S A A ++ P+ +PP SPP+ P+ SPP S P
Sbjct: 238 SPPSKAAAPAPATTPPVATPPAATPPAATPPAVTPVSSPPA---PVPVSSPPAPVPVSSP 408
Query: 571 GAATTSEAAQI------EQAPAPEVGSSSYYNMLPNAIEPSEFLL 609
A + A + APAP+ + + A PS LL
Sbjct: 409 PALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHTAPAPSPALL 543
Score = 40.4 bits (93), Expect = 0.004
Identities = 41/162 (25%), Positives = 59/162 (36%), Gaps = 7/162 (4%)
Frame = +1
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATE----PT 483
A + +T P T AS + + P T A P A+ A T P
Sbjct: 181 ASSPNAATATPPVSSPTVASPPSKAAAPAPATTPPVATPPAATPPAATPPAVTPVSSPPA 360
Query: 484 AVPASSPAS---AGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSP 540
VP SSP + + A A + A SA V A + T K K++ P
Sbjct: 361 PVPVSSPPAPVPVSSPPALAPTTPAPVVAPSAEVPAPAPKSKKKTK---KSKKHTAPAPS 531
Query: 541 PRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIE 582
P PP+P P+ +P + S PG A + + + E
Sbjct: 532 PALLGPPAP-------PVGAPGSSQDASSPGPAVSEDESGAE 636
>TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (47%)
Length = 870
Score = 47.8 bits (112), Expect = 2e-05
Identities = 48/153 (31%), Positives = 58/153 (37%), Gaps = 5/153 (3%)
Frame = +3
Query: 431 QAEST--KAPKRRRLTKASSDAGT--SKPGAQPTTAAAAPKG-KNVAEASVAAATEPTAV 485
QA ST K+P K+S A T S P A PTT AAP AS AT P A
Sbjct: 195 QAPSTVPKSPAPVASPKSSPPAATPASTPAASPTTTPAAPAPVTKPPAASPPPATPPPAT 374
Query: 486 PASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDA 545
P +P + A SS A A T A S P K+ + +P
Sbjct: 375 PPPAPVPVSSPPAPVPVSSPPAPVPVAA--PTTPVAPSPAPKHKKKGKKHGAPAPSPLLG 548
Query: 546 PPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEA 578
PP+PP+ P P PG A E+
Sbjct: 549 PPAPPT-----GAPGPSEDASSPGPGTAANDES 632
Score = 45.1 bits (105), Expect = 1e-04
Identities = 42/140 (30%), Positives = 55/140 (39%), Gaps = 2/140 (1%)
Frame = +3
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEA--SVAAATEPTAVPASSPASAGATAAFAAESSIGAT 508
G P A P+ A P A+A +V + P A P SSP +A + AA + T
Sbjct: 129 GGQSPAAAPSNTPATPAAATPAQAPSTVPKSPAPVASPKSSPPAATPASTPAASPT---T 299
Query: 509 AASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKS 568
+A TK A+S P P +PP P S P P P P
Sbjct: 300 TPAAPAPVTKPPAASPPPA------TPPPATPPPAPVPVSSP------PAPVPVSSPPAP 443
Query: 569 CPGAATTSEAAQIEQAPAPE 588
P AA T+ A +PAP+
Sbjct: 444 VPVAAPTTPVA---PSPAPK 494
>CO985160
Length = 777
Score = 47.4 bits (111), Expect = 3e-05
Identities = 46/151 (30%), Positives = 66/151 (43%), Gaps = 17/151 (11%)
Frame = -1
Query: 391 LEDMNFTNAELLKARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDA 450
+E TNA A + A S T A + A +T A A++ A
Sbjct: 768 VESAAATNAATTSADAKPTAASSAVATTAAATSHATTATSSAPTTAATTSS--AAAAAAA 595
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASS--PASAGATAAFAAESS---- 504
+S P TT++AA + A+ AAAT +A A++ PA+A AA AA S+
Sbjct: 594 TSSAPTTAATTSSAAAAAATTSSAAAAAATTSSAATAAATGPAAAACAAATAAASANPAT 415
Query: 505 -----------IGATAASAGVNATKAAASSD 524
ATAA++ V AT AAA++D
Sbjct: 414 AAATSTSTTPTTTATAAASAVTATAAAAATD 322
Score = 35.0 bits (79), Expect = 0.15
Identities = 29/65 (44%), Positives = 33/65 (50%), Gaps = 5/65 (7%)
Frame = +2
Query: 462 AAAAPKGKNVAEASVAAA----TEPTAVPASSPASAG-ATAAFAAESSIGATAASAGVNA 516
AAA V E VAAA E AV A+ A+AG AA AAE + A AA+ V A
Sbjct: 362 AAAVAVVVGVVEVEVAAAVAGFAEAAAVAAAQAAAAGPVAAAVAAEEVVAAAAAAEEVVA 541
Query: 517 TKAAA 521
AAA
Sbjct: 542 AAAAA 556
Score = 33.1 bits (74), Expect = 0.57
Identities = 33/121 (27%), Positives = 51/121 (41%), Gaps = 1/121 (0%)
Frame = -1
Query: 474 ASVAAATEPTAVPASSPASAGATA-AFAAESSIGATAASAGVNATKAAASSDTPIGDKEK 532
A+ +A +PTA ++ +A AT+ A A SS TAA+ A AAA+S P
Sbjct: 741 ATTSADAKPTAASSAVATTAAATSHATTATSSAPTTAATTSSAAAAAAATSSAPTTAATT 562
Query: 533 ENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSS 592
+ + A + +T A + G + AA T+ AA A A +S
Sbjct: 561 SSAAAAAATTSSAAAAAATTSSAATAAA---TGPAAAACAAATA-AASANPATAAATSTS 394
Query: 593 S 593
+
Sbjct: 393 T 391
Score = 29.3 bits (64), Expect = 8.3
Identities = 21/63 (33%), Positives = 28/63 (44%)
Frame = +2
Query: 462 AAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAA 521
AAA + A VAAA V A++ A+ AA AA + A A A AAA
Sbjct: 434 AAAVAAAQAAAAGPVAAAVAAEEVVAAAAAAEEVVAAAAAAEEVVAAVVGAEEVAAAAAA 613
Query: 522 SSD 524
+ +
Sbjct: 614 AEE 622
>TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {Arabidopsis
thaliana;} , partial (22%)
Length = 978
Score = 47.0 bits (110), Expect = 4e-05
Identities = 37/126 (29%), Positives = 50/126 (39%), Gaps = 10/126 (7%)
Frame = +1
Query: 472 AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSD----TPI 527
A ASV A + PT + A+SP+S G A +S T+ ++ A + TP
Sbjct: 61 APASVPANSPPTTLAATSPSS-GTPPAVTQAASPPTTSNPPTISQPPAIVAPKPAPVTPP 237
Query: 528 GDKEKENETPKSPPRQDAPPSPPSTHDAG------PMPSPPHQGEKSCPGAATTSEAAQI 581
K +PK+P Q PP PP P+P PP P A+
Sbjct: 238 APKVAPASSPKAPTPQTPPPQPPKVSPVAAPTLPPPLPPPPVATPPPLPPPKIAPTPAKT 417
Query: 582 EQAPAP 587
APAP
Sbjct: 418 SPAPAP 435
>TC228914 similar to UP|DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (Dach1),
partial (3%)
Length = 784
Score = 46.6 bits (109), Expect = 5e-05
Identities = 44/189 (23%), Positives = 75/189 (39%), Gaps = 14/189 (7%)
Frame = +3
Query: 414 PKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPK------ 467
P +T + G A + + S A KRR S A ++ P T++ AAP
Sbjct: 171 PSPSTTPTSSAPGNAPHSSPS--ASKRRPTPSGPSSAASTSPKPTSTSSRAAPSKSPFTW 344
Query: 468 -GKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGV-------NATKA 519
++ + + A+ P P +S +S + A+ A SS TA++ + T
Sbjct: 345 PSASLGTSMSSPASPPPPAPNASTSSTTSAASPALASSAANTASATTAPSLPSIPSRTTP 524
Query: 520 AASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAA 579
+ TP + +P + P++ SP + + SPP E P A T
Sbjct: 525 TTARSTPSSSNPTSSTSPTATPKRTRVSSPTPSSSSTSRSSPPSPKE---PTATVTVSHT 695
Query: 580 QIEQAPAPE 588
+ E+A P+
Sbjct: 696 RGEEANTPQ 722
Score = 32.0 bits (71), Expect = 1.3
Identities = 19/57 (33%), Positives = 25/57 (43%)
Frame = +3
Query: 537 PKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
P + PR +PPS P PSP S PG A S + ++ P P SS+
Sbjct: 129 PPASPRSSSPPSSP--------PSPSTTPTSSAPGNAPHSSPSASKRRPTPSGPSSA 275
Score = 30.8 bits (68), Expect = 2.8
Identities = 32/151 (21%), Positives = 56/151 (36%), Gaps = 3/151 (1%)
Frame = +3
Query: 453 SKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASA 512
S+ + TTA + AS +++ P++ P+ S + A SS A+
Sbjct: 72 SQTPTRTTTATRRTTTSTLPPASPRSSSPPSSPPSPSTTPTSSAPGNAPHSSPSASKRRP 251
Query: 513 GVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDA---GPMPSPPHQGEKSC 569
+ +AAS+ + P S + AP P T + G S P
Sbjct: 252 TPSGPSSAAST----------SPKPTSTSSRAAPSKSPFTWPSASLGTSMSSPASPPPPA 401
Query: 570 PGAATTSEAAQIEQAPAPEVGSSSYYNMLPN 600
P A+T+S + A A +++ P+
Sbjct: 402 PNASTSSTTSAASPALASSAANTASATTAPS 494
>TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(AtAGP4) (AT5g10430/F12B17_220), partial (50%)
Length = 966
Score = 46.2 bits (108), Expect = 7e-05
Identities = 36/115 (31%), Positives = 42/115 (36%)
Frame = +2
Query: 437 APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGAT 496
AP + T S S P PTT A P A+ A P +PA A T
Sbjct: 206 APSQPPTTTPSPPPPRSAPAPAPTTPATPPPATPPPAATPPPAATPPPAATPTPAPAPPT 385
Query: 497 AAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
AA SS A+ S T + S +TP G P P A P PPS
Sbjct: 386 AAPTPASSPAASPPSPSPTVTPSPTSPNTPPG--------PSPGPSGSAEPPPPS 526
Score = 41.6 bits (96), Expect = 0.002
Identities = 40/137 (29%), Positives = 54/137 (39%), Gaps = 3/137 (2%)
Frame = +2
Query: 448 SDAGTSKPGAQPTTAAAAPKGKNV-AEASVAAATEPTAVP--ASSPASAGATAAFAAESS 504
+ A + P PTT + P ++ A A AT P A P A++P A A +
Sbjct: 188 AQAPGAAPSQPPTTTPSPPPPRSAPAPAPTTPATPPPATPPPAATPPPAATPPPAATPTP 367
Query: 505 IGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQ 564
A +A A+ AAS +P TP SP + PP GP P P
Sbjct: 368 APAPPTAAPTPASSPAASPPSP-----SPTVTP-SPTSPNTPP--------GPSPGPSGS 505
Query: 565 GEKSCPGAATTSEAAQI 581
E P AA ++ A I
Sbjct: 506 AEPPPPSAAFSASKAFI 556
Score = 37.0 bits (84), Expect = 0.040
Identities = 27/96 (28%), Positives = 38/96 (39%)
Frame = +2
Query: 492 SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
S T F+ + GA + ++ + P + T SPP + P+P
Sbjct: 95 STATTTTFSLDMGSGAVQLFLILGLLASSCLAQAPGAAPSQPPTTTPSPPPPRSAPAPAP 274
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
T A P P+ P + P AAT AA APAP
Sbjct: 275 TTPATPPPATPPPA-ATPPPAATPPPAATPTPAPAP 379
Score = 29.6 bits (65), Expect = 6.3
Identities = 25/119 (21%), Positives = 36/119 (30%)
Frame = +2
Query: 474 ASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKE 533
AS A P A P+ P + + + + T + AT A++ P
Sbjct: 173 ASSCLAQAPGAAPSQPPTTTPSPPPPRSAPAPAPTTPATPPPATPPPAATPPPAA----- 337
Query: 534 NETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSS 592
+PP P P+ A P P+ P T P P G S
Sbjct: 338 -----TPPPAATPTPAPAPPTAAPTPASSPAASPPSPSPTVTPSPTSPNTPPGPSPGPS 499
>TC215397 weakly similar to UP|Q8S9J6 (Q8S9J6) AT5g10770/T30N20_40, partial
(76%)
Length = 1447
Score = 45.4 bits (106), Expect = 1e-04
Identities = 31/97 (31%), Positives = 44/97 (44%)
Frame = +1
Query: 474 ASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKE 533
A+ AA+ P P SS S+ A A A+SA A+ A+A++ +P +K
Sbjct: 394 ATSAASASP*PPPTSSTTSSSAAARTTR-------ASSAAPPASLASAATPSPSSNKLPP 552
Query: 534 NETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCP 570
N SP PP+PP+T + P PP S P
Sbjct: 553 NTVKSSPTASPPPPAPPATSPSAP---PPQAATSSTP 654
Score = 35.4 bits (80), Expect = 0.12
Identities = 36/147 (24%), Positives = 56/147 (37%), Gaps = 17/147 (11%)
Frame = +1
Query: 432 AESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPK-GKNVAEASVAAATEPTAVPASSP 490
+ +T + R T+ASS A + + T + ++ K N ++S A+ P A PA+SP
Sbjct: 436 SSTTSSSAAARTTRASSAAPPASLASAATPSPSSNKLPPNTVKSSPTASPPPPAPPATSP 615
Query: 491 ASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD------ 544
+ A AA SS + S + + +P ++ P SPP
Sbjct: 616 S---APPPQAATSSTPPSPPSPAAPPSTVSILLPSP*AVSNSQSPPPPSPPAVQSSTLAP 786
Query: 545 ----------APPSPPSTHDAGPMPSP 561
AP PPS P P
Sbjct: 787 *SPASPQPPTAPSDPPSVKACQNTPPP 867
Score = 35.0 bits (79), Expect = 0.15
Identities = 42/171 (24%), Positives = 64/171 (36%), Gaps = 4/171 (2%)
Frame = +1
Query: 438 PKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATA 497
P R T +S+ S+ +A+ P + +S AAA A A+ PAS + A
Sbjct: 340 PPRHAYTASSTATVPSQLATSAASASP*PPPTSSTTSSSAAARTTRASSAAPPASLASAA 519
Query: 498 AFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDA----PPSPPSTH 553
+ S+ N K++ ++ P +P +PP Q A PPSPPS
Sbjct: 520 TPSPSSN------KLPPNTVKSSPTASPP--PPAPPATSPSAPPPQAATSSTPPSPPSP- 672
Query: 554 DAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEP 604
+PP P S + P V SS+ P + +P
Sbjct: 673 -----AAPPSTVSILLPSP*AVSNSQSPPPPSPPAVQSSTLAP*SPASPQP 810
>TC225560 similar to UP|Q9FUL8 (Q9FUL8) Arabinogalactan protein, partial
(48%)
Length = 1030
Score = 45.4 bits (106), Expect = 1e-04
Identities = 46/164 (28%), Positives = 61/164 (37%)
Frame = +2
Query: 413 SPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVA 472
+P T A + S K+ K ++ S P A PTT AAP A
Sbjct: 170 TPAAATPAQAPSTPNSPAPVASPKSSPPASSPKPATATPASSPAASPTTTPAAPAP---A 340
Query: 473 EASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK 532
AA+ P VP SSP + SS A A A A + + K K
Sbjct: 341 TKPPAASPPPAPVPVSSPPAP------VPVSSPPAPVPVAAPTTPVAPAPAPSKHKKKGK 502
Query: 533 ENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTS 576
++ P P PP+PP+ P PS + S PG AT +
Sbjct: 503 KHGAPAPSPSLLGPPAPPT---GAPGPSE----DASSPGPATAA 613
Score = 42.7 bits (99), Expect = 7e-04
Identities = 37/138 (26%), Positives = 51/138 (36%)
Frame = +2
Query: 472 AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKE 531
A S AT A PA +P++ + A A+ S ++ AT A++ + +P
Sbjct: 146 AAPSNTPATPAAATPAQAPSTPNSPAPVASPKSSPPASSPKPATATPASSPAASP----- 310
Query: 532 KENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGS 591
TP +P PP+ P+ SPP S P A A APAP
Sbjct: 311 --TTTPAAPAPATKPPAASPPPAPVPVSSPPAPVPVSSPPAPVPVAAPTTPVAPAPAPSK 484
Query: 592 SSYYNMLPNAIEPSEFLL 609
A PS LL
Sbjct: 485 HKKKGKKHGAPAPSPSLL 538
Score = 40.0 bits (92), Expect = 0.005
Identities = 47/167 (28%), Positives = 61/167 (36%), Gaps = 25/167 (14%)
Frame = +2
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEA--------------SVAAATEP---TAVPASSPASA 493
G P A P+ A P A+A S A+ P TA PASSPA++
Sbjct: 128 GGQSPAAAPSNTPATPAAATPAQAPSTPNSPAPVASPKSSPPASSPKPATATPASSPAAS 307
Query: 494 GATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSP-PST 552
T A + AAS A +S P+ P + P P+P PS
Sbjct: 308 PTTTPAAPAPATKPPAASP-PPAPVPVSSPPAPVPVSSPPAPVPVAAPTTPVAPAPAPSK 484
Query: 553 H------DAGPMPSPPHQGEKSCP-GAATTSEAAQIEQAPAPEVGSS 592
H P PSP G + P GA SE A +P P ++
Sbjct: 485 HKKKGKKHGAPAPSPSLLGPPAPPTGAPGPSEDA---SSPGPATAAN 616
Score = 32.3 bits (72), Expect = 0.98
Identities = 25/77 (32%), Positives = 31/77 (39%), Gaps = 1/77 (1%)
Frame = +2
Query: 512 AGVNA-TKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCP 570
AGV + AAA S+TP S P AP + P + P P S P
Sbjct: 119 AGVGGQSPAAAPSNTPATPAAATPAQAPSTPNSPAPVASPKSSPPASSPKPATATPASSP 298
Query: 571 GAATTSEAAQIEQAPAP 587
A+ T+ A APAP
Sbjct: 299 AASPTTTPA----APAP 337
>TC220292 weakly similar to UP|O94096 (O94096) Kexin, partial (4%)
Length = 1094
Score = 45.1 bits (105), Expect = 1e-04
Identities = 35/121 (28%), Positives = 50/121 (40%), Gaps = 9/121 (7%)
Frame = +1
Query: 455 PGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGV 514
P AQP AA + A + +++T T SSP++ +A+ + T A +
Sbjct: 394 P*AQPQHAATRRTKSDAAAPTPSSSTRLTTPTRSSPSTRTPSAS*SDPLRC*RTRACPPI 573
Query: 515 NATKAAASSDTPIGDKEKENETPK---------SPPRQDAPPSPPSTHDAGPMPSPPHQG 565
T+ SS TP + TP S PR APP P+ + P P PPH
Sbjct: 574 KCTRG--SSSTPPSPSTSQAPTPSCISTAPPRSSSPRSTAPPPAPAIPTSRPPPPPPHAR 747
Query: 566 E 566
E
Sbjct: 748 E 750
>TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast nucleoid DNA
binding protein (CND41), partial (69%)
Length = 1808
Score = 44.7 bits (104), Expect = 2e-04
Identities = 38/149 (25%), Positives = 61/149 (40%), Gaps = 1/149 (0%)
Frame = +1
Query: 458 QPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNAT 517
QP A V S A + + P P S+ +++ AA+++ + A A+
Sbjct: 670 QPPQKHAYTASSTVTVRSPLATSAASGSP*PPPTSSTTSSSVAAKTTRASLVAPP---AS 840
Query: 518 KAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPG-AATTS 576
A+A++ +P +K SP PP+PP+ + P P P S P AA S
Sbjct: 841 SASAATPSPSSNKPPPYIVKSSPTASPPPPAPPAASPSAPPPLPTSSTPPSPPSPAAPPS 1020
Query: 577 EAAQIEQAPAPEVGSSSYYNMLPNAIEPS 605
A+ +P S S P A++ S
Sbjct: 1021TASTSPASPWAAPNSQSPPPPSPPAVQSS 1107
Score = 40.8 bits (94), Expect = 0.003
Identities = 43/165 (26%), Positives = 68/165 (41%), Gaps = 2/165 (1%)
Frame = +1
Query: 432 AESTKAPKRRRLTKASSDA--GTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASS 489
+ +T + + T+AS A +S A P+ ++ P + ++S A+ P A PA+S
Sbjct: 772 SSTTSSSVAAKTTRASLVAPPASSASAATPSPSSNKPP-PYIVKSSPTASPPPPAPPAAS 948
Query: 490 PASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSP 549
P++ + S + AA +T A+ P +SPP PPSP
Sbjct: 949 PSAPPPLPTSSTPPSPPSPAAPPSTASTSPASPWAAP---------NSQSPP----PPSP 1089
Query: 550 PSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSY 594
P+ + P P P P +A S A P P G+S Y
Sbjct: 1090PAVQSSTPAP*SPASPPPPTPPSAPPSGKA-CPNTPPP--GNSPY 1215
Score = 35.0 bits (79), Expect = 0.15
Identities = 34/136 (25%), Positives = 52/136 (38%), Gaps = 2/136 (1%)
Frame = +1
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAA-TEPTAVP 486
A+ S AP + A+ ++KP P ++P A AA+ + P +P
Sbjct: 799 AKTTRASLVAPPASSASAATPSPSSNKP--PPYIVKSSPTASPPPPAPPAASPSAPPPLP 972
Query: 487 ASS-PASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDA 545
SS P S + AA + +S + A N+ S + +P SPP
Sbjct: 973 TSSTPPSPPSPAAPPSTASTSPASPWAAPNSQSPPPPSPPAVQSSTPAP*SPASPPPPTP 1152
Query: 546 PPSPPSTHDAGPMPSP 561
P +PPS P P
Sbjct: 1153PSAPPSGKACPNTPPP 1200
>CA853599 weakly similar to PIR|F86379|F86 protein F21J9.28 [imported] -
Arabidopsis thaliana, partial (58%)
Length = 389
Score = 44.3 bits (103), Expect = 2e-04
Identities = 37/111 (33%), Positives = 43/111 (38%), Gaps = 8/111 (7%)
Frame = -1
Query: 491 ASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPP 550
A GA A S T SA + AT AAS+ + TP R PSP
Sbjct: 389 AGDGAGAGQRRRRSWSWTT*SARLTATAMAASASRSSSSSTQRASTPTRSWRTSRTPSPS 210
Query: 551 STHDAGPMPSPPHQGEKSCPGAATTS--------EAAQIEQAPAPEVGSSS 593
ST A PSPP SC +A T+ AA + A AP SS
Sbjct: 209 STSTA-TAPSPPRSSTPSCAASARTAPSPSAAG*SAASMATATAPSTSRSS 60
>TC218404 UP|Q9MAX7 (Q9MAX7) Epsilon2-COP (Fragment), partial (49%)
Length = 485
Score = 44.3 bits (103), Expect = 2e-04
Identities = 28/93 (30%), Positives = 42/93 (45%)
Frame = +2
Query: 472 AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKE 531
A ++ T+P + A+SP S T + A SS AT+ SA +++ + + P K
Sbjct: 86 ATTCISGRTKPPSTAATSPTSPKKTPSSATPSSTAATSPSASFSSSSPRSMTTLPHLSKP 265
Query: 532 KENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQ 564
+ SPP P PPS +P PP Q
Sbjct: 266 SNSSLSISPP--PTPRIPPSRVSRSGLPIPPLQ 358
>TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyrate
dehydrogenase , partial (76%)
Length = 1230
Score = 44.3 bits (103), Expect = 2e-04
Identities = 50/180 (27%), Positives = 70/180 (38%), Gaps = 3/180 (1%)
Frame = +2
Query: 438 PKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTA-VPAS-SPASAGA 495
P R T SS GTS P P A A AS A+ PT+ VPA+ SP+S
Sbjct: 8 PNSARGTLPSS--GTSWPRRSPL*AHLTRAWAGSARASWASPCAPTSSVPATPSPSSTAP 181
Query: 496 TAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDA 555
+ S+ G T+ + A SS ++ +PP +PPS P+ +
Sbjct: 182 PPRPSPSSTSGPTSPPPPTPSQPAPTSSSPSSATPRTSAQSSSTPPPAPSPPSAPAASSS 361
Query: 556 G-PMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNR 614
P P+PP + P +AA P P ++ EP + AG NR
Sbjct: 362 T*PPPNPPSPSKSRTP---LPPKAATQSTPPCPAATAA-------RRTEP*QSSPAGKNR 511
>TC228275 weakly similar to UP|Q9M3D0 (Q9M3D0) Lipase-like protein, partial
(25%)
Length = 1153
Score = 44.3 bits (103), Expect = 2e-04
Identities = 32/127 (25%), Positives = 59/127 (46%), Gaps = 4/127 (3%)
Frame = +3
Query: 437 APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVP-ASSPASAGA 495
A R++ T T P ++P TA ++P +++ ++ + P+ P A++P AG+
Sbjct: 33 ATPRKKNTTPPKAFATPSPTSKPPTAKSSPSPSSLSTSNPTKSKPPSL*PTATAPTPAGS 212
Query: 496 TAAFAAESSIGATAASAGVNATKAA---ASSDTPIGDKEKENETPKSPPRQDAPPSPPST 552
+ A+ S GAT +S ++ AA +S+ + K +P S A P+ S
Sbjct: 213 SRKSASTSPPGATPSSPPTSSATAAPTVSSATSATWTKSPPPPSPSSSTSAIATPTKTSR 392
Query: 553 HDAGPMP 559
H + P
Sbjct: 393 HSSSASP 413
Score = 39.3 bits (90), Expect = 0.008
Identities = 36/136 (26%), Positives = 53/136 (38%), Gaps = 1/136 (0%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATK 518
P T+ A P+ KN A PT+ P ++ +S + SS+ + N TK
Sbjct: 18 PLTSGATPRKKNTTPPKAFATPSPTSKPPTAKSSP-------SPSSLSTS------NPTK 158
Query: 519 AAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMP-SPPHQGEKSCPGAATTSE 577
+ S P TP R+ A SPP G P SPP + P ++ +
Sbjct: 159 SKPPSL*PTATAP----TPAGSSRKSASTSPP-----GATPSSPPTSSATAAPTVSSATS 311
Query: 578 AAQIEQAPAPEVGSSS 593
A + P P SS+
Sbjct: 312 ATWTKSPPPPSPSSST 359
Score = 37.4 bits (85), Expect = 0.030
Identities = 35/136 (25%), Positives = 57/136 (41%), Gaps = 10/136 (7%)
Frame = +3
Query: 423 KKRGGAENQAESTKAPKRRRLTKASSDAGTS----------KPGAQPTTAAAAPKGKNVA 472
+K+ +A +T +P + T SS + +S P PT A P G +
Sbjct: 42 RKKNTTPPKAFATPSPTSKPPTAKSSPSPSSLSTSNPTKSKPPSL*PTATAPTPAGSSRK 221
Query: 473 EASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK 532
AS T P SSP ++ ATAA S+ AT + + ++++S K
Sbjct: 222 SAS----TSPPGATPSSPPTSSATAAPTVSSATSATWTKSPPPPSPSSSTSAIATPTKTS 389
Query: 533 ENETPKSPPRQDAPPS 548
+ + S P +D+P S
Sbjct: 390 RHSSSAS-PWEDSPRS 434
>TC217534 similar to GB|AAB62826.1|2252827|IG005I10 coded for by A. thaliana
cDNA N96903 {Arabidopsis thaliana;} , partial (98%)
Length = 1147
Score = 43.5 bits (101), Expect = 4e-04
Identities = 37/137 (27%), Positives = 60/137 (43%), Gaps = 2/137 (1%)
Frame = +3
Query: 453 SKPGAQPTTAAAAPK--GKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAA 510
S P +A+++P + + + A + PT AS+ +S+ ATA + AT+
Sbjct: 135 SSPSRGSPSASSSPS*GSTSTSLSQNALSATPTRCSAST-SSSAATALLRLPPAPPATST 311
Query: 511 SAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCP 570
SA +SS P + + T + PPS PS A P+PP +S
Sbjct: 312 SATTAPLSTPSSSPLPSAARSPASPTASASSPASFPPSLPSPSAATAPPTPPE--SRSSS 485
Query: 571 GAATTSEAAQIEQAPAP 587
AT+S A + ++A P
Sbjct: 486 RGATSSSARRAQRAVNP 536
>TC228085 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (17%)
Length = 947
Score = 43.5 bits (101), Expect = 4e-04
Identities = 43/153 (28%), Positives = 69/153 (44%), Gaps = 9/153 (5%)
Frame = +1
Query: 767 LAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLED 826
L + EK+ E L + + D+E + ++ + EA +AE A++ + + LE
Sbjct: 1 LEKDEKRDPEKLRIEREDLERRQKEEKARLQAEAKAAEEAQRKAEAEAAAEAKRKRELER 180
Query: 827 ELATEKAKAIEAREQAADIALDNR--ERGFYLAKDQAQHLFPNFDFSAMGVMKEITAA-- 882
E A + A++ E+ DI ++ E L+ +HL P+F KE T+A
Sbjct: 181 EAARQ---ALQKMEKTVDINENSHFLEDLEMLSAVHDEHL-PSF--------KEETSADQ 324
Query: 883 -----GLVGPDDPPLIDQNLWTATEEEEEEEEE 910
G + PL L+ EEEEEEEEE
Sbjct: 325 PQDGLGGIKLQGNPLEQLGLYMKEEEEEEEEEE 423
>TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/k19m22_160),
partial (36%)
Length = 827
Score = 43.1 bits (100), Expect = 6e-04
Identities = 31/89 (34%), Positives = 39/89 (42%), Gaps = 11/89 (12%)
Frame = -3
Query: 529 DKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEK---SCP--------GAATTSE 577
D N +P SPP APP PP T P P+PPH+G + SCP A T+
Sbjct: 459 DSSPPNPSP-SPP-SSAPPPPPRTVPLPPSPTPPHRGRRHLFSCPSSSRRRRRSAGTSP* 286
Query: 578 AAQIEQAPAPEVGSSSYYNMLPNAIEPSE 606
A + +AP P + P A P E
Sbjct: 285 ARRASEAPPPPPRQQLASSPGPTANAPPE 199
>TC213885 weakly similar to UP|Q9VUB7 (Q9VUB7) CG32133-PA, partial (4%)
Length = 540
Score = 42.7 bits (99), Expect = 7e-04
Identities = 30/83 (36%), Positives = 48/83 (57%), Gaps = 4/83 (4%)
Frame = +1
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASV-AAATEPTAVPASSPASAGATAAFAAES- 503
A++ A TS ++AAAA + A A+ ++AT PA++ ++ ATAA A+ S
Sbjct: 64 ATTAAATSHAATTTSSAAAAAAASSAATAAATSSATTAATAPAAAASATPATAAAASTST 243
Query: 504 --SIGATAASAGVNATKAAASSD 524
+ ATAA++ V AT A A++D
Sbjct: 244 TPTTTATAAASAVAATAATAATD 312
Score = 42.4 bits (98), Expect = 0.001
Identities = 30/97 (30%), Positives = 43/97 (43%), Gaps = 1/97 (1%)
Frame = +1
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAAT-EPTAVP 486
A + A +T A +S A + + TAAA A A AAA+ P
Sbjct: 46 ATSSAVATTAAATSHAATTTSSAAAAAAASSAATAAATSSATTAATAPAAAASATPATAA 225
Query: 487 ASSPASAGATAAFAAESSIGATAASAGVNATKAAASS 523
A+S ++ T A AA S++ ATAA+A + SS
Sbjct: 226 AASTSTTPTTTATAAASAVAATAATAATDGWGRNGSS 336
Score = 40.0 bits (92), Expect = 0.005
Identities = 27/81 (33%), Positives = 42/81 (51%), Gaps = 2/81 (2%)
Frame = +1
Query: 448 SDAGTSKPGAQPT--TAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSI 505
S A T+ A+PT ++A A + A+ ++ A ASS A+A AT++ ++
Sbjct: 7 SAAATAPADAKPTATSSAVATTAAATSHAATTTSSAAAAAAASSAATAAATSSATTAATA 186
Query: 506 GATAASAGVNATKAAASSDTP 526
A AASA AA++S TP
Sbjct: 187 PAAAASATPATAAAASTSTTP 249
Score = 38.9 bits (89), Expect = 0.010
Identities = 26/74 (35%), Positives = 38/74 (51%), Gaps = 2/74 (2%)
Frame = +1
Query: 452 TSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASA--GATAAFAAESSIGATA 509
T+ A TTAAA ++ AAA +A A++ +SA ATA AA S+ ATA
Sbjct: 43 TATSSAVATTAAATSHAATTTSSAAAAAAASSAATAAATSSATTAATAPAAAASATPATA 222
Query: 510 ASAGVNATKAAASS 523
A+A + T ++
Sbjct: 223 AAASTSTTPTTTAT 264
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,967,348
Number of Sequences: 63676
Number of extensions: 659771
Number of successful extensions: 12026
Number of sequences better than 10.0: 620
Number of HSP's better than 10.0 without gapping: 6655
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9737
length of query: 919
length of database: 12,639,632
effective HSP length: 106
effective length of query: 813
effective length of database: 5,889,976
effective search space: 4788550488
effective search space used: 4788550488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0590c.8


