

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.5
(441 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
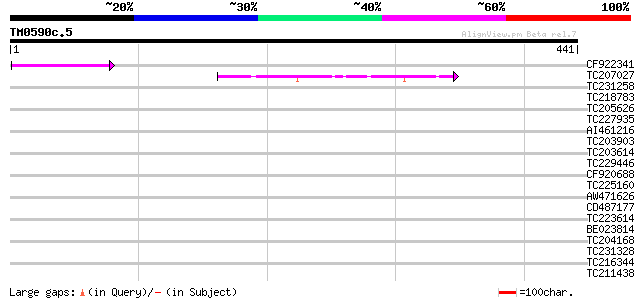
Score E
Sequences producing significant alignments: (bits) Value
CF922341 46 3e-05
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 43 3e-04
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 40 0.002
TC218783 similar to UP|Q40363 (Q40363) NuM1 protein, partial (19%) 36 0.039
TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g7539... 33 0.20
TC227935 32 0.44
AI461216 homologue to GP|13702121|dbj SA2112 {Staphylococcus aur... 32 0.57
TC203903 homologue to UP|Q94LX2 (Q94LX2) Beta-conglycinin alpha ... 31 1.3
TC203614 UP|Q94LX2 (Q94LX2) Beta-conglycinin alpha subunit, comp... 31 1.3
TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 31 1.3
CF920688 31 1.3
TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, comp... 30 1.7
AW471626 similar to SP|P11827|GLCX Beta-conglycinin alpha' chai... 30 2.2
CD487177 homologue to GP|10946375|gb| ribulose-1 5-bisphosphate ... 30 2.2
TC223614 similar to UP|Q950Y2 (Q950Y2) Ymf62, partial (6%) 30 2.2
BE023814 30 2.8
TC204168 UP|Q7XXT2 (Q7XXT2) Prepro beta-conglycinin alpha prime ... 30 2.8
TC231328 GB|AAF34631.1|6979760|AF215712 protamine 2 {Gorilla gor... 29 3.7
TC216344 weakly similar to UP|Q9SMQ5 (Q9SMQ5) Proliferating-cell... 29 3.7
TC211438 UP|GLCX_SOYBN (P11827) Beta-conglycinin, alpha' chain p... 28 8.2
>CF922341
Length = 675
Score = 46.2 bits (108), Expect = 3e-05
Identities = 22/80 (27%), Positives = 36/80 (44%)
Frame = +1
Query: 2 PDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPR 61
P K L D Y G + PK+HL + KM A + + F + A+ W+T L
Sbjct: 406 PPKFKVLDFDKYKGTTCPKNHLKMYCQKMGAYAKDEELLIHSFQESLTGVAVTWYTNLEP 585
Query: 62 GSISNFRDFSSKFLVQFSAN 81
+ +++D F+ Q+ N
Sbjct: 586 SRVHSWKDLMVAFVRQYQYN 645
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 42.7 bits (99), Expect = 3e-04
Identities = 45/213 (21%), Positives = 92/213 (43%), Gaps = 25/213 (11%)
Frame = +3
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRD 221
DEE + K+K K + G T +++KK ED G +++ K K K ++ + D
Sbjct: 126 DEEKKEKKKKEKKDKDGKTDAEKIKKK---EDDGKDDEEKKEKKKKYDKIDAXKVKGKED 296
Query: 222 D-------------------YEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVD 262
D E++ + +++E+ D TD +G +D +
Sbjct: 297 DGKDEGNKEKKDKEKGDGDGEEKKEKKDKEKEKKKEKKDKDEETDTLKE-KGKNDEG--E 467
Query: 263 EPEAPKYQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIR------AGYQGNRQGQW 316
+ E K + +D K+ + H+ D ++ +++ +R +G ++ +
Sbjct: 468 DDEGNKKKKKDKKEKE--KDHKKEKKDKEEGEKEDSKVEVSVRDIDIEEIKKEGEKEDKG 641
Query: 317 RNGGDQNKAHKREEERADTKGKKKQESAAIATK 349
++GG + K K++E++ D K KKK+ + TK
Sbjct: 642 KDGGKEVKEKKKKEDK-DKKEKKKKVTGKDKTK 737
Score = 32.3 bits (72), Expect = 0.44
Identities = 37/180 (20%), Positives = 70/180 (38%), Gaps = 2/180 (1%)
Frame = +3
Query: 176 EKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHR 235
E+ K+ KK + +DKGD KGK +D E + K +
Sbjct: 15 EEKKEKDKKEKKKKENKDKGDKTDVEKVKGK-------------ENDGEDDEEKKEKKKK 155
Query: 236 QREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRDANPKKWCEFHRSAGHDTDDCWT 295
++++ D G +DA + + E +D KK + +D D
Sbjct: 156 EKKDKD------------GKTDAEKIKKKED---DGKDDEEKK----EKKKKYDKIDAXK 278
Query: 296 LQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGK--KKQESAAIATKGADD 353
++ + D G + + + +G + K K+++E+ K K K +E+ + KG +D
Sbjct: 279 VKGKEDDGKDEGNKEKKDKEKGDGDGEEKKEKKDKEKEKKKEKKDKDEETDTLKEKGKND 458
Score = 28.5 bits (62), Expect = 6.3
Identities = 15/57 (26%), Positives = 28/57 (48%)
Frame = +3
Query: 154 ARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFK 210
++ + D + E K++ K +KG K VK+ + EDK ++++ GK K
Sbjct: 567 SKVEVSVRDIDIEEIKKEGEKEDKGKDGGKEVKEKKKKEDKDKKEKKKKVTGKDKTK 737
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 40.4 bits (93), Expect = 0.002
Identities = 29/96 (30%), Positives = 38/96 (39%), Gaps = 4/96 (4%)
Frame = +2
Query: 18 DPKDHLLYFN----TKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 73
DP HL F+ T D + + FP + + A W L SI N+ D
Sbjct: 575 DPHKHLKEFHIVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRSIFNWDDLKRV 754
Query: 74 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARY 109
FL +F D+ IRQ ESL EY R+
Sbjct: 755 FLEKFFPASRTTTIRKDISGIRQLSRESLYEYWERF 862
>TC218783 similar to UP|Q40363 (Q40363) NuM1 protein, partial (19%)
Length = 870
Score = 35.8 bits (81), Expect = 0.039
Identities = 33/105 (31%), Positives = 44/105 (41%), Gaps = 5/105 (4%)
Frame = +3
Query: 117 EDEEPRA-CALAFKNGLLPGGLNSKLTRK--PARSMGEMRARASTYILDEEDEAFKRKRA 173
EDE+P A A+ KN G S L +K PA S + +EDEA K K A
Sbjct: 99 EDEKPAAKVAVPSKNQSAKNGTLSTLAKKGKPAASSSSSESSDDN---SDEDEAPKTKVA 269
Query: 174 KL--EKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQL 216
+ G S K+ + S + +GKS KPT +L
Sbjct: 270 PAAGKNGHASTKKTQPSESSDSDSSDSDSSSDEGKSKKKPTTAKL 404
>TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g75390), partial
(34%)
Length = 1184
Score = 33.5 bits (75), Expect = 0.20
Identities = 27/98 (27%), Positives = 41/98 (41%)
Frame = +2
Query: 131 GLLPGGLNSKLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRS 190
G+ G S + + R T I DEE EA +R + E SP R ++ S
Sbjct: 440 GIFGGRRRSGCDGREKEEADGVEPRVRTAIADEEAEAARRPIRRSE----SPSRRERALS 607
Query: 191 GEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRP 228
E KG+G R + + + +P+ D+ RRP
Sbjct: 608 AEHKGEGGSVR*DRSR--------ERHPQSTDHGTRRP 697
>TC227935
Length = 681
Score = 32.3 bits (72), Expect = 0.44
Identities = 27/95 (28%), Positives = 39/95 (40%)
Frame = +2
Query: 144 KPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPG 203
+ R E RARA+ D D +R+ + GD + K D G G R G
Sbjct: 212 RSGREAREGRARAA----DNGDIR*RRRGSVRRDGDAGSEHKKLD-----SGKGSAPRDG 364
Query: 204 KGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHRQRE 238
S + T +PRR R P+Q+K ++E
Sbjct: 365 ADVSRSRGTLAGYHPRRGALPNRTPYQTKESGRQE 469
>AI461216 homologue to GP|13702121|dbj SA2112 {Staphylococcus aureus subsp.
aureus N315}, partial (3%)
Length = 410
Score = 32.0 bits (71), Expect = 0.57
Identities = 28/111 (25%), Positives = 53/111 (47%)
Frame = -2
Query: 160 ILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPR 219
I +E+D +K+KRAK E + + ++ K + D Q PT E +
Sbjct: 310 IENEDDGKYKQKRAK-ETEEATEMQLLKTATDSPPEDNVQST--------VPTSEIVKGE 158
Query: 220 RDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQ 270
++ Q++ +++ +ET +V++TD + + +G D D+ EA K Q
Sbjct: 157 TEEISQQQTYKN------DET-VVIDTDKTTLQQGIEDGGNFDQQEATKIQ 26
>TC203903 homologue to UP|Q94LX2 (Q94LX2) Beta-conglycinin alpha subunit,
partial (31%)
Length = 903
Score = 30.8 bits (68), Expect = 1.3
Identities = 24/99 (24%), Positives = 40/99 (40%)
Frame = +3
Query: 174 KLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKS 233
K+EK + + + R + + Q+PG+ K E PR + + +P Q +
Sbjct: 201 KVEKEECEEGEIPRPRPRPQHPEREPQQPGE-----KEEDEDEQPRPIPFPRPQPRQEEE 365
Query: 234 HRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPR 272
H QREE + + RG + DE E + R
Sbjct: 366 HEQREEQEWPRKEE----KRGEKGSEEEDEDEDEEQDER 470
>TC203614 UP|Q94LX2 (Q94LX2) Beta-conglycinin alpha subunit, complete
Length = 1983
Score = 30.8 bits (68), Expect = 1.3
Identities = 24/99 (24%), Positives = 40/99 (40%)
Frame = +3
Query: 174 KLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKS 233
K+EK + + + R + + Q+PG+ K E PR + + +P Q +
Sbjct: 222 KVEKEECEEGEIPRPRPRPQHPEREPQQPGE-----KEEDEDEQPRPIPFPRPQPRQEEE 386
Query: 234 HRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPR 272
H QREE + + RG + DE E + R
Sbjct: 387 HEQREEQEWPRKEE----KRGEKGSEEEDEDEDEEQDER 491
>TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(20%)
Length = 850
Score = 30.8 bits (68), Expect = 1.3
Identities = 21/85 (24%), Positives = 34/85 (39%), Gaps = 14/85 (16%)
Frame = +1
Query: 170 RKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYP----------- 218
RK+ K K +PK K+ R + Q+P K + KP + Q P
Sbjct: 250 RKKKKKTKNPKTPKAKKRKRKLRNNKRSGHQKP*KSNRIQKPWRLQTTPK*TAAMKETTK 429
Query: 219 ---RRDDYEQRRPWQSKSHRQREET 240
RR ++ PW+S +R + +
Sbjct: 430 KRKRRISLSKKSPWRSFWNRSQRSS 504
>CF920688
Length = 521
Score = 30.8 bits (68), Expect = 1.3
Identities = 16/56 (28%), Positives = 27/56 (47%)
Frame = +1
Query: 320 GDQNKAHKREEERADTKGKKKQESAAIATKGADDTFARHSGPPVGTINTIAGGFGG 375
G K K+++++ K KKK++ + K +R GPP G I+ + GG
Sbjct: 133 GPXKKKKKKKKKKKKKKKKKKKKKKKLKKKKKKKKSSRPGGPPKGAISANSLHMGG 300
>TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, complete
Length = 2152
Score = 30.4 bits (67), Expect = 1.7
Identities = 18/68 (26%), Positives = 31/68 (45%)
Frame = +3
Query: 174 KLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKS 233
K+EK + + + R + + Q+PG+ K E PR + + +P Q +
Sbjct: 261 KVEKEECEEGEIPRPRPRPQHPEREPQQPGE-----KEEDEDEQPRPIPFPRPQPRQEEE 425
Query: 234 HRQREETD 241
H QREE +
Sbjct: 426 HEQREEQE 449
>AW471626 similar to SP|P11827|GLCX Beta-conglycinin alpha' chain precursor.
[Soybean] {Glycine max}, partial (26%)
Length = 517
Score = 30.0 bits (66), Expect = 2.2
Identities = 25/90 (27%), Positives = 40/90 (43%), Gaps = 4/90 (4%)
Frame = +2
Query: 154 ARASTYILDEEDE----AFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVF 209
AR +DEE+E AF R R + P+R ++ +++ +G+Q RP
Sbjct: 164 ARCDLLKVDEEEECEKGAFPRPRPQ------HPEREREQHGEKEEDEGEQPRPFP----- 310
Query: 210 KPTKEQLYPRRDDYEQRRPWQSKSHRQREE 239
+PR R+P Q + H Q+EE
Sbjct: 311 -------FPR-----PRQPHQEEEHEQKEE 364
>CD487177 homologue to GP|10946375|gb| ribulose-1 5-bisphosphate carboxylase
small subunit rbcS1 {Glycine max}, partial (31%)
Length = 634
Score = 30.0 bits (66), Expect = 2.2
Identities = 16/35 (45%), Positives = 19/35 (53%)
Frame = -3
Query: 350 GADDTFARHSGPPVGTINTIAGGFGGGGDTHAARK 384
G DDT A G PVG I + GG G GG+ R+
Sbjct: 620 GDDDTGAARVGGPVGEIGELEGG-GFGGEVRRVRR 519
>TC223614 similar to UP|Q950Y2 (Q950Y2) Ymf62, partial (6%)
Length = 500
Score = 30.0 bits (66), Expect = 2.2
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = -1
Query: 315 QWRNGGDQNKAHKREEERADTKGKKKQESAAIATKGADDTFARHSG 360
+W+N ++N H + E +GKK+++ A TK T H G
Sbjct: 191 RWKNNSNKNPVHTLQIEMFLQQGKKQKDQADTDTKNNHGTKGWHRG 54
>BE023814
Length = 401
Score = 29.6 bits (65), Expect = 2.8
Identities = 21/95 (22%), Positives = 45/95 (47%), Gaps = 1/95 (1%)
Frame = +1
Query: 134 PGGLNSKLTRKPARSMG-EMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGE 192
PG L PA + ++ + D+ ++ +RKR + E+ + ++ +K S E
Sbjct: 13 PGSSKPSLGDTPAEEENVSLPSQPAMKTSDDSEDESRRKRRREERKEEKREKREKHHSRE 192
Query: 193 DKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRR 227
D+ K+++ +G S +++ + D E+RR
Sbjct: 193 DRKQEKREKHERGHSRDSDGRKR---HKKDKERRR 288
>TC204168 UP|Q7XXT2 (Q7XXT2) Prepro beta-conglycinin alpha prime subunit,
complete
Length = 2055
Score = 29.6 bits (65), Expect = 2.8
Identities = 34/180 (18%), Positives = 72/180 (39%), Gaps = 1/180 (0%)
Frame = +2
Query: 154 ARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTK 213
AR + ++EE+E + + + P+R ++ +++ +G+Q RP
Sbjct: 224 ARCNLLKVEEEEECEEGQIPRPRP--QHPERERQQHGEKEEDEGEQPRPFP--------- 370
Query: 214 EQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPK-YQPR 272
+PR R+P Q + H Q+EE + + G DE E P+ +QP
Sbjct: 371 ---FPR-----PRQPHQEEEHEQKEEHEWHRKEEKHG---GKGSEEEQDEREHPRPHQPH 517
Query: 273 DANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEER 332
+K H+ H + + + D+ + +++ Q G + + +R + +
Sbjct: 518 QKEEEKHEWQHKQEKHQGKESEEEEEDQDE----DEEQDKESQESEGSESQREPRRHKNK 685
>TC231328 GB|AAF34631.1|6979760|AF215712 protamine 2 {Gorilla gorilla;} ,
partial (12%)
Length = 541
Score = 29.3 bits (64), Expect = 3.7
Identities = 22/68 (32%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Frame = -1
Query: 169 KRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRP 228
+ +R L GD + +RV++ SGE QQR +P KE+ RR +RR
Sbjct: 175 RHRRLALTAGDKNRERVERGESGESFRRATQQRS-------RPRKER---RRSCRHRRRS 26
Query: 229 --WQSKSH 234
W+ K H
Sbjct: 25 CRWRKKVH 2
>TC216344 weakly similar to UP|Q9SMQ5 (Q9SMQ5) Proliferating-cell nucleolar
antigen-like protein, partial (12%)
Length = 726
Score = 29.3 bits (64), Expect = 3.7
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Frame = +2
Query: 148 SMGEMRARASTYILD---EEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGK 204
++G + RAS ++ E E +R +LEK + K+R ++KGD Q+ GK
Sbjct: 335 AIGCWKGRASLTVMVGALECQELLERLIMRLEKVTEKDSSMHKNRPSDNKGDEAQESNGK 514
Query: 205 GKSV 208
+ V
Sbjct: 515 NEDV 526
>TC211438 UP|GLCX_SOYBN (P11827) Beta-conglycinin, alpha' chain precursor,
complete
Length = 1959
Score = 28.1 bits (61), Expect = 8.2
Identities = 30/136 (22%), Positives = 55/136 (40%), Gaps = 1/136 (0%)
Frame = +1
Query: 154 ARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTK 213
AR + ++EE+E + + + P+R ++ +++ +G+Q RP
Sbjct: 202 ARCNLLKVEEEEECEEGQIPRPRP--QHPERERQQHGEKEEDEGEQPRPFP--------- 348
Query: 214 EQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPK-YQPR 272
+PR R+P Q + H Q+EE + + G DE E P+ +QP
Sbjct: 349 ---FPR-----PRQPHQEEEHEQKEEHEWHRKEEKHG---GKGSEEEQDEREHPRPHQPH 495
Query: 273 DANPKKWCEFHRSAGH 288
+K H+ H
Sbjct: 496 QKEEEKHEWQHKQEKH 543
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,730,270
Number of Sequences: 63676
Number of extensions: 188084
Number of successful extensions: 1335
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 1304
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1328
length of query: 441
length of database: 12,639,632
effective HSP length: 100
effective length of query: 341
effective length of database: 6,272,032
effective search space: 2138762912
effective search space used: 2138762912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0590c.5


