

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.10
(993 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
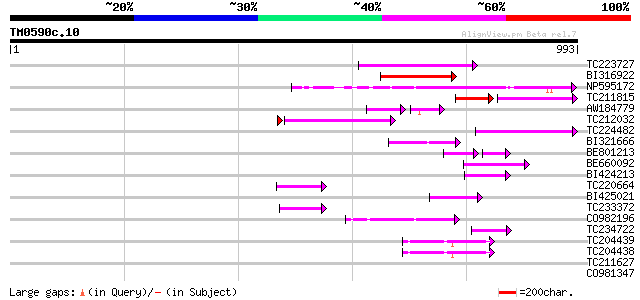
Score E
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 183 4e-46
BI316922 135 9e-32
NP595172 polyprotein [Glycine max] 127 2e-29
TC211815 78 5e-28
AW184779 64 2e-19
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 92 7e-19
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 86 8e-17
BI321666 82 2e-15
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 52 5e-11
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 62 1e-09
BI424213 57 4e-08
TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 57 5e-08
BI425021 54 3e-07
TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 52 1e-06
CO982196 50 6e-06
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 49 8e-06
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 49 1e-05
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 49 1e-05
TC211627 42 0.001
CO981347 41 0.003
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 183 bits (464), Expect = 4e-46
Identities = 87/209 (41%), Positives = 125/209 (59%), Gaps = 1/209 (0%)
Frame = +1
Query: 612 QREASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRA 671
+R A+ + + LY+R L+CV + ++ EVHEG +H G ++A K+LRA
Sbjct: 217 RRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTHANGHAMARKILRA 396
Query: 672 GFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPF-PTAR 730
G+YW T+ DC V++C +CQ FAD APP L MS+PWPF+MWG+D++G P A
Sbjct: 397 GYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMWGIDVIGAIEPKAS 576
Query: 731 AQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQ 790
+FILVA+DYFTKW+EA + +V F K I+CR+G+PR I++DNGT ++
Sbjct: 577 NGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRKIITDNGTNLNNKM 756
Query: 791 TREFCKEMGIQMRFASVEHPQTNGQVESA 819
E C+E IQ + P+ N VE A
Sbjct: 757 MGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>BI316922
Length = 405
Score = 135 bits (340), Expect = 9e-32
Identities = 61/134 (45%), Positives = 91/134 (67%)
Frame = +3
Query: 649 EVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVT 708
E+H G+C H + + +VLR G+Y T+RK C ++VK+C+EC F ++S +EL
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 709 MSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRI 768
+ APWPFA+ GVD++ PFP ++ Q+K++LV +D FTKWIE E +A I+ A + F + I
Sbjct: 183 IVAPWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNI 362
Query: 769 VCRFGIPRAIVSDN 782
VC FGIP ++SDN
Sbjct: 363 VC*FGIPNTLISDN 404
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 127 bits (319), Expect = 2e-29
Identities = 134/515 (26%), Positives = 223/515 (43%), Gaps = 16/515 (3%)
Frame = +1
Query: 494 LIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKL 553
+IRTD + +++ + + Q P +L + L + +EY P +NQ ADAL+++
Sbjct: 2863 IIRTDQRSLKSLMDQSLQT--PEQQAWLHKF-----LGYDFKIEYKPGKDNQAADALSRM 3021
Query: 554 ASTRKPGNNKSVIQETLAYPSIEGELMACVSRGRTWMDPIISILEGDPAEVEQCTKEQQR 613
+ ++E R R DP + L T +Q
Sbjct: 3022 FMLAWSEPHSIFLEEL---------------RARLISDPHLKQLME--------TYKQGA 3132
Query: 614 EASHYTLIDGHLYRRGFSTPLLKCVPPEKYEA---IMSEVHEGVCASHIG-GRSLACKVL 669
+ASHYT+ +G LY + + V P + E I+ E H H G R+LA L
Sbjct: 3133 DASHYTVREGLLYWKD------RVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLAR--L 3288
Query: 670 RAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW---GVDLVGPF 726
+A FYWP +++D ++++C CQ + P L + P P +W +D +
Sbjct: 3289 KAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL--PIPQQVWEDVAMDFITGL 3462
Query: 727 PTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQ 785
P + + I+V +D TK+ PL ++K+V + IV GIPR+IVSD
Sbjct: 3463 PNSFG-LSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRV 3639
Query: 786 FSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAV 845
F+S+ + K G + +S HPQ++GQ E N+ + LR E W+ LP
Sbjct: 3640 FTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWA 3819
Query: 846 LWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSET 905
+ YNT + TPFR YG + P + TR ++ A + +L
Sbjct: 3820 EFWYNTAYHMSLGMTPFRALYGRE---PPTL------TRQACSIDDPAEVREQLTDRDAL 3972
Query: 906 RDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVL----KWRSGA----PGNKLTPNWEG 957
+ I T +Q + + + + Q+GD VL +R + KL+ + G
Sbjct: 3973 LAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFG 4152
Query: 958 PYRIIKVLGNGAYHLEELDGRRLPRSFNGLSLRYF 992
P++++ +G+ AY LE R+ F+ L+ F
Sbjct: 4153 PFKVLAKIGDVAYKLELPSAARIHPVFHVSQLKPF 4257
>TC211815
Length = 704
Score = 77.8 bits (190), Expect(2) = 5e-28
Identities = 49/143 (34%), Positives = 76/143 (52%), Gaps = 3/143 (2%)
Frame = +2
Query: 854 QSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRE 913
QSTT ETPF + YG+ MLP+E+ E N+ + ++LDL+ + R++ I
Sbjct: 230 QSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIMT 409
Query: 914 TAMKQRVAAKFNSRVRVREMQVG---DLVLKWRSGAPGNKLTPNWEGPYRIIKVLGNGAY 970
A KQR+ FNS++ G + + + +K T N EGP++I NGAY
Sbjct: 410 *AFKQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVRSKFTTN*EGPFKIRHNSKNGAY 589
Query: 971 HLEELDGRRLPRSFNGLSLRYFY 993
LEEL G+ + R +N + L+ +Y
Sbjct: 590 KLEELSGKVVLRIWNSMHLKVYY 658
Score = 66.2 bits (160), Expect(2) = 5e-28
Identities = 30/67 (44%), Positives = 45/67 (66%)
Frame = +3
Query: 781 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 840
DNG QF++ + EF + I+ R SV+HPQTN + E+AN+VIL L++ L A G W++
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLLDGANGRWVE 182
Query: 841 ELPAVLW 847
+L +LW
Sbjct: 183 DLVEILW 203
>AW184779
Length = 432
Score = 63.9 bits (154), Expect(2) = 2e-19
Identities = 28/69 (40%), Positives = 41/69 (58%)
Frame = +1
Query: 625 LYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMD 684
LY+R LL+CV + E ++ EVHEG +H ++A K+LR G+YW T+ DC
Sbjct: 16 LYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDCCI 195
Query: 685 FVKQCKECQ 693
V +C +CQ
Sbjct: 196 HVWKCHKCQ 222
Score = 50.8 bits (120), Expect(2) = 2e-19
Identities = 26/70 (37%), Positives = 38/70 (54%), Gaps = 10/70 (14%)
Frame = +3
Query: 702 PPKELVTMSAPWPFA---------MWGVDLVGPF-PTARAQMKFILVAVDYFTKWIEAEP 751
P + + M P P+ MWG+D++G P A FILVA+DYFTKW+EA
Sbjct: 219 PVRPSLIMLTPHPYL*MSWQHLGHMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVS 398
Query: 752 LAKITSAKIV 761
A +T + ++
Sbjct: 399 YASVTRSVVI 428
Score = 30.0 bits (66), Expect = 5.2
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +2
Query: 693 QVFADLSKAPPKELVTMSAPWPF 715
Q FAD APP L ++APWP+
Sbjct: 224 QTFADNVNAPPIPLNVLAAPWPY 292
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 92.4 bits (228), Expect(2) = 7e-19
Identities = 61/199 (30%), Positives = 100/199 (49%), Gaps = 5/199 (2%)
Frame = +2
Query: 482 LKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPR 541
++ A + ++ L + DS LV +Q++G + +DPNLI Y ++ L F E+ +V
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 542 AENQRADALAKLASTRK--PGNNKSVIQETLAYPSIEGELMACVSRGRTWMDPIISILEG 599
ENQ ADALA L S + P + I+ L+ G+ W I +E
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLVEEERDGKPWYFDIKRYVES 565
Query: 600 DPAEVEQCTKEQQRE---ASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCA 656
+E +++R+ A+ + + LY+R LL CV ++ E ++ EVHEG
Sbjct: 566 KEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFG 745
Query: 657 SHIGGRSLACKVLRAGFYW 675
+H G ++A K+LRAG+YW
Sbjct: 746 THSNGHAMARKILRAGYYW 802
Score = 20.8 bits (42), Expect(2) = 7e-19
Identities = 8/9 (88%), Positives = 8/9 (88%)
Frame = +1
Query: 469 TNNQAEYEA 477
TNN AEYEA
Sbjct: 166 TNNMAEYEA 192
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 85.9 bits (211), Expect = 8e-17
Identities = 57/183 (31%), Positives = 94/183 (51%), Gaps = 6/183 (3%)
Frame = +1
Query: 817 ESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLP--V 874
E+AN+ I + +++ K W + LP L Y T+ +++T TPF + YG++A+LP V
Sbjct: 1 EAANKNIKKIIQKMTVSYKD-WHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 875 EIDNFTWRTRPGFEEENQANMAV-ELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVREM 933
E+ + G +E A +L+L+ R A +QR+ + F+ +V +R+
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 934 QVGDLVLKWRSGAPGN---KLTPNWEGPYRIIKVLGNGAYHLEELDGRRLPRSFNGLSLR 990
GDLVLK S A + K PN+EGP+ + + GA L +DG LP N ++
Sbjct: 358 HEGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVVK 537
Query: 991 YFY 993
+Y
Sbjct: 538 RYY 546
>BI321666
Length = 430
Score = 81.6 bits (200), Expect = 2e-15
Identities = 41/126 (32%), Positives = 69/126 (54%)
Frame = +2
Query: 664 LACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLV 723
++ VL++ FY P++ KD + C +CQ +SK L T+ F WG+D V
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFV 181
Query: 724 GPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNG 783
GPFP + + ++ILV VDY +KW+EA K + ++ F K+I R G+P ++ + G
Sbjct: 182 GPFPPSFSN-EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGG 358
Query: 784 TQFSSS 789
+ ++
Sbjct: 359 SHLCNA 376
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 52.4 bits (124), Expect(2) = 5e-11
Identities = 23/61 (37%), Positives = 36/61 (58%)
Frame = +3
Query: 760 IVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESA 819
++ F K I RFG+PR ++SD G+ F SQ + K ++ + + HPQTNGQ + +
Sbjct: 18 VIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNGQAKVS 197
Query: 820 N 820
N
Sbjct: 198 N 200
Score = 34.3 bits (77), Expect(2) = 5e-11
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = +2
Query: 829 RRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEI 876
+ +A ++ W +L LW+ T +++ TPF+M Y LPVE+
Sbjct: 227 KNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVEL 370
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 62.0 bits (149), Expect = 1e-09
Identities = 39/118 (33%), Positives = 62/118 (52%), Gaps = 3/118 (2%)
Frame = -3
Query: 796 KEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQS 855
K+ G+ R ++ HPQTNGQ E +NR I R L + + ++ W L LW++ T ++
Sbjct: 361 KKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAYKA 182
Query: 856 TTRETPFRMTYGVDAMLPVEIDNFT-WRTRPGFEEENQANMAVELDL--LSETRDEAH 910
+P+R+ +G LPVEI++ W + +QA +L L L E R EA+
Sbjct: 181 PIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRLEAY 8
>BI424213
Length = 426
Score = 57.0 bits (136), Expect = 4e-08
Identities = 26/80 (32%), Positives = 48/80 (59%)
Frame = +1
Query: 797 EMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQST 856
++G ++ F++ HPQT+GQ + NR + LR L +W + LP V ++YN T
Sbjct: 31 KLGTKLLFSTTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEFAYNRGVHRT 210
Query: 857 TRETPFRMTYGVDAMLPVEI 876
T+++PF + YG + + P+++
Sbjct: 211 TKQSPFEVVYGFNPLTPLDL 270
>TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (31%)
Length = 724
Score = 56.6 bits (135), Expect = 5e-08
Identities = 34/87 (39%), Positives = 48/87 (55%)
Frame = +1
Query: 468 ATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYL 527
ATNN AEY A+I G+K A + + I+ DS+LV Q+ G+++VK+ NL + L
Sbjct: 61 ATNNAAEYRAMILGMKYALKKGFTGIRIQGDSKLVCMQIDGSWKVKNENLSTLYNVAKEL 240
Query: 528 MTLFQEVVVEYVPRAENQRADALAKLA 554
F + +V R N ADA A LA
Sbjct: 241 KDKFSSFQISHVLRNFNSDADAQANLA 321
>BI425021
Length = 426
Score = 53.9 bits (128), Expect = 3e-07
Identities = 36/94 (38%), Positives = 50/94 (52%), Gaps = 1/94 (1%)
Frame = -1
Query: 736 ILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRF-GIPRAIVSDNGTQFSSSQTREF 794
I V V+ F+K I L +A +V + IV + G PR+IVSD F S ++
Sbjct: 336 IFVVVNRFSKGIHLGTLPTSHTAHMVASLFLNIVIKLHGFPRSIVSDRDPLFISHFWQDL 157
Query: 795 CKEMGIQMRFASVEHPQTNGQVESANRVILRGLR 828
+ G +R +S HPQT+GQ E NRVI + LR
Sbjct: 156 FRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQYLR 55
>TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (12%)
Length = 561
Score = 52.0 bits (123), Expect = 1e-06
Identities = 27/82 (32%), Positives = 45/82 (53%)
Frame = +1
Query: 473 AEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQ 532
AEY +LI GL + + ++++ DS LV NQ++G +++K+ N+ + L F
Sbjct: 1 AEYRSLILGLXHXXKKGYKXIIVQGDSLLVCNQIQGLWKIKNQNMGTLCXEAKELKDKFL 180
Query: 533 EVVVEYVPRAENQRADALAKLA 554
+ ++PR N ADA A LA
Sbjct: 181 SFKISHIPREYNSEADAQANLA 246
>CO982196
Length = 812
Score = 49.7 bits (117), Expect = 6e-06
Identities = 53/204 (25%), Positives = 85/204 (40%), Gaps = 4/204 (1%)
Frame = +1
Query: 589 WMDPIISILEGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMS 648
W + I + LE E+ Q + + Y + G LY F L+ K ++
Sbjct: 220 WEEEIQAYLE--LYEIYQGILTKTTKKPGYAIRGGKLY---FKDRLVLSKNSTKIPLLLK 384
Query: 649 EVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVT 708
E+ + H G +V F W ++K D+V C+ C+ + +P L
Sbjct: 385 ELQDSPLGGHSGFFRTFKRVANVVF-WQGMKKTTRDYVAACEICRRNKTSTLSPAGLL*L 561
Query: 709 MSAPWPFAMW---GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAK-IVNFY 764
+ P P +W +D +G P A+ + ILV VD TK+ L+ +AK + +
Sbjct: 562 L--PIPTKVWTDISMDFIGGLPKAQGKDN-ILVVVDRLTKYAHFFALSHPYTAKEVAELF 732
Query: 765 WKRIVCRFGIPRAIVSDNGTQFSS 788
K +V G P +IVSD F S
Sbjct: 733 IKELVRLHGFPASIVSDXXRLFMS 804
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 49.3 bits (116), Expect = 8e-06
Identities = 23/70 (32%), Positives = 38/70 (53%)
Frame = -3
Query: 809 HPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGV 868
HPQTNGQ E +N+ I R L + ++ W +L W+Y ++ +PF++ YG
Sbjct: 231 HPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYGK 52
Query: 869 DAMLPVEIDN 878
L VE+++
Sbjct: 51 ACHLSVELEH 22
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 48.9 bits (115), Expect = 1e-05
Identities = 46/166 (27%), Positives = 76/166 (45%), Gaps = 6/166 (3%)
Frame = +1
Query: 689 CKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPT-ARAQMKFILVAVDYFTKWI 747
C ECQ+ + K ++L + + +DL+GP + ++ V VD F+++
Sbjct: 2179 CGECQIGKQV-KMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFT 2355
Query: 748 EAEPLAKITSAKIVNFYWKRIVCRFG-----IPRAIVSDNGTQFSSSQTREFCKEMGIQM 802
+ + + V +K + R + + I SD+G +F +S+ EFC GI
Sbjct: 2356 WVNFIREKSETFEV---FKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITH 2526
Query: 803 RFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWS 848
F++ PQ NG VE NR L+ R + AK ELP LW+
Sbjct: 2527 EFSAAITPQQNGIVERKNRT-LQEAARVMLHAK-----ELPYNLWA 2646
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 48.9 bits (115), Expect = 1e-05
Identities = 46/166 (27%), Positives = 76/166 (45%), Gaps = 6/166 (3%)
Frame = +1
Query: 689 CKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPT-ARAQMKFILVAVDYFTKWI 747
C ECQ+ + K ++L + + +DL+GP + ++ V VD F+++
Sbjct: 2182 CGECQIGKQV-KMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFT 2358
Query: 748 EAEPLAKITSAKIVNFYWKRIVCRFG-----IPRAIVSDNGTQFSSSQTREFCKEMGIQM 802
+ + + V +K + R + + I SD+G +F +S+ EFC GI
Sbjct: 2359 WVNFIREKSDTFEV---FKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITH 2529
Query: 803 RFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWS 848
F++ PQ NG VE NR L+ R + AK ELP LW+
Sbjct: 2530 EFSAAITPQQNGIVERKNRT-LQEAARVMLHAK-----ELPYNLWA 2649
>TC211627
Length = 1034
Score = 42.4 bits (98), Expect = 0.001
Identities = 25/77 (32%), Positives = 37/77 (47%)
Frame = +2
Query: 786 FSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAV 845
F S E G ++RF++ HPQT+GQ E NR++ + LR + + W L
Sbjct: 5 FISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSLA 184
Query: 846 LWSYNTTEQSTTRETPF 862
YNT+ S +PF
Sbjct: 185 E*CYNTSVHSGIGFSPF 235
>CO981347
Length = 624
Score = 40.8 bits (94), Expect = 0.003
Identities = 27/85 (31%), Positives = 38/85 (43%), Gaps = 2/85 (2%)
Frame = +2
Query: 776 RAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAK 835
+ + +DNG +F Q EFC+++GI+ P NG E N IL +R L A+
Sbjct: 164 KVLRTDNGLEFVLEQFNEFCRKIGIKRHKIVPHTP*QNGLAERMNMTILERVRCMLLSAR 343
Query: 836 GAWLDELPAVLW--SYNTTEQSTTR 858
LP W + NTT R
Sbjct: 344 ------LPKTFWGEAANTTSYLINR 400
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.336 0.143 0.468
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,205,559
Number of Sequences: 63676
Number of extensions: 687227
Number of successful extensions: 6650
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 6200
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6566
length of query: 993
length of database: 12,639,632
effective HSP length: 107
effective length of query: 886
effective length of database: 5,826,300
effective search space: 5162101800
effective search space used: 5162101800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0590c.10


