

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590b.4
(283 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
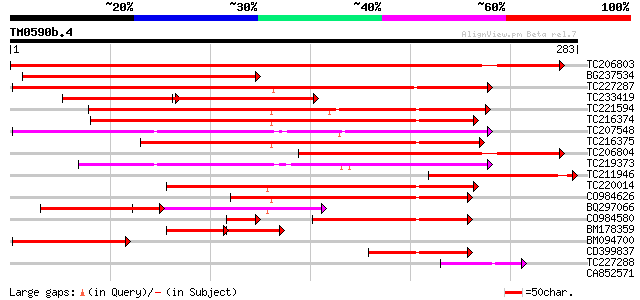
Score E
Sequences producing significant alignments: (bits) Value
TC206803 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gat... 342 9e-95
BG237534 similar to PIR|T52574|T52 cyclic nucleotide and calmodu... 227 4e-60
TC227287 similar to UP|CG17_ARATH (Q8L7Z0) Probable cyclic nucle... 225 2e-59
TC233419 homologue to UP|CNG4_ARATH (Q94AS9) Cyclic nucleotide-g... 131 1e-57
TC221594 similar to UP|Q9M7Q8 (Q9M7Q8) Cyclic nucleotide-gated c... 192 1e-49
TC216374 similar to UP|CNG6_ARATH (O82226) Probable cyclic nucle... 173 7e-44
TC207548 similar to UP|CG20_ARATH (Q9LD37) Probable cyclic nucle... 155 1e-38
TC216375 similar to UP|CNG6_ARATH (O82226) Probable cyclic nucle... 151 4e-37
TC206804 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gat... 142 1e-34
TC219373 similar to UP|CG20_ARATH (Q9LD37) Probable cyclic nucle... 135 2e-32
TC211946 similar to UP|CNG4_ARATH (Q94AS9) Cyclic nucleotide-gat... 130 5e-31
TC220014 homologue to UP|Q8H6U1 (Q8H6U1) Cyclic nucleotide-gated... 130 7e-31
CO984626 100 1e-21
BQ297066 homologue to GP|24943196|gb cyclic nucleotide-gated cha... 81 6e-16
CO984580 72 4e-15
BM178359 similar to PIR|T51519|T515 cyclic nucleotide-gated cati... 50 4e-12
BM094700 similar to GP|8131898|gb| cyclic nucleotide-binding tra... 55 5e-08
CD399837 similar to GP|24943192|gb cyclic nucleotide-gated chann... 50 9e-07
TC227288 similar to UP|CG17_ARATH (Q8L7Z0) Probable cyclic nucle... 43 1e-04
CA852571 similar to GP|8131898|gb|A cyclic nucleotide-binding tr... 39 0.003
>TC206803 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gated ion
channel 2 (AtCNGC2) (Cyclic nucleotide-and
calmodulin-regulated ion channel 2) (DEFENSE NO DEATH
1), partial (37%)
Length = 1018
Score = 342 bits (878), Expect = 9e-95
Identities = 168/277 (60%), Positives = 204/277 (72%)
Frame = +2
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL A +KK+ MQLR R++EWWMR+RQLP LRQRVR++ERQRW AM G DE +MI++L
Sbjct: 44 VFLHAVMAKKRKMQLRCRDMEWWMRRRQLPSRLRQRVRHFERQRWAAMGGEDEMEMIKDL 223
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIK HLCLDL+R+VPLF +MDDL+L+NICDRVK LVF+K E I REGDPV RM+
Sbjct: 224 PEGLRRDIKRHLCLDLIRKVPLFHNMDDLILDNICDRVKPLVFSKDEKIIREGDPVPRMV 403
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
FVVRG ++ +Q L G+ + L PG F GDELLSWCLRRPFI+RLP SSAT V LE++E
Sbjct: 404 FVVRGRIKRNQSLSKGMVASSILEPGGFLGDELLSWCLRRPFIDRLPASSATFVCLESSE 583
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AFGL+A +RY+T HFRY F E++KR+ARYYS WRTWAAV IQ AWRRY+ R
Sbjct: 584 AFGLDANHLRYITDHFRYKFANERLKRTARYYSSNWRTWAAVNIQFAWRRYRQRTK---- 751
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEE 277
P P E RL Y A+ S +P+D E
Sbjct: 752 ---GPVTPVRDTNGGTERRLLQYAAMFMSIRPHDHLE 853
>BG237534 similar to PIR|T52574|T52 cyclic nucleotide and
calmodulin-regulated ion channel [imported] -
Arabidopsis thaliana, partial (17%)
Length = 359
Score = 227 bits (579), Expect = 4e-60
Identities = 107/119 (89%), Positives = 113/119 (94%)
Frame = +3
Query: 7 TSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRR 66
TSKKQAM LRMRN+EWWM KR+LPQG RQRVRNYER RW A RGVDECQMI+NLP+GLRR
Sbjct: 3 TSKKQAMLLRMRNIEWWMSKRRLPQGFRQRVRNYERMRWAATRGVDECQMIKNLPEGLRR 182
Query: 67 DIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRG 125
DIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETIT+EGDPV+RMLFVVRG
Sbjct: 183 DIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITKEGDPVQRMLFVVRG 359
>TC227287 similar to UP|CG17_ARATH (Q8L7Z0) Probable cyclic nucleotide-gated
ion channel 17 (Cyclic nucleotide-and
calmodulin-regulated ion channel 17), partial (43%)
Length = 1743
Score = 225 bits (574), Expect = 2e-59
Identities = 118/242 (48%), Positives = 156/242 (63%), Gaps = 2/242 (0%)
Frame = +1
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + +L+ R+ E WMR RQLP+ LR RVR + + +W A RGVDE ++R LP
Sbjct: 310 YLQSITIRLEEWRLKQRDTEEWMRHRQLPEDLRSRVRRFVQYKWLATRGVDEETILRALP 489
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDI+ HLCLDLVR+VP F MDD +L+ IC+R+ S + T+G I REGDPV MLF
Sbjct: 490 ADLRRDIQRHLCLDLVRRVPFFSQMDDQLLDAICERLVSSLSTQGTYIVREGDPVTEMLF 669
Query: 122 VVRGHLQSS--QDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L SS R G + L PG+F G+ELLSW L LP S+ T+ L
Sbjct: 670 IIRGRLDSSTTNGGRSGFFNSIILRPGDFCGEELLSWALLPKSTINLPSSTRTVKALSEV 849
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L AED+++V FR K K++ + R+YS WRTWAA IQ AWRRYK R+++
Sbjct: 850 EAFALRAEDLKFVANQFRRLHSK-KLQHTFRFYSHHWRTWAACFIQAAWRRYKKRMTMKD 1026
Query: 240 LS 241
LS
Sbjct: 1027LS 1032
>TC233419 homologue to UP|CNG4_ARATH (Q94AS9) Cyclic nucleotide-gated ion
channel 4 (AtCNGC4) (Cyclic nucleotide-and
calmodulin-regulated ion channel 4) (AtHLM1), partial
(18%)
Length = 394
Score = 131 bits (329), Expect(2) = 1e-57
Identities = 64/73 (87%), Positives = 68/73 (92%)
Frame = +1
Query: 82 LFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYC 141
L+ +MDDLVLENICDRVKSL+FTKGETI REGDPV+RMLFVVRGHLQSSQ LRDGVKS C
Sbjct: 175 LYFNMDDLVLENICDRVKSLIFTKGETIAREGDPVQRMLFVVRGHLQSSQVLRDGVKSCC 354
Query: 142 TLGPGNFSGDELL 154
LGPGNFSGDELL
Sbjct: 355 MLGPGNFSGDELL 393
Score = 110 bits (274), Expect(2) = 1e-57
Identities = 50/59 (84%), Positives = 54/59 (90%)
Frame = +3
Query: 27 RQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQVPLFQH 85
R+LP G RQRVRNYERQRW AMRGVDE +M +NLP+GLRRDIKYHLCLDLVRQVPLFQH
Sbjct: 12 RRLPLGFRQRVRNYERQRWAAMRGVDEFEMTKNLPEGLRRDIKYHLCLDLVRQVPLFQH 188
>TC221594 similar to UP|Q9M7Q8 (Q9M7Q8) Cyclic nucleotide-gated
calmodulin-binding ion channel, partial (29%)
Length = 881
Score = 192 bits (488), Expect = 1e-49
Identities = 100/204 (49%), Positives = 135/204 (66%), Gaps = 3/204 (1%)
Frame = +2
Query: 40 YERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVK 99
Y++ +W A RGVDE +++ LP LRRDIK HLCLDLVR VPLF MD+ +L+ IC+R+K
Sbjct: 5 YDQYKWLATRGVDEEALLKGLPVDLRRDIKRHLCLDLVRGVPLFDQMDERMLDAICERLK 184
Query: 100 SLVFTKGETITREGDPVKRMLFVVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWC 157
+ T+G + REGDPV MLF++RGHL S + R G + C +GPG+F G+ELL+W
Sbjct: 185 PALCTEGTFLVREGDPVNEMLFIIRGHLDSYTTNGGRAGFFNSCRIGPGDFCGEELLTWA 364
Query: 158 L-RRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGW 216
L RP + LP S+ T+ ++ EAF L AED+++V FR K+ V+ R+YS W
Sbjct: 365 LDPRPSV-ILPSSTRTVKSISEVEAFALIAEDLKFVASQFRRLHSKQ-VRHKFRFYSHHW 538
Query: 217 RTWAAVAIQLAWRRYKHRLSLTSL 240
RTWAA IQ AWRR+K R + L
Sbjct: 539 RTWAACFIQAAWRRHKKRKQVAEL 610
>TC216374 similar to UP|CNG6_ARATH (O82226) Probable cyclic nucleotide-gated
ion channel 6 (AtCNGC6) (Cyclic nucleotide- and
calmodulin-regulated ion channel 6), partial (33%)
Length = 1063
Score = 173 bits (439), Expect = 7e-44
Identities = 89/196 (45%), Positives = 127/196 (64%), Gaps = 2/196 (1%)
Frame = +1
Query: 41 ERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKS 100
++ +W A RGVDE ++++LP+ LRRDIK HLCL LVR+VPLF+ MD+ +L+ IC+R+K
Sbjct: 1 DQYKWLATRGVDEENLVQSLPKDLRRDIKRHLCLALVRRVPLFESMDERLLDAICERLKP 180
Query: 101 LVFTKGETITREGDPVKRMLFVVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCL 158
+FT+ I REGDPV MLF++RG L+S + R G + L +F G+ELL+W L
Sbjct: 181 CLFTENTYIVREGDPVDEMLFIIRGRLESVTTDGGRSGFFNRGFLKEADFCGEELLTWAL 360
Query: 159 RRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRT 218
LP S+ T+ L EAF L A+++++V FR ++ V+ + R+YS WRT
Sbjct: 361 DPKSGSNLPSSTRTVKALMEVEAFALTADELKFVASQFRRLHSRQ-VQHTFRFYSQQWRT 537
Query: 219 WAAVAIQLAWRRYKHR 234
WAA IQ AWRRY +
Sbjct: 538 WAACFIQAAWRRYSKK 585
>TC207548 similar to UP|CG20_ARATH (Q9LD37) Probable cyclic nucleotide-gated
ion channel 20, chloroplast precursor (Cyclic
nucleotide-binding transporter 1), partial (40%)
Length = 1315
Score = 155 bits (393), Expect = 1e-38
Identities = 98/251 (39%), Positives = 143/251 (56%), Gaps = 11/251 (4%)
Frame = +1
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
FL A ++ MQLR R++E WM R+LP+ LR+RVR+ ER W A RGV+E ++ N+
Sbjct: 421 FLQALGRRRLEMQLRGRDVEQWMSHRRLPEDLRRRVRHAERYSWAATRGVNEEILLENMQ 600
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ L+ DI+ HL V++V +F MD+ +L+ IC+R+K + KG + +G V++M+F
Sbjct: 601 EDLQTDIRRHL-FKFVKKVRIFALMDEPILDAICERLKQKTYIKGSKVLSQGSLVEKMVF 777
Query: 122 VVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFI-----------ERLPPSS 170
VVRG L+S D DG + L G+ G+ELL+W L + +RL S+
Sbjct: 778 VVRGTLESFGD--DG--TMVPLSEGDACGEELLTWYLEHSSVSTDGKKVRVQGQRL-LSN 942
Query: 171 ATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRR 230
T+ L EAF L A D+ +T F V+ + RY SP WR+ AA IQ+AWR
Sbjct: 943 RTVRCLTNVEAFSLRAADLEELTILFTRFLRNPHVQGALRYVSPYWRSLAANRIQVAWRY 1122
Query: 231 YKHRLSLTSLS 241
K RLS + S
Sbjct: 1123RKKRLSRANTS 1155
>TC216375 similar to UP|CNG6_ARATH (O82226) Probable cyclic nucleotide-gated
ion channel 6 (AtCNGC6) (Cyclic nucleotide- and
calmodulin-regulated ion channel 6), partial (31%)
Length = 896
Score = 151 bits (381), Expect = 4e-37
Identities = 80/174 (45%), Positives = 110/174 (62%), Gaps = 2/174 (1%)
Frame = +2
Query: 66 RDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRG 125
RDIK HLCL LVR+VPLF+ MD+ +L+ IC+R+K +FT+ I REGDPV MLF++RG
Sbjct: 2 RDIKRHLCLALVRRVPLFESMDERLLDAICERLKPCLFTESTYIVREGDPVDEMLFIIRG 181
Query: 126 HLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFG 183
L+S + R G + L +F G+ELL+W L LP S+ T+ L EAF
Sbjct: 182 RLESVTTDGGRSGFFNRGFLKEADFCGEELLTWALDPKSGSNLPSSTRTVKALTEVEAFA 361
Query: 184 LEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSL 237
L AE++++V FR ++ V+ + R+YS WRTWAA IQ AWRRY R ++
Sbjct: 362 LTAEELKFVASQFRRLHSRQ-VQHTFRFYSQQWRTWAACFIQAAWRRYSKRKTM 520
>TC206804 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gated ion
channel 2 (AtCNGC2) (Cyclic nucleotide-and
calmodulin-regulated ion channel 2) (DEFENSE NO DEATH
1), partial (16%)
Length = 701
Score = 142 bits (359), Expect = 1e-34
Identities = 73/133 (54%), Positives = 85/133 (63%)
Frame = +1
Query: 145 PGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEK 204
PG F GDELLSWCLRRPFI+RLP SSAT V LE++ AFGL+A ++RY+T HFRY F E+
Sbjct: 1 PGGFLGDELLSWCLRRPFIDRLPASSATFVCLESS*AFGLDANNLRYITDHFRYKFANER 180
Query: 205 VKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLSFIRPRRPASRCTSMEEDRLRLYT 264
+KR+ARYYS WRTWAAV IQ AWRRY+ P P E L Y
Sbjct: 181 LKRTARYYSSNWRTWAAVNIQFAWRRYRQMTK-------GPVIPVRDTNGGTERMLLQYA 339
Query: 265 ALLTSPKPNDQEE 277
AL S P+D E
Sbjct: 340 ALFMSLMPHDHLE 378
>TC219373 similar to UP|CG20_ARATH (Q9LD37) Probable cyclic nucleotide-gated
ion channel 20, chloroplast precursor (Cyclic
nucleotide-binding transporter 1), partial (27%)
Length = 1151
Score = 135 bits (340), Expect = 2e-32
Identities = 85/217 (39%), Positives = 122/217 (56%), Gaps = 10/217 (4%)
Frame = +2
Query: 35 QRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENI 94
+RVR ER W A RGV+E ++ NLP+ L+RDI+ HL V+++ LF MD+ +L+ I
Sbjct: 8 RRVRRAERYNWAATRGVNEEMLMENLPEDLQRDIRRHL-FKFVKKIRLFALMDEPILDAI 184
Query: 95 CDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELL 154
C+R++ + KG I +G V++M+FVVRG L+S + DG + L G+ G+ELL
Sbjct: 185 CERLRQKTYIKGSKILSQGGLVEKMVFVVRGKLESIGE--DGTR--IPLSEGDSCGEELL 352
Query: 155 SWCLRRPFIE------RLPP----SSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEK 204
+W L + RLP S+ T+ L E+F L A D+ VT F
Sbjct: 353 TWYLEHSSVSTDGRKVRLPGQRLVSNRTVRCLTNVESFSLSASDIEEVTILFTRFLRSPC 532
Query: 205 VKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLS 241
V+ + RY SP WR+ AA IQ+AWR K RLS + S
Sbjct: 533 VQGALRYESPYWRSLAATRIQVAWRYRKKRLSRVNSS 643
>TC211946 similar to UP|CNG4_ARATH (Q94AS9) Cyclic nucleotide-gated ion
channel 4 (AtCNGC4) (Cyclic nucleotide-and
calmodulin-regulated ion channel 4) (AtHLM1), partial
(10%)
Length = 357
Score = 130 bits (328), Expect = 5e-31
Identities = 63/74 (85%), Positives = 66/74 (89%)
Frame = +3
Query: 210 RYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLSFIRPRRPASRCTSMEEDRLRLYTALLTS 269
RYYSP WRTWAAVAIQLAWRRY+HRL+LTSLSFIRPRRP SR TSMEEDRLRLYTA+LTS
Sbjct: 9 RYYSPAWRTWAAVAIQLAWRRYQHRLTLTSLSFIRPRRPLSRSTSMEEDRLRLYTAMLTS 188
Query: 270 PKPNDQEEYDDFDF 283
PKPN DDFDF
Sbjct: 189 PKPNQ----DDFDF 218
>TC220014 homologue to UP|Q8H6U1 (Q8H6U1) Cyclic nucleotide-gated channel C
(Fragment), partial (41%)
Length = 963
Score = 130 bits (327), Expect = 7e-31
Identities = 67/158 (42%), Positives = 101/158 (63%), Gaps = 2/158 (1%)
Frame = +2
Query: 79 QVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHL--QSSQDLRDG 136
+VP+F++MD+ +L+ +CDR+K +++T+ I REGDPV MLF++RG L ++ R G
Sbjct: 5 RVPMFENMDEQLLDAMCDRLKPVLYTEESCIAREGDPVDEMLFIMRGKLLTVTTNGGRTG 184
Query: 137 VKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHF 196
+ L G+F G+ELL+W L LP S+ T+ TL EAF L+A+D+++V F
Sbjct: 185 FFNSEYLKAGDFCGEELLTWALDPQSSSNLPISTRTVQTLSEVEAFALKADDLKFVASQF 364
Query: 197 RYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHR 234
R K+ ++ + R+YS WRTWAA IQ AWRRY +
Sbjct: 365 RRLHSKQ-LRHTFRFYSQQWRTWAACFIQAAWRRYSKK 475
>CO984626
Length = 817
Score = 100 bits (248), Expect = 1e-21
Identities = 55/123 (44%), Positives = 75/123 (60%), Gaps = 2/123 (1%)
Frame = -2
Query: 111 REGDPVKRMLFVVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPP 168
REGDPV MLF++RG L + + R G + L G+F G+ELL+W L LP
Sbjct: 816 REGDPVGEMLFIMRGKLLTVTTNGGRTGFFNSEYLKAGDFCGEELLTWALDPQSSSNLPI 637
Query: 169 SSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAW 228
S+ T+ TL EAF L+A+D+++V FR K+ ++ + R+YS WRTWAA IQ AW
Sbjct: 636 STRTVQTLSEVEAFALKADDLKFVASQFRRLHSKQ-LRHTFRFYSQQWRTWAACFIQAAW 460
Query: 229 RRY 231
RRY
Sbjct: 459 RRY 451
>BQ297066 homologue to GP|24943196|gb cyclic nucleotide-gated channel C
{Phaseolus vulgaris}, partial (25%)
Length = 454
Score = 80.9 bits (198), Expect = 6e-16
Identities = 37/62 (59%), Positives = 45/62 (71%)
Frame = +2
Query: 16 RMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLD 75
+ R+ E WM R LP GLR+R+R YE+ RW RGVDE +IRNLP+ LRRDIK HLCL
Sbjct: 2 KRRDAEQWMSHRLLPDGLRERIRRYEQYRWQETRGVDEDNLIRNLPKDLRRDIKRHLCLA 181
Query: 76 LV 77
L+
Sbjct: 182LL 187
Score = 68.6 bits (166), Expect = 3e-12
Identities = 36/99 (36%), Positives = 57/99 (57%), Gaps = 2/99 (2%)
Frame = +1
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
QG ++ K P+F+ MD+ +L+ +CD +K +++T+ I REGDPV MLF
Sbjct: 139 QGFKKGHKASSLFGFADDCPMFEKMDEQLLDAMCDLLKPVLYTEESYIVREGDPVDEMLF 318
Query: 122 VVRGHL--QSSQDLRDGVKSYCTLGPGNFSGDELLSWCL 158
++RG L ++ R G + L G+F G+ELL+W L
Sbjct: 319 IMRGKLLTMTTNGGRTGFFNSEYLKAGDFCGEELLTWAL 435
>CO984580
Length = 752
Score = 71.6 bits (174), Expect(2) = 4e-15
Identities = 35/80 (43%), Positives = 49/80 (60%)
Frame = -1
Query: 152 ELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARY 211
ELL+W L LP S+ T+ T+ EAF L A+D+++V FR K+ ++ + R+
Sbjct: 608 ELLTWALDPNSSSNLPISTRTVETISEVEAFALTADDLKFVASQFRRLHSKQ-LQHAFRF 432
Query: 212 YSPGWRTWAAVAIQLAWRRY 231
YS W+TWAA IQ AWRRY
Sbjct: 431 YSSQWKTWAATFIQAAWRRY 372
Score = 26.9 bits (58), Expect(2) = 4e-15
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = -2
Query: 109 ITREGDPVKRMLFVVRG 125
I RE DPV MLF++RG
Sbjct: 742 IVREEDPVDEMLFIMRG 692
>BM178359 similar to PIR|T51519|T515 cyclic nucleotide-gated cation channel -
Arabidopsis thaliana, partial (8%)
Length = 435
Score = 50.1 bits (118), Expect(2) = 4e-12
Identities = 21/31 (67%), Positives = 27/31 (86%)
Frame = +3
Query: 79 QVPLFQHMDDLVLENICDRVKSLVFTKGETI 109
QVPLF ++DDL+L+NICDRVK LVF+K E +
Sbjct: 144 QVPLFHNLDDLILDNICDRVKPLVFSKDEKV 236
Score = 38.1 bits (87), Expect(2) = 4e-12
Identities = 16/29 (55%), Positives = 22/29 (75%)
Frame = +1
Query: 109 ITREGDPVKRMLFVVRGHLQSSQDLRDGV 137
I REGDPV RM+F+VRG ++ +Q L G+
Sbjct: 328 IIREGDPVPRMVFIVRGRIKRNQSLSKGM 414
>BM094700 similar to GP|8131898|gb| cyclic nucleotide-binding transporter 1
{Arabidopsis thaliana}, partial (17%)
Length = 427
Score = 54.7 bits (130), Expect = 5e-08
Identities = 26/59 (44%), Positives = 36/59 (60%)
Frame = +1
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
FL A ++ M LR R++E WM R L + LR++VR ER W A RGV+E ++ NL
Sbjct: 250 FLQALERRRLEMTLRRRDVEQWMSHRHLAEDLRRKVRQAERYNWAATRGVNEEMLLENL 426
>CD399837 similar to GP|24943192|gb cyclic nucleotide-gated channel A
{Phaseolus vulgaris}, partial (34%)
Length = 532
Score = 50.4 bits (119), Expect = 9e-07
Identities = 24/52 (46%), Positives = 34/52 (65%)
Frame = -3
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRY 231
EAF L A+D+++V FR K+ ++ + R+YS W+TWAA IQ A RRY
Sbjct: 530 EAFALMADDLKFVASQFRRLHSKQ-LQHAFRFYSSQWKTWAATFIQAAXRRY 378
>TC227288 similar to UP|CG17_ARATH (Q8L7Z0) Probable cyclic nucleotide-gated
ion channel 17 (Cyclic nucleotide-and
calmodulin-regulated ion channel 17), partial (9%)
Length = 558
Score = 43.1 bits (100), Expect = 1e-04
Identities = 20/43 (46%), Positives = 26/43 (59%)
Frame = +2
Query: 216 WRTWAAVAIQLAWRRYKHRLSLTSLSFIRPRRPASRCTSMEED 258
WRTWAA IQ AWRRYK R+++ LS +R P + E +
Sbjct: 32 WRTWAACFIQAAWRRYKKRITMKDLS-LRESIPLDETVASERE 157
>CA852571 similar to GP|8131898|gb|A cyclic nucleotide-binding transporter 1
{Arabidopsis thaliana}, partial (5%)
Length = 672
Score = 38.5 bits (88), Expect = 0.003
Identities = 24/54 (44%), Positives = 31/54 (56%)
Frame = +3
Query: 105 KGETITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCL 158
KG I G V +MLFVVRG L+S + DG + L G+ G+ELL+W L
Sbjct: 57 KGSRILSHGAVVDKMLFVVRGKLESIGE--DGTR--IPLSEGDACGEELLTWYL 206
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,267,926
Number of Sequences: 63676
Number of extensions: 188926
Number of successful extensions: 1386
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 1349
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1362
length of query: 283
length of database: 12,639,632
effective HSP length: 96
effective length of query: 187
effective length of database: 6,526,736
effective search space: 1220499632
effective search space used: 1220499632
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0590b.4


