

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.12
(583 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
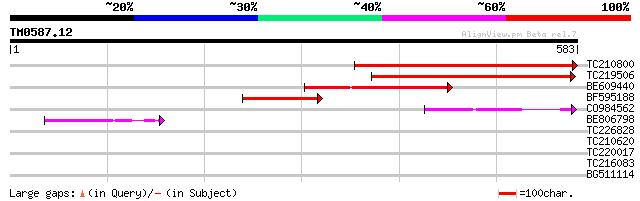
TC210800 similar to UP|Q9M8Z1 (Q9M8Z1) T6K12.3 protein (AT3g0435... 440 e-124
TC219506 similar to UP|Q9M8Z1 (Q9M8Z1) T6K12.3 protein (AT3g0435... 335 4e-92
BE609440 weakly similar to GP|20197184|gb expressed protein {Ara... 181 8e-46
BF595188 similar to GP|15450691|gb| AT3g04350/T6K12_3 {Arabidops... 147 2e-35
CO984562 130 2e-30
BE806798 similar to GP|20197184|gb| expressed protein {Arabidops... 70 3e-12
TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 31 1.3
TC210620 31 1.8
TC220017 similar to UP|Q7PC53 (Q7PC53) Chitinase B, partial (3%) 31 1.8
TC216083 similar to UP|Q84P21 (Q84P21) 4-coumarate-CoA ligase-li... 29 5.1
BG511114 29 6.7
>TC210800 similar to UP|Q9M8Z1 (Q9M8Z1) T6K12.3 protein (AT3g04350/T6K12_3),
partial (35%)
Length = 986
Score = 440 bits (1132), Expect = e-124
Identities = 204/229 (89%), Positives = 219/229 (95%)
Frame = +1
Query: 355 VKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAFSKVGEHVGDWE 414
++ GDLK AKLYVHVKPALGGTFTD+AMWVFCPFNGPSTLK GITS AFSKVGEHVGDWE
Sbjct: 10 IQAGDLKSAKLYVHVKPALGGTFTDIAMWVFCPFNGPSTLKIGITSRAFSKVGEHVGDWE 189
Query: 415 HFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASFPHPGTYIQ 474
HFTLRICNFSGELWSIYFSQHSGGKWVD+Y+LEYI+GNKA+VYSSK+GHAS+PHPGTY+Q
Sbjct: 190 HFTLRICNFSGELWSIYFSQHSGGKWVDAYELEYIDGNKAVVYSSKNGHASYPHPGTYLQ 369
Query: 475 GSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVYGSKT 534
GSSKLGIGIRNDA SNLYVDSS+QYE+VAAEYLGDVVREPQWLQ+MREWGPKIVY SKT
Sbjct: 370 GSSKLGIGIRNDATRSNLYVDSSVQYELVAAEYLGDVVREPQWLQFMREWGPKIVYDSKT 549
Query: 535 ELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDERW 583
ELDKIINALP LR SFV+L+KKLPVELYGEEGPTGPKEKNNWIGDERW
Sbjct: 550 ELDKIINALPGGLRNSFVSLIKKLPVELYGEEGPTGPKEKNNWIGDERW 696
>TC219506 similar to UP|Q9M8Z1 (Q9M8Z1) T6K12.3 protein (AT3g04350/T6K12_3),
partial (36%)
Length = 995
Score = 335 bits (858), Expect = 4e-92
Identities = 152/210 (72%), Positives = 176/210 (83%), Gaps = 1/210 (0%)
Frame = +3
Query: 373 LGGTFTDLAMWVFCPFNGPSTLKFGITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYF 432
LGG FTD+ MWVFCPFNGP+TLK + +I SK+GEHVGDWEHFTLRI NF+GELWS+YF
Sbjct: 3 LGGAFTDIVMWVFCPFNGPATLKVALMNIEMSKIGEHVGDWEHFTLRISNFTGELWSVYF 182
Query: 433 SQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNL 492
SQHSGG W+ ++DLE+ GNK IVYSSK GHASFPHPGTY+QGSSKLGIG+RNDA S
Sbjct: 183 SQHSGGGWIHAFDLEFNKGNKPIVYSSKDGHASFPHPGTYLQGSSKLGIGVRNDAAPSKF 362
Query: 493 YVDSSIQYEIVAAEYLGD-VVREPQWLQYMREWGPKIVYGSKTELDKIINALPFRLRISF 551
VDSSI+Y+IVAAEYLGD V+ EP WLQYMREWGP IVY S++E++KIIN LP +R S
Sbjct: 363 IVDSSIKYQIVAAEYLGDGVIAEPCWLQYMREWGPTIVYDSRSEIEKIINLLPLFVRFSV 542
Query: 552 VNLVKKLPVELYGEEGPTGPKEKNNWIGDE 581
NL + P ELYGEEGPTGPKEK+NW+GDE
Sbjct: 543 ENLFELFPTELYGEEGPTGPKEKDNWLGDE 632
>BE609440 weakly similar to GP|20197184|gb expressed protein {Arabidopsis
thaliana}, partial (26%)
Length = 461
Score = 181 bits (459), Expect = 8e-46
Identities = 87/153 (56%), Positives = 107/153 (69%), Gaps = 1/153 (0%)
Frame = +1
Query: 304 WFFSNRAMLFRKGV-STGEAIDAGGSNLPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKG 362
WFFSN A+L++KG S +I G+NLP DG +W+DLP D + +E VK GDLK
Sbjct: 1 WFFSNGALLYKKGQESKPVSISPNGANLPQDPNIDGAYWVDLPADSTN-KERVKKGDLKS 177
Query: 363 AKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAFSKVGEHVGDWEHFTLRICN 422
A YVHVKP LGGTFTD+AMWVF PFNGP+ K ++ K+GEHVGDWEH TLR+ N
Sbjct: 178 AISYVHVKPMLGGTFTDIAMWVFYPFNGPARAKVEFLTVNLGKIGEHVGDWEHVTLRVSN 357
Query: 423 FSGELWSIYFSQHSGGKWVDSYDLEYINGNKAI 455
F+GEL +YFSQHS G W DS LE+ +GNK +
Sbjct: 358 FNGELKHVYFSQHSKGVWFDSSQLEFQSGNKPL 456
>BF595188 similar to GP|15450691|gb| AT3g04350/T6K12_3 {Arabidopsis
thaliana}, partial (10%)
Length = 249
Score = 147 bits (370), Expect = 2e-35
Identities = 68/82 (82%), Positives = 75/82 (90%)
Frame = +2
Query: 240 VSVGTFFCSSCVNMGEQLPVVCLKNLNPVLPAMPCLQQIHALIEHYGPTVYFHPEEVYLP 299
VSVGTFFC+SC N G++L VVCLKNLNPVLPAMP L IHALIEHYGPTV+FHPEEVYLP
Sbjct: 2 VSVGTFFCTSCWNNGDELSVVCLKNLNPVLPAMPRLH*IHALIEHYGPTVFFHPEEVYLP 181
Query: 300 SSVDWFFSNRAMLFRKGVSTGE 321
SSVDWFF+N A+L+RKGVSTGE
Sbjct: 182 SSVDWFFNNGALLYRKGVSTGE 247
>CO984562
Length = 623
Score = 130 bits (327), Expect = 2e-30
Identities = 66/156 (42%), Positives = 84/156 (53%)
Frame = -1
Query: 427 LWSIYFSQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRND 486
LW +YFSQHS G+W D+ +LE+ NGN+ + YSS GHA FP PG +QG G+G+RND
Sbjct: 623 LWKVYFSQHSKGQWEDASELEFQNGNRPVAYSSLHGHALFPKPGLVMQGMR--GLGVRND 450
Query: 487 ACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVYGSKTELDKIINALPFR 546
A S+ +D + +EIVAAEYLG +REP WL + WGPK
Sbjct: 449 AAKSDAVMDMATWFEIVAAEYLGSQIREPPWLNFCMNWGPK------------------- 327
Query: 547 LRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDER 582
EGP GPK+K+ W GDER
Sbjct: 326 -------------------EGPKGPKQKDMWKGDER 276
>BE806798 similar to GP|20197184|gb| expressed protein {Arabidopsis
thaliana}, partial (17%)
Length = 412
Score = 70.1 bits (170), Expect = 3e-12
Identities = 46/125 (36%), Positives = 67/125 (52%), Gaps = 1/125 (0%)
Frame = +1
Query: 36 YSVIVHVCVITGQGFASGIINLGE-LEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPD 94
Y + H G GFA+GII+LG L V +++F VW + F++P G+ +
Sbjct: 61 YWINCHGF*FVGGGFATGIIDLGGGLLVSQISTFNKVWTTYEGGPNNLGATFFEPTGLSE 240
Query: 95 GFHVLSHYCQPSYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNA 154
GF +L YCQP+ NKPL G VLV ++ + S+ AL +P+DY LVW N
Sbjct: 241 GFFMLGCYCQPN-NKPLHGCVLVGKDNSSTSN-----------GALAEPVDYKLVW--NT 378
Query: 155 GSKEI 159
S++I
Sbjct: 379 KSQKI 393
>TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (47%)
Length = 1488
Score = 31.2 bits (69), Expect = 1.3
Identities = 24/75 (32%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = -2
Query: 330 LPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKLYVH--VKPALGGTFTDLAMWVFCP 387
LP+G +D W + CD Q F++H D L +H V+PA T+ + CP
Sbjct: 875 LPLGFNHDRGTWYSV-CDGFPQ--FLRHLDAL*GPL*LHESVQPASEATYVGQYRPMLCP 705
Query: 388 FNGPSTLKFGITSIA 402
F S + SIA
Sbjct: 704 FTQASPFPIPVKSIA 660
>TC210620
Length = 698
Score = 30.8 bits (68), Expect = 1.8
Identities = 17/64 (26%), Positives = 28/64 (43%)
Frame = +1
Query: 143 PLDYTLVWCSNAGSKEIVTSSAYFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLT 202
P+ Y LVW E + W P+ P+G+ A G + +P D + C+ L
Sbjct: 28 PMGYDLVW---RNCPEDYVTPLSIWHPRAPDGFVAPGCVAIAGYLEPEPDLVYCIAESLV 198
Query: 203 DKCE 206
++ E
Sbjct: 199 EETE 210
>TC220017 similar to UP|Q7PC53 (Q7PC53) Chitinase B, partial (3%)
Length = 825
Score = 30.8 bits (68), Expect = 1.8
Identities = 14/35 (40%), Positives = 23/35 (65%)
Frame = +1
Query: 317 VSTGEAIDAGGSNLPIGGTNDGEFWIDLPCDDDDQ 351
V +A++AGG+++ G +DGE ++ DDDDQ
Sbjct: 67 VGQSDALNAGGADVDGGNDDDGEEEDEVDDDDDDQ 171
>TC216083 similar to UP|Q84P21 (Q84P21) 4-coumarate-CoA ligase-like protein,
partial (80%)
Length = 1411
Score = 29.3 bits (64), Expect = 5.1
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 4/59 (6%)
Frame = -1
Query: 317 VSTGEAIDAGGS--NLPIGGTNDGEFWIDLPCDDDDQREFVKHGD--LKGAKLYVHVKP 371
VS E +D G +L +G +DG P + DQ +VKHGD +G L +HV+P
Sbjct: 847 VSGAEPLDRGEHVVHLELGEHDDGGAGSQKPRGEGDQAVYVKHGDRAYEGFVL-LHVEP 674
>BG511114
Length = 357
Score = 28.9 bits (63), Expect = 6.7
Identities = 14/26 (53%), Positives = 15/26 (56%)
Frame = +2
Query: 47 GQGFASGIINLGELEVCSVTSFELVW 72
G A G I LGE+EV V FE VW
Sbjct: 278 GGSXACGRICLGEIEVIKVNKFEKVW 355
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,037,389
Number of Sequences: 63676
Number of extensions: 519281
Number of successful extensions: 2394
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 2374
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2387
length of query: 583
length of database: 12,639,632
effective HSP length: 102
effective length of query: 481
effective length of database: 6,144,680
effective search space: 2955591080
effective search space used: 2955591080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0587.12


