

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0546.13
(1118 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
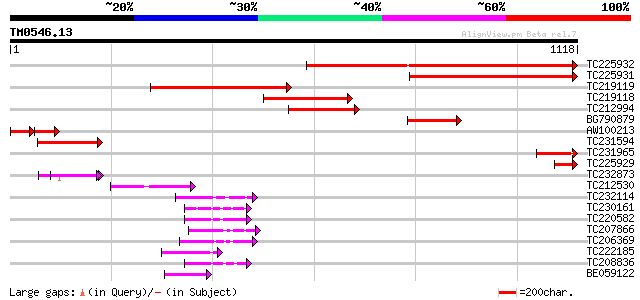
Sequences producing significant alignments: (bits) Value
TC225932 1030 0.0
TC225931 637 0.0
TC219119 homologue to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-term... 565 e-161
TC219118 similar to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-termin... 342 4e-94
TC212994 similar to UP|Q9FPT1 (Q9FPT1) Ubiquitin-specific protea... 220 2e-57
BG790879 174 2e-43
AW100213 similar to GP|10178116|db ubiquitin carboxyl-terminal h... 96 3e-39
TC231594 139 9e-33
TC231965 130 4e-30
TC225929 100 4e-21
TC232873 weakly similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, par... 78 2e-14
TC212530 similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific protea... 69 2e-11
TC232114 similar to UP|Q8VZF5 (Q8VZF5) AT3g14400/MLN21_18, parti... 61 2e-09
TC230161 similar to UP|Q9C585 (Q9C585) Ubiquitin-specific protea... 59 9e-09
TC220582 similar to UP|Q9C585 (Q9C585) Ubiquitin-specific protea... 59 9e-09
TC207866 homologue to UP|O24454 (O24454) Ubiquitin-specific prot... 56 1e-07
TC206369 similar to UP|Q9FPS3 (Q9FPS3) Ubiquitin-specific protea... 53 7e-07
TC222185 weakly similar to UP|Q9FKP5 (Q9FKP5) Similarity to ubiq... 53 9e-07
TC208836 similar to UP|Q9ZSB5 (Q9ZSB5) F3H7.5 protein, partial (... 52 1e-06
BE059122 51 3e-06
>TC225932
Length = 2244
Score = 1030 bits (2664), Expect = 0.0
Identities = 501/533 (93%), Positives = 518/533 (96%)
Frame = +1
Query: 586 YFDLVDHDKVRSFRVQKQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTP 645
YFDLVDHDKVRSFRVQKQ SFNLFK+EVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLT
Sbjct: 1 YFDLVDHDKVRSFRVQKQTSFNLFKDEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTH 180
Query: 646 AEEAQSVGQVREVSNKVHNAELKLFLEVELGPDLRPIAPSDKTKDDILLFFKLYDPEKEE 705
EEAQSVGQ+REVSNKVHNA+LKLFLEVELG DLRPIAP DKTKDDILLFFKLYD EKEE
Sbjct: 181 MEEAQSVGQLREVSNKVHNAQLKLFLEVELGLDLRPIAPPDKTKDDILLFFKLYDTEKEE 360
Query: 706 LRYVGRLFVKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRAS 765
LRYVGRLFVK+TGKPSEILTRLN+MAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRAS
Sbjct: 361 LRYVGRLFVKATGKPSEILTRLNKMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRAS 540
Query: 766 QLEDGDIICFQKAPAMDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRL 825
QLEDGDIICFQKAPA+D+E HVRYPDVPSYLEYVHNRQVVHFRSL+KPKEDDFCLEMSRL
Sbjct: 541 QLEDGDIICFQKAPAIDNE-HVRYPDVPSYLEYVHNRQVVHFRSLEKPKEDDFCLEMSRL 717
Query: 826 YTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDI 885
YTYDDVVEKVAQQL L+DPS IRLTPHNCYSQQPKPQPIKYRGV+HLSDMLVHYNQTSDI
Sbjct: 718 YTYDDVVEKVAQQLGLEDPSIIRLTPHNCYSQQPKPQPIKYRGVEHLSDMLVHYNQTSDI 897
Query: 886 LYYEILDIPLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHP 945
LYYE+LDIPLPELQGLKTLKVAF+HATKDEVV HTIRLPKQSTVGDVLDDLKTKVELS P
Sbjct: 898 LYYEVLDIPLPELQGLKTLKVAFHHATKDEVVIHTIRLPKQSTVGDVLDDLKTKVELSDP 1077
Query: 946 NAELRLLEVFYHKIYKVFPPNEKIETINDQYWTLRAEEVPEEEKNLGPHDRLIHVYHFTK 1005
AELRLLEVFYHKIYKVFPPNEKIE+INDQYWTLRAEE+PEEEKNLGPHDRLIHVYHFTK
Sbjct: 1078 EAELRLLEVFYHKIYKVFPPNEKIESINDQYWTLRAEEIPEEEKNLGPHDRLIHVYHFTK 1257
Query: 1006 DTSQNQMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPEYL 1065
DT+QNQMQIQNFGEPFFLVI EGETL EIKVRIQKKLQVPDDEF KWKFAFF+LGRPEYL
Sbjct: 1258 DTAQNQMQIQNFGEPFFLVIHEGETLAEIKVRIQKKLQVPDDEFVKWKFAFFSLGRPEYL 1437
Query: 1066 QDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
QDSDIVS+RFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN
Sbjct: 1438 QDSDIVSSRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1596
>TC225931
Length = 1469
Score = 637 bits (1644), Expect = 0.0
Identities = 305/330 (92%), Positives = 319/330 (96%)
Frame = +2
Query: 789 YPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIR 848
YPDVPSYLEYVHNRQVVHFRSLD+PKEDDF LEMSRL+TYDDVVE+VAQQL LDDPSKIR
Sbjct: 2 YPDVPSYLEYVHNRQVVHFRSLDRPKEDDFFLEMSRLFTYDDVVERVAQQLGLDDPSKIR 181
Query: 849 LTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEILDIPLPELQGLKTLKVAF 908
LTPHNCYSQQPKPQPIKYRGV+HLSDMLVHYNQTSDILYYE+LDIPLPELQGLKTLKVAF
Sbjct: 182 LTPHNCYSQQPKPQPIKYRGVEHLSDMLVHYNQTSDILYYEVLDIPLPELQGLKTLKVAF 361
Query: 909 YHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHPNAELRLLEVFYHKIYKVFPPNEK 968
+HATK+EVV HTIRLPKQSTVGDVLDDLKTKVELS P AELRLLEVFYHKIYKVFPPNEK
Sbjct: 362 HHATKEEVVIHTIRLPKQSTVGDVLDDLKTKVELSDPEAELRLLEVFYHKIYKVFPPNEK 541
Query: 969 IETINDQYWTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQNQMQIQNFGEPFFLVIREG 1028
IE+INDQYWTLRAEE+PEEEKNLG HDRLIHVYHF K+T+QNQMQIQNFGEPFFLVI EG
Sbjct: 542 IESINDQYWTLRAEEIPEEEKNLGSHDRLIHVYHFNKETAQNQMQIQNFGEPFFLVIHEG 721
Query: 1029 ETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPEYLQDSDIVSNRFQRRDVYGAWEQYL 1088
ETL EIKVRIQKKLQVPDDEF KWKFAF +LGRPEYLQDSD+VS+RFQRRDVYGAWEQYL
Sbjct: 722 ETLDEIKVRIQKKLQVPDDEFCKWKFAFLSLGRPEYLQDSDVVSSRFQRRDVYGAWEQYL 901
Query: 1089 GLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
GLEHTDNAPKRSYAVNQNRHTFEKPVKIYN
Sbjct: 902 GLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 991
>TC219119 homologue to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-terminal
hydrolase, partial (25%)
Length = 836
Score = 565 bits (1455), Expect = e-161
Identities = 271/278 (97%), Positives = 275/278 (98%)
Frame = +2
Query: 278 MQHDVQELNRVLCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQ 337
MQHDVQELNRV CEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQ
Sbjct: 2 MQHDVQELNRVXCEKLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQ 181
Query: 338 LDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEY 397
LDVKGC DVYASFDKYVEVE LEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEY
Sbjct: 182 LDVKGCPDVYASFDKYVEVERLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEY 361
Query: 398 DFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYY 457
DFMRDTMVKINDRYEFPL+LDLDR++GKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYY
Sbjct: 362 DFMRDTMVKINDRYEFPLQLDLDRENGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYY 541
Query: 458 AFIRPTLSDQWYKFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYM 517
AFIRPTLS+QWYKFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYM
Sbjct: 542 AFIRPTLSEQWYKFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYM 721
Query: 518 LVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEH 555
LVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEH
Sbjct: 722 LVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEH 835
>TC219118 similar to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-terminal
hydrolase, partial (16%)
Length = 566
Score = 342 bits (878), Expect = 4e-94
Identities = 166/177 (93%), Positives = 173/177 (96%)
Frame = +3
Query: 500 PGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAEHLRERLKKEQEEKEHKKKE 559
P FNNTPFKFTKYSNAYMLVYIRE+DKDK+ICNVDEKDIAEHLRERLKKEQEEKEHKKKE
Sbjct: 36 PXFNNTPFKFTKYSNAYMLVYIRESDKDKIICNVDEKDIAEHLRERLKKEQEEKEHKKKE 215
Query: 560 KAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRVQKQMSFNLFKEEVAKEFGI 619
KAEAHLYTIIKVAR+E+LKEQIGKDIYFDLVDHDKVRSFRVQKQ SFNLFKEEVAKE+GI
Sbjct: 216 KAEAHLYTIIKVARDEELKEQIGKDIYFDLVDHDKVRSFRVQKQTSFNLFKEEVAKEYGI 395
Query: 620 PVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSVGQVREVSNKVHNAELKLFLEVELG 676
PVQFQR+WLWAKRQNHTYRPNRPLT EEAQSVGQ+REVSNKVHNAELKLFLEVELG
Sbjct: 396 PVQFQRYWLWAKRQNHTYRPNRPLTHIEEAQSVGQLREVSNKVHNAELKLFLEVELG 566
>TC212994 similar to UP|Q9FPT1 (Q9FPT1) Ubiquitin-specific protease 12,
partial (12%)
Length = 423
Score = 220 bits (561), Expect = 2e-57
Identities = 107/140 (76%), Positives = 124/140 (88%)
Frame = +2
Query: 550 QEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRVQKQMSFNLF 609
Q+EKE K+KEKAEAHLYT IKVA +EDL+EQIG +I FDLVD+DKVRSFRVQ QM F +F
Sbjct: 2 QDEKELKRKEKAEAHLYTTIKVACDEDLREQIGNNILFDLVDYDKVRSFRVQIQMPFMVF 181
Query: 610 KEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSVGQVREVSNKVHNAELKL 669
KEE+AKEFGIP+Q+QRFWLWAKRQN+TYRPNR LTP EEAQSVG +REVS K +NAELKL
Sbjct: 182 KEEIAKEFGIPIQYQRFWLWAKRQNNTYRPNRALTPQEEAQSVGLLREVSTKANNAELKL 361
Query: 670 FLEVELGPDLRPIAPSDKTK 689
FLE+E+G DLRPI P +K+K
Sbjct: 362 FLELEMGQDLRPIPPPEKSK 421
>BG790879
Length = 327
Score = 174 bits (441), Expect = 2e-43
Identities = 80/107 (74%), Positives = 94/107 (87%)
Frame = +2
Query: 785 EHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDDVVEKVAQQLNLDDP 844
E RYPDVPS+L YVHNR VV FR+L+KPKED+F LE+++L TYD+VVE+VAQ + L DP
Sbjct: 5 ERYRYPDVPSFLVYVHNRLVVRFRTLEKPKEDEFSLELTKLDTYDNVVEEVAQHIGLSDP 184
Query: 845 SKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEIL 891
SKIRLT HNCYSQQPKPQ IKYRG++HLSDML+H NQTSDILYYE+L
Sbjct: 185 SKIRLTSHNCYSQQPKPQSIKYRGMEHLSDMLIHSNQTSDILYYEVL 325
>AW100213 similar to GP|10178116|db ubiquitin carboxyl-terminal hydrolase
{Arabidopsis thaliana}, partial (5%)
Length = 408
Score = 95.5 bits (236), Expect(2) = 3e-39
Identities = 41/49 (83%), Positives = 46/49 (93%)
Frame = +3
Query: 50 EEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYL 98
++P SRFTW+IDNFSRMN KKLYSE+FVVGGYKWRVLIFP+GNNVDYL
Sbjct: 261 KDPSTSRFTWKIDNFSRMNTKKLYSEIFVVGGYKWRVLIFPRGNNVDYL 407
Score = 86.3 bits (212), Expect(2) = 3e-39
Identities = 43/49 (87%), Positives = 44/49 (89%), Gaps = 1/49 (2%)
Frame = +2
Query: 1 MTVMTPAPIDQQ-EDEEMLVPHTDLPENNHQPMEVVAQPEAAPTVESQP 48
MTVMTPAPIDQQ EDEEMLVPHTD ENNHQPMEV AQP+AA TVESQP
Sbjct: 110 MTVMTPAPIDQQHEDEEMLVPHTDWAENNHQPMEVAAQPDAANTVESQP 256
>TC231594
Length = 1086
Score = 139 bits (349), Expect = 9e-33
Identities = 63/128 (49%), Positives = 90/128 (70%)
Frame = +3
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWS 115
+FTW I NFS+++ KKLYSE F + + W++L +PKG +V YLS+YLD A NLP+GWS
Sbjct: 81 KFTWTISNFSKLDSKKLYSEKF*LDHHTWKILAYPKGTDVGYLSIYLD-AGVVNLPFGWS 257
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
++A F A++NQ +K T K+T QFNA E WGF F+ L EL D S G+++NDT ++
Sbjct: 258 KFANFKFALINQANDKMTKIKETTKQFNATEIAWGFPKFIHLDELNDSSSGFMVNDTCII 437
Query: 176 EAEVLVRR 183
E ++LV +
Sbjct: 438 EVQILVSK 461
>TC231965
Length = 710
Score = 130 bits (326), Expect = 4e-30
Identities = 64/80 (80%), Positives = 67/80 (83%)
Frame = +2
Query: 1039 QKKLQVPDDEFEKWKFAFFALGRPEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPK 1098
QKKLQVPD+EF KWKFAF + GR EYLQDSDIVS RFQRRD GAWEQYLGLEHTDNA K
Sbjct: 2 QKKLQVPDEEFSKWKFAFLSFGRXEYLQDSDIVSARFQRRDXXGAWEQYLGLEHTDNASK 181
Query: 1099 RSYAVNQNRHTFEKPVKIYN 1118
RS A NQNRH EK VKIY+
Sbjct: 182 RSNAANQNRH--EKAVKIYD 235
>TC225929
Length = 664
Score = 100 bits (249), Expect = 4e-21
Identities = 45/45 (100%), Positives = 45/45 (100%)
Frame = +3
Query: 1074 RFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
RFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN
Sbjct: 3 RFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 137
>TC232873 weakly similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, partial (28%)
Length = 954
Score = 78.2 bits (191), Expect = 2e-14
Identities = 40/107 (37%), Positives = 62/107 (57%), Gaps = 5/107 (4%)
Frame = +1
Query: 80 GGYKWRVLIFPKGNNVD----YLSMYLDVADSTNLPYGWSRYAQFSLAVVNQIQNKYTVR 135
GGYKW + I+P GN ++S+YL + DS++LP W A + + N I ++Y
Sbjct: 1 GGYKWSLSIYPTGNTKGGGEGHVSIYLVLMDSSSLPVDWEVNAIVNFSAYNFIDDEYVAT 180
Query: 136 KDTQH-QFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEVLV 181
+DT +F+ +++WG F+ + DPS GYL++DT V AEV V
Sbjct: 181 QDTNVLRFHVLKTEWGVAKFIDIDTFNDPSNGYLMDDTCVFGAEVFV 321
Score = 73.9 bits (180), Expect = 4e-13
Identities = 40/131 (30%), Positives = 71/131 (53%), Gaps = 3/131 (2%)
Frame = +1
Query: 58 TWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKG---NNVDYLSMYLDVADSTNLPYGW 114
+W+ DNFS + K SE FV G Y+W+++++P G + +S++L + ST LP
Sbjct: 385 SWKFDNFSLAKLDKYESESFVGGNYRWKLILYPNGIFEGKGNSISLFLTLEVST-LPPNT 561
Query: 115 SRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLV 174
+ +L QI + + +F++ S WG + L +L DP+ G+L+NDT +
Sbjct: 562 KLVVECTLRAKKQISGHH-AQTGFCRKFSSSNSTWGTRQLVALAKLTDPNSGFLVNDTCI 738
Query: 175 VEAEVLVRRIV 185
+EAE + ++
Sbjct: 739 LEAEFTILGLI 771
>TC212530 similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific protease 26,
partial (21%)
Length = 685
Score = 68.6 bits (166), Expect = 2e-11
Identities = 54/175 (30%), Positives = 82/175 (46%), Gaps = 9/175 (5%)
Frame = +3
Query: 200 GLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTT---ENDMPAGSIPLALQSLFYKLQYS 256
GL N GATCY NS+LQ LY FR+ ++ + ++ + L +Q K+ +
Sbjct: 174 GLTNLGATCYANSILQCLYMNKSFREGMFSVERDVLQQHPVLDQLARLFVQLHISKMAFI 353
Query: 257 DTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGT---IQKLFEG- 312
D+S K L D+ +Q D E +L LE + + V +Q LF G
Sbjct: 354 DSSPFVKTLE-------LDNGVQQDSHEFLTLLLSLLERCLSHSKVPKATTIVQDLFRGS 512
Query: 313 --HHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKY 365
H +C S++ E FY+L+L+VKG + S + Y+ E L GDN+Y
Sbjct: 513 VSHVTTCSQCGRDSEASSKMEDFYELELNVKGLKSLDESLE*YLTAEELNGDNQY 677
>TC232114 similar to UP|Q8VZF5 (Q8VZF5) AT3g14400/MLN21_18, partial (26%)
Length = 659
Score = 61.2 bits (147), Expect = 2e-09
Identities = 51/164 (31%), Positives = 76/164 (46%), Gaps = 2/164 (1%)
Frame = +2
Query: 327 STRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYG-LQDAKKGVLFIDFP 385
S + + D+ LDV + + S K+ + E L+G+ KY + L AKK + + P
Sbjct: 2 SNKVDEIMDISLDVFHSNSLKDSMQKFFQPEVLDGNKKYKCDSCKKLVAAKKQMSILQAP 181
Query: 386 PVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVL 445
+L +QLKRFE KI+ F L L K A ++ + Y L +
Sbjct: 182 NILVIQLKRFEGILGG----KIDKAVAFEEVLVLSSFMCK-----ASQDPQPEYKLFGTI 334
Query: 446 VHSG-GVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQ 488
VHSG GHYYA+I+ + +WY DD VT + L E+
Sbjct: 335 VHSGYSPESGHYYAYIKDAMG-RWYCCDDSCVTVATLQEVLSEK 463
>TC230161 similar to UP|Q9C585 (Q9C585) Ubiquitin-specific protease-like
protein, partial (18%)
Length = 1067
Score = 59.3 bits (142), Expect = 9e-09
Identities = 40/134 (29%), Positives = 69/134 (50%), Gaps = 3/134 (2%)
Frame = +3
Query: 346 VYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDF---PPVLQLQLKRFEYDFMRD 402
+Y + +++ EPL ++ ++ G ++ ++ +D P +L + LKRF+Y R
Sbjct: 153 LYKCLEAFLQEEPLGPEDMWYCP--GCKEHRQASKKLDLWRLPEILVIHLKRFQYS--RY 320
Query: 403 TMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRP 462
K+ +FP++ + D Y++ D + YTL++V H G + GGHY AF+
Sbjct: 321 LKNKLETYVDFPVD---NLDLSAYITYGNDESYH--YTLYAVSNHYGSMGGGHYTAFVHR 485
Query: 463 TLSDQWYKFDDERV 476
DQWY FDD V
Sbjct: 486 G-GDQWYDFDDSHV 524
>TC220582 similar to UP|Q9C585 (Q9C585) Ubiquitin-specific protease-like
protein, partial (16%)
Length = 1009
Score = 59.3 bits (142), Expect = 9e-09
Identities = 41/132 (31%), Positives = 67/132 (50%), Gaps = 1/132 (0%)
Frame = +3
Query: 346 VYASFDKYVEVEPLEGDNKYHAEQYGL-QDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTM 404
+Y + +++ EPL ++ ++ Q A K + P +L + LKRF Y R
Sbjct: 39 IYKCLEAFLKEEPLGPEDMWYCPNCKKPQQAYKKLDLWRLPEILVVHLKRFSYS--RYFK 212
Query: 405 VKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTL 464
K+ +F + D D Y++ +++ N Y L+++ H GG+ GGHY AF+R
Sbjct: 213 NKLETFVDFQIN---DLDLSTYVAHGNNQS-SNRYVLYAISCHYGGLGGGHYTAFVRYGY 380
Query: 465 SDQWYKFDDERV 476
D+WY FDD RV
Sbjct: 381 -DKWYDFDDSRV 413
>TC207866 homologue to UP|O24454 (O24454) Ubiquitin-specific protease
(AtUBP3) (AT4g39910/T5J17_80) , partial (43%)
Length = 784
Score = 55.8 bits (133), Expect = 1e-07
Identities = 45/144 (31%), Positives = 71/144 (49%), Gaps = 2/144 (1%)
Frame = +3
Query: 353 YVEVEPLEGDNKYHAEQY-GLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRY 411
+ E L ++K ++ LQ+A+K + P +L + LKRF+Y K++ R
Sbjct: 39 FSSTETLNAEDKLFCDKCCSLQEAQKRMKIKKPPHILVIHLKRFKYMEQLGRYKKLSYRV 218
Query: 412 EFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQWYK 470
FPLEL L DAD Y+L +V+VH G G + GHY + ++ + W
Sbjct: 219 VFPLELKLSN-----TVEDADIE----YSLFAVVVHVGSGPNHGHYVSLVKS--HNHWLF 365
Query: 471 FDDERVTKEDTKRALEEQYGGEEE 494
FDDE V D + A++ +G +E
Sbjct: 366 FDDENVEMID-ESAVQTFFGSSQE 434
>TC206369 similar to UP|Q9FPS3 (Q9FPS3) Ubiquitin-specific protease 24,
partial (47%)
Length = 1305
Score = 53.1 bits (126), Expect = 7e-07
Identities = 43/155 (27%), Positives = 66/155 (41%), Gaps = 2/155 (1%)
Frame = +1
Query: 336 LQLDV--KGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLK 393
L LD+ H + + + E LEG + G+ A+K V + P ++ L L
Sbjct: 466 LHLDIYPDAVHTIEDALHLFSAPETLEGYRTSLTAKAGVVTARKSVQIVTLPKIMILHLM 645
Query: 394 RFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHG 453
RF Y T K++ FPLEL L RD +SP + Y L + + H G
Sbjct: 646 RFGYGSQGST--KLHKPVHFPLELVLGRD--LLVSPSTEGRK---YELVATITHHGMEPS 804
Query: 454 GHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQ 488
+Y + +W +FDD+ V T + L +Q
Sbjct: 805 KGHYTADAQYPNGRWLRFDDQSVFAIGTNKVLHDQ 909
>TC222185 weakly similar to UP|Q9FKP5 (Q9FKP5) Similarity to ubiquitin
carboxyl-terminal hydrolase, partial (26%)
Length = 804
Score = 52.8 bits (125), Expect = 9e-07
Identities = 37/125 (29%), Positives = 66/125 (52%), Gaps = 4/125 (3%)
Frame = +2
Query: 299 GTVVEGT--IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKG-CHDVYASFDKYVE 355
G++ E T + + F G+ ++ I+C+ KS R+E DL ++++G + + ++
Sbjct: 14 GSLEEDTTLMGQTFGGYLLSKIKCMRCGGKSERQERMMDLTVEIEGEITTLVEALRRFTS 193
Query: 356 VEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFP 414
E L+G+NKYH + + AKK + + P VL + LKRF+ K+N +FP
Sbjct: 194 TETLDGENKYHCVRCKSYEKAKKKLTVSEAPNVLTVALKRFQ----SGKFGKLNKPIQFP 361
Query: 415 LELDL 419
L+L
Sbjct: 362 EILNL 376
>TC208836 similar to UP|Q9ZSB5 (Q9ZSB5) F3H7.5 protein, partial (17%)
Length = 1117
Score = 52.0 bits (123), Expect = 1e-06
Identities = 40/135 (29%), Positives = 67/135 (49%), Gaps = 3/135 (2%)
Frame = +1
Query: 346 VYASFDKYVEVEPLEGDNKYHAEQYGL-QDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTM 404
+++ + ++ EPL D+ ++ + + A K + P +L LKRF Y R
Sbjct: 127 LFSCLEAFLTEEPLGPDDMWYCPRCKEHRQATKKLDLWKLPEILVFHLKRFSYS--RYLK 300
Query: 405 VKINDRYEFPLE-LDLDRDDGKYL-SPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRP 462
K++ FP+ LDL KY+ S D V +LY + + H GG+ GGHY A+ +
Sbjct: 301 NKLDTFVNFPIHNLDLT----KYVKSKDGPSYVYDLYAISN---HYGGLGGGHYTAYCKL 459
Query: 463 TLSDQWYKFDDERVT 477
++W+ FDD V+
Sbjct: 460 IDENKWFHFDDSHVS 504
>BE059122
Length = 387
Score = 50.8 bits (120), Expect = 3e-06
Identities = 27/93 (29%), Positives = 49/93 (52%), Gaps = 1/93 (1%)
Frame = +3
Query: 306 IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKY 365
+ K F+G N C+ + + R E+F+DL LD++ + + + E L ++K+
Sbjct: 81 VHKNFQGILTNETRCLRCETVTARDETFFDLSLDIEQNSSITSCLKNFSSTETLNAEDKF 260
Query: 366 HAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEY 397
++ LQ+A+K + P VL + LKRF+Y
Sbjct: 261 FCDKCCSLQEAQKRMKIKKPPHVLVIHLKRFKY 359
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,394,217
Number of Sequences: 63676
Number of extensions: 621484
Number of successful extensions: 3426
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 3212
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3370
length of query: 1118
length of database: 12,639,632
effective HSP length: 107
effective length of query: 1011
effective length of database: 5,826,300
effective search space: 5890389300
effective search space used: 5890389300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0546.13


