

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0545.3
(2087 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
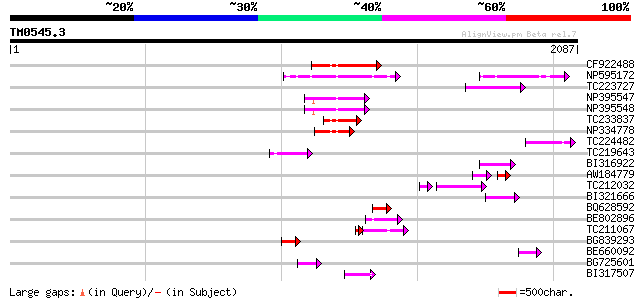
Score E
Sequences producing significant alignments: (bits) Value
CF922488 199 1e-50
NP595172 polyprotein [Glycine max] 197 5e-50
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 156 1e-37
NP395547 reverse transcriptase [Glycine max] 151 3e-36
NP395548 reverse transcriptase [Glycine max] 143 7e-34
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 136 8e-32
NP334778 reverse transcriptase [Glycine max] 129 2e-29
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 105 3e-22
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 102 1e-21
BI316922 101 3e-21
AW184779 60 1e-16
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 65 1e-16
BI321666 78 5e-14
BQ628592 72 2e-12
BE802896 66 1e-10
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 56 2e-10
BG839293 65 2e-10
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 62 3e-09
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 62 3e-09
BI317507 59 3e-08
>CF922488
Length = 741
Score = 199 bits (506), Expect = 1e-50
Identities = 105/255 (41%), Positives = 163/255 (63%)
Frame = +3
Query: 1112 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA* 1171
V+K++GK+ +C+D+RDLN A+PKD++ +P ++VD+ S +DG+SGYNQI IA
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 1172 EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDD 1231
ED+ KT F GT+ + M FGLKN G ATYQR M +F D + ++VY+DD
Sbjct: 183 EDMEKTTFIT--LWGTFCYKAMSFGLKNVG-----ATYQRAMVALF*DMMHKEIEVYMDD 341
Query: 1232 IVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKA 1291
++VKS + + HL++LRK F R+RKY L++NP KC F V + L F+ +GIE++ NK
Sbjct: 342 MIVKSRTEEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKV 521
Query: 1292 KAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKA 1351
K IL+ + P ++KQ+Q LG++N++ RFI+ L + + L K +W+ + A
Sbjct: 522 KVILEMAKPHTEKQVQGFLGRLNYIVRFIS*LIATCEPL--FILLCKNQFVKWDHDC*VA 695
Query: 1352 FDELKKYLSNPPVMI 1366
F+ +K+ L NP V++
Sbjct: 696 FERIKQCLINPHVLV 740
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 197 bits (500), Expect = 5e-50
Identities = 142/428 (33%), Positives = 214/428 (49%)
Frame = +1
Query: 1009 DGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKP 1068
D L+ PT I DPEL LL + FA P ++ +P+K+ P
Sbjct: 1645 DTLKDLPTNI----DPEL----AILLHTYAQVFAVPASLPPQREQDHA---IPLKQGSGP 1791
Query: 1069 VKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDL 1128
VK P R+ +I++ I+ +L I+ + L ++ V KK+G R C D+R L
Sbjct: 1792 VKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSPFSLP-ILLVKKKDGSWRFCTDYRAL 1968
Query: 1129 NAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTY 1188
NA T KD + MP + ++D G +Y S LD SGY+QI + ED KTAFR G Y
Sbjct: 1969 NAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHH--GHY 2142
Query: 1189 EWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRK 1248
EW+VMPFGL NA AT+Q +MN IF + F+ V+ DDI++ S S HL HL
Sbjct: 2143 EWLVMPFGLTNAP-----ATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLES 2307
Query: 1249 SFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQS 1308
+ ++++ L KC+FG D+LG V G+ + K +A+LD P + KQL+
Sbjct: 2308 VLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRG 2487
Query: 1309 LLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPP 1368
LG + RRFI + + + LL ++D F W E + AF +LKK ++ PV+ P
Sbjct: 2488 FLGLTGYYRRFIKSYANIAGPLTDLL---QKDSFLWNNEAEAAFVKLKKAMTEAPVLSLP 2658
Query: 1369 IKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFS 1428
+P L A+ +G++L Q N I Y S+ L + + + L + +
Sbjct: 2659 DFSQPFILETDASGIGVGAVLGQ-----NGHPIAYFSKKLAPRMQKQSAYTRELLAITEA 2823
Query: 1429 CVKLKYYI 1436
K ++Y+
Sbjct: 2824 LSKFRHYL 2847
Score = 107 bits (267), Expect = 5e-23
Identities = 97/341 (28%), Positives = 150/341 (43%), Gaps = 9/341 (2%)
Frame = +1
Query: 1730 HDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSII 1789
H G H AG + YWP + +D Y + C CQ+ +PA L +
Sbjct: 3235 HSSPIGGH-AGITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL- 3408
Query: 1790 KPWPFRGW---AIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQ-NVTQETVIEFIQ 1845
P P + W A+D I + +S I+V +D TK+ IPL+ + + V E
Sbjct: 3409 -PIPQQVWEDVAMDFITGL--PNSFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFM 3579
Query: 1846 NHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILIS 1905
+HIV G+P SI +D+ VF + G L S+ Y+ Q++GQ E NK L
Sbjct: 3580 SHIVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEM 3759
Query: 1906 LIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQR 1965
++ PK W ++L + Y + + G TPFR YG+E P + Q+C I
Sbjct: 3760 YLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGRE---PPTLTRQACSIDD 3930
Query: 1966 QEEIPSEVYWNMMLDEMVNLDEERLLA-LDV-LTRQKDRIAKAYNKKVRDRSFITGDYVW 2023
E+ ++ D + LLA L + LTR + + + +KK D SF GD V
Sbjct: 3931 PAEVREQL-----------TDRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVL 4077
Query: 2024 KVILPMDKKDRVY---GKWTPNWEGPFTVEKTLPNNAYVIK 2061
+ P + V K + + GPF V + + AY ++
Sbjct: 4078 VKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLE 4200
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 156 bits (394), Expect = 1e-37
Identities = 79/223 (35%), Positives = 117/223 (52%)
Frame = +1
Query: 1677 EYLQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGA 1736
EYL + R ++ A + + G+ L+K+N + L+C+ +A + H+G G
Sbjct: 175 EYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGT 354
Query: 1737 HQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRG 1796
H G M + R G YW T+ DC + R C CQ + + P L+ + PWPF
Sbjct: 355 HANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSM 534
Query: 1797 WAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPE 1856
W ID+IG I P +S H++I+VA+DYFTKWVEA +V + V+ FI+ I+ R+GLP
Sbjct: 535 WGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPR 714
Query: 1857 SITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAA 1899
I TD GT + + E + I+ TPY + N VE A
Sbjct: 715 KIITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 151 bits (381), Expect = 3e-36
Identities = 94/260 (36%), Positives = 131/260 (50%), Gaps = 18/260 (6%)
Frame = +1
Query: 1084 IKEEIERLLKCKFIRTARYVDWLANVVPVIKKNG------------------KMRVCIDF 1125
+++E+ +LL+ I W++ V V KK G + R+CID+
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 1126 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGAL 1185
R LN AT KD Y +P + M+ A + LDGYSGYNQI + +D KTAF CP ++
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 1186 GTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLH 1245
Y MPFGL NA T+QR M IF D +E ++V++DD S L +
Sbjct: 361 FAYRR--MPFGLCNAS-----TTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLAN 519
Query: 1246 LRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQ 1305
L K +R K L +N KC F V G LG + K+GIE+ K K I PP + K
Sbjct: 520 LEKVLQRCEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKG 699
Query: 1306 LQSLLGKINFLRRFITNLSE 1325
+ S LG + F RRFI + ++
Sbjct: 700 IHSFLGHVGFYRRFIKDFTK 759
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 143 bits (361), Expect = 7e-34
Identities = 88/260 (33%), Positives = 133/260 (50%), Gaps = 18/260 (6%)
Frame = +1
Query: 1084 IKEEIERLLKCKFIRTARYVDWLANVVPVIKKNG------------------KMRVCIDF 1125
+++E+ +LL+ I W++ V+ V KK G ++CID+
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 1126 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGAL 1185
R LN AT KD + +P + M++ AGH Y LD Y GYNQI + +D K AF CP
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCP--F 354
Query: 1186 GTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLH 1245
G + + +PFGL NA T+Q M IF D +E ++V++DD V PS + L
Sbjct: 355 GVFAYRRIPFGLCNAP-----TTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKK 519
Query: 1246 LRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQ 1305
L +R + L +N KC F V G LG + +GIE+++ K I PP++ K
Sbjct: 520 LEMVLQRCVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKG 699
Query: 1306 LQSLLGKINFLRRFITNLSE 1325
++S LG+ F RRFI + ++
Sbjct: 700 IRSFLGQARFYRRFIKDFTK 759
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 136 bits (343), Expect = 8e-32
Identities = 71/139 (51%), Positives = 94/139 (67%)
Frame = +2
Query: 1156 SLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNT 1215
S +DG+SGYNQI +A EDV KT F GT+ + VM FGLKN G ATYQR M
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVT--LWGTFSYRVMAFGLKNTG-----ATYQRAMVA 163
Query: 1216 IFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFL 1275
+FHD + ++VY+DD++ KS + HL++L K F R++KY LK+NP KC FGV +G L
Sbjct: 164 LFHDMMHKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLL 343
Query: 1276 GFVVHKKGIEINKNKAKAI 1294
GF+V +KGIEI+ K KA+
Sbjct: 344 GFIVSQKGIEIDPEKVKAL 400
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 129 bits (323), Expect = 2e-29
Identities = 63/150 (42%), Positives = 97/150 (64%)
Frame = +3
Query: 1120 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAF 1179
R+C+D+RDLN A+PKD + +P ++++ + A S +DG+SGYNQI +A ED+ KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
GT+ + VM FGLKN G ATY R M +F D + ++ Y+D+++ KS
Sbjct: 183 IT--LWGTFCYKVMSFGLKNFG-----ATYHRAMVALFQDMMHKEIEAYVDEMIAKSRME 341
Query: 1240 DGHLLHLRKSFERMRKYGLKMNPLKCAFGV 1269
+ HL++L+ F ++RKY L++NP KC FG+
Sbjct: 342 EEHLVNLQNLFGQLRKYRLRLNPRKCVFGL 431
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 105 bits (261), Expect = 3e-22
Identities = 66/184 (35%), Positives = 94/184 (50%)
Frame = +1
Query: 1897 EAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEV 1956
EAANK + +I+K + K WHE L L YR S +TG TPF L YG EAVLP EV
Sbjct: 1 EAANKNIKKIIQK-MTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 1957 YLQSCRIQRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSF 2016
+ S RI + + + D++ ++ +RL A+ + R+ A++KKV R F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 2017 ITGDYVWKVILPMDKKDRVYGKWTPNWEGPFTVEKTLPNNAYVIKELGNQRQCVTINGKY 2076
GD V K + K R GKW PN+EGPF V++ A V+ + + +N
Sbjct: 358 HEGDLVLKKMSHAVKDHR--GKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDV 531
Query: 2077 LKAY 2080
+K Y
Sbjct: 532 VKRY 543
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 102 bits (255), Expect = 1e-21
Identities = 55/156 (35%), Positives = 90/156 (57%), Gaps = 1/156 (0%)
Frame = +3
Query: 958 EQSVDECEFEDLRNLRLDCIYDDGPLGFEKPVVDVSKKMEA-QDPLEEVDLGDGLEKRPT 1016
EQ +++ E E ++ L P E+ V ++M Q+ E VDLG G KR
Sbjct: 870 EQKMNQTEDEGNEDVGL-------PPELERMVAHEDQEMGPHQEETELVDLGSGSGKREV 1028
Query: 1017 YISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKKPVKQLPRRF 1076
I I +++ ++ LL++++D FAW Y +MPGL+ ++V+ +LP+ + PVKQ RR
Sbjct: 1029 KIGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQHRLPLNPECSPVKQKLRRM 1208
Query: 1077 HPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPV 1112
P+ +KIKEE+++ F+ ARY W+AN+VP+
Sbjct: 1209 KPETSLKIKEEVKK*FDAGFLAVARYPKWVANIVPI 1316
>BI316922
Length = 405
Score = 101 bits (252), Expect = 3e-21
Identities = 48/132 (36%), Positives = 75/132 (56%)
Frame = +3
Query: 1730 HDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSII 1789
H G+CG H M + R G Y T+ K C EY + C++C K I H+ ELH+I+
Sbjct: 9 HRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHNIV 188
Query: 1790 KPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIV 1849
PWPF +D++ P S +Q KY++V +D FTKW+E + ++ V +F+ +IV
Sbjct: 189 APWPFAI*GVDILRPF-PLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNIV 365
Query: 1850 YRFGLPESITTD 1861
FG+P ++ +D
Sbjct: 366 C*FGIPNTLISD 401
>AW184779
Length = 432
Score = 59.7 bits (143), Expect(2) = 1e-16
Identities = 25/45 (55%), Positives = 33/45 (72%)
Frame = +3
Query: 1797 WAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETVI 1841
W ID+IG I P +S H +I+VA+DYFTKWVEA+ +VT+ VI
Sbjct: 294 WGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVI 428
Score = 47.4 bits (111), Expect(2) = 1e-16
Identities = 23/71 (32%), Positives = 34/71 (47%)
Frame = +1
Query: 1702 NELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDC 1761
N L+K+N + LL+C+ +A + H+G G H M + R G YW T+ DC
Sbjct: 10 NILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDC 189
Query: 1762 IEYARGCQDCQ 1772
+ C CQ
Sbjct: 190 CIHVWKCHKCQ 222
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 65.5 bits (158), Expect(2) = 1e-16
Identities = 50/191 (26%), Positives = 87/191 (45%), Gaps = 6/191 (3%)
Frame = +2
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
K + + GDS LV+ QL E + +L Y L F+EI HV ENQ A+ L
Sbjct: 233 KLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA*EENQMADAL 412
Query: 1630 AQIASGYMVDKLKLTELIKIKEKLSPSDLDIMCIDNLTSNDWRKPIVEYLQN---PVGST 1686
A + S + + I+ + + P+ ++ + W I Y+++ P+ ++
Sbjct: 413 ATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLV-EEERDGKPWYFDIKRYVESKEYPLEAS 589
Query: 1687 D---RKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKM 1743
D R+ + A + + G+ L+K+N + LL C++ + + H+G G H G M
Sbjct: 590 DNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFGTHSNGHAM 769
Query: 1744 KWILFRQGLYW 1754
+ R G YW
Sbjct: 770 ARKILRAGYYW 802
Score = 41.2 bits (95), Expect(2) = 1e-16
Identities = 18/47 (38%), Positives = 28/47 (59%)
Frame = +1
Query: 1510 WKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEA 1556
W ++FD +S+ G G+G ++SP F R+ +C+NN EYEA
Sbjct: 52 WIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEA 192
>BI321666
Length = 430
Score = 77.8 bits (190), Expect = 5e-14
Identities = 41/122 (33%), Positives = 64/122 (51%)
Frame = +2
Query: 1753 YWPTIMKDCIEYARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQ 1812
Y P+I KD +A+ C CQ+ + LH+I++ F W ID +G P+ S +
Sbjct: 32 YLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVGPFPPSFSNE 211
Query: 1813 HKYIIVAVDYFTKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVA 1872
YI+V VDY +KWVEA+ Q + VI+F++ I R G+P + + G+ +A
Sbjct: 212 --YILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGSHLCNA*LA 385
Query: 1873 AF 1874
F
Sbjct: 386 RF 391
>BQ628592
Length = 423
Score = 72.4 bits (176), Expect = 2e-12
Identities = 31/72 (43%), Positives = 51/72 (70%), Gaps = 1/72 (1%)
Frame = -1
Query: 1336 LKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDED 1395
L K W + +Q+AF+++K+ L+NP V++PP+ GRP LY++ DE++G +L Q D+
Sbjct: 423 LPKNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
Query: 1396 G-NERAIFYLSR 1406
G E+AI+YLS+
Sbjct: 243 GKKEQAIYYLSK 208
>BE802896
Length = 416
Score = 66.2 bits (160), Expect = 1e-10
Identities = 46/137 (33%), Positives = 71/137 (51%)
Frame = -2
Query: 1308 SLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIP 1367
S LG F RRFI + + S+LL+ KE F + + + AFD LK+ L P++
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQ--KEVEFDFNDKCK*AFDCLKRALITTPIIQA 242
Query: 1368 PIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYF 1427
P P +L A++ +G +LAQ+ D R I+Y SR L+ A+ YT EK L + F
Sbjct: 241 PDWTAPFELMCDASNYALGVVLAQK-IDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVF 65
Query: 1428 SCVKLKYYIKPIDVMVF 1444
+ K Y+ ++V+
Sbjct: 64 ALEKFHSYLLGTRIIVY 14
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 56.2 bits (134), Expect(2) = 2e-10
Identities = 43/169 (25%), Positives = 76/169 (44%), Gaps = 2/169 (1%)
Frame = +1
Query: 1300 PTSKK--QLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKK 1357
PT K ++S G +F RRF+ N S + L+ KK F W + ++AF LK+
Sbjct: 97 PTLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELV--KKNMAFTWGEKQEQAFALLKE 270
Query: 1358 YLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTM 1417
L+ PV+ P + +L A+ + ++L Q I Y S L+ A Y
Sbjct: 271 KLTKAPVLALPDFSKTFELECDASGVGVRAVLLQ-----GGHPIAYFSEKLHSATLNYPT 435
Query: 1418 IEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKW 1466
+K L + ++++ + ++ S + +K++ K L+ R KW
Sbjct: 436 YDKELYALIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKW 582
Score = 29.3 bits (64), Expect(2) = 2e-10
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = +2
Query: 1274 FLGFVVHKKGIEINKNKAKAILDTSPP 1300
F GFVV + G++++ K KAI + PP
Sbjct: 23 FSGFVVGRNGVQMDPEKIKAIQEWPPP 103
>BG839293
Length = 781
Score = 65.5 bits (158), Expect = 2e-10
Identities = 30/73 (41%), Positives = 47/73 (64%)
Frame = +1
Query: 999 QDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVEL 1058
Q+ E VDLG G KR I I +++ ++ LLK+++D FAW Y +MPGL+ ++V+
Sbjct: 511 QEETELVDLGIGSGKREVKIGTGITAPIREELIILLKDYQDIFAWSYQDMPGLSSDIVQH 690
Query: 1059 KLPIKEDKKPVKQ 1071
+LP+ + PVKQ
Sbjct: 691 QLPLNPECSPVKQ 729
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 62.0 bits (149), Expect = 3e-09
Identities = 28/84 (33%), Positives = 46/84 (54%)
Frame = -3
Query: 1873 AFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNS 1932
A + +G+ STPY+ Q NGQ E +N+ + +++K V K W L LWA+R +
Sbjct: 370 ALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTA 191
Query: 1933 PTEATGTTPFRLAYGQEAVLPAEV 1956
G +P+R+ +G+ LP E+
Sbjct: 190 YKAPIGMSPYRVVFGKACHLPVEI 119
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 61.6 bits (148), Expect = 3e-09
Identities = 36/89 (40%), Positives = 47/89 (52%)
Frame = -3
Query: 1059 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGK 1118
KL I D K V Q R+ + + +E+ +L FIR Y L +VV V K NGK
Sbjct: 274 KLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNGK 95
Query: 1119 MRVCIDFRDLNAATPKDEYHMPIAEMMVD 1147
R+C D+ DLN A PKD Y +P + M D
Sbjct: 94 WRICTDYIDLN*ACPKDAYPLPNIDHMTD 8
>BI317507
Length = 359
Score = 58.5 bits (140), Expect = 3e-08
Identities = 32/114 (28%), Positives = 58/114 (50%)
Frame = -1
Query: 1231 DIVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNK 1290
+I++ SP H++HL + ++K L N KC F ++LG V+ K + ++ NK
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 1291 AKAILDTSPPTSKKQLQSLLGKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRW 1344
K++++ P + K++ S L + R+FI + + L L K D F+W
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAP--RPLTDLTKNDGFKW 18
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.333 0.143 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 91,188,445
Number of Sequences: 63676
Number of extensions: 1300932
Number of successful extensions: 9781
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 9500
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9759
length of query: 2087
length of database: 12,639,632
effective HSP length: 112
effective length of query: 1975
effective length of database: 5,507,920
effective search space: 10878142000
effective search space used: 10878142000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0545.3


