

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
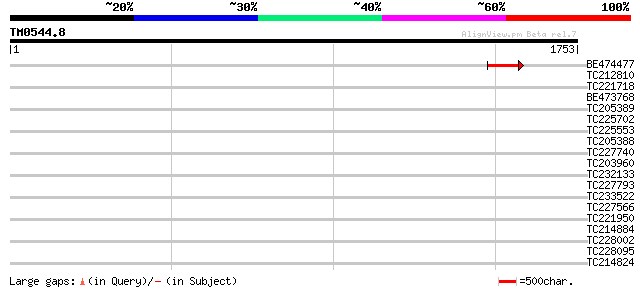
Query= TM0544.8
(1753 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE474477 138 2e-32
TC212810 similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD12035p), pa... 36 0.17
TC221718 35 0.29
BE473768 34 0.64
TC205389 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2... 33 0.84
TC225702 UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (98%) 33 1.1
TC225553 weakly similar to UP|Q06106 (Q06106) Ypr112cp, partial ... 32 3.2
TC205388 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partia... 31 5.4
TC227740 similar to UP|Q8LBD3 (Q8LBD3) Cytochrome-b5 reductase-l... 30 7.1
TC203960 UP|EF1A_SOYBN (P25698) Elongation factor 1-alpha (EF-1-... 30 7.1
TC232133 weakly similar to UP|Q9V5S5 (Q9V5S5) CG33473-PB (Transc... 30 7.1
TC227793 30 7.1
TC233522 30 7.1
TC227566 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (7%) 30 7.1
TC221950 similar to UP|Q9FL07 (Q9FL07) RING zinc finger protein-... 30 9.3
TC214884 similar to UP|P93169 (P93169) Early light-induced prote... 30 9.3
TC228002 30 9.3
TC228095 similar to GB|AAO42974.1|28416887|BT004728 At3g53470 {A... 30 9.3
TC214824 UP|Q9MAZ9 (Q9MAZ9) Nonclathrin coat protein zeta1-COP, ... 30 9.3
>BE474477
Length = 354
Score = 138 bits (348), Expect = 2e-32
Identities = 62/115 (53%), Positives = 84/115 (72%), Gaps = 2/115 (1%)
Frame = +3
Query: 1477 LFETTLCEIISWETLEARTSLDPKGYLSTYNEKDIQLICCQSDASTLWLVPPIVQARFMK 1536
L++TTLCE + W+ L + + DP GYL YN+ DIQLICCQ+DASTLWLVP +V+ R ++
Sbjct: 9 LYQTTLCERLQWDLLNSNINPDPYGYLGAYNKNDIQLICCQADASTLWLVPLVVRTRLIQ 188
Query: 1537 SLSWNMDLAF--SWEFTRDRPRGKEVVKYELTIQEQDLPKSSEVTEVFNGTSNSF 1589
SL WN+D+ +W +RDRP+GKE+VKYE + Q LP S+V +V NG+ NSF
Sbjct: 189 SLEWNIDMEIFSTWILSRDRPKGKEIVKYEKAVDPQYLPTRSDVQKVLNGSMNSF 353
>TC212810 similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD12035p), partial (11%)
Length = 407
Score = 35.8 bits (81), Expect = 0.17
Identities = 20/70 (28%), Positives = 35/70 (49%)
Frame = -1
Query: 236 PNETTACTPQNKQAETSTSATIKRLARSRSCPTVNSALSQGTDSGIEGDSTKKLRQLHFW 295
P +TTA ++Q +S S ++ A RS + S +E S++K + FW
Sbjct: 284 PIKTTALNSASQQRNSSRS--LRSHAMERSASPEEEVEEEEESSEVEDKSSEKRSPMCFW 111
Query: 296 ESSKDSLKWN 305
+SS+ S+ W+
Sbjct: 110 KSSRSSISWS 81
>TC221718
Length = 860
Score = 35.0 bits (79), Expect = 0.29
Identities = 15/29 (51%), Positives = 19/29 (64%)
Frame = -3
Query: 889 VRHIWSRMRSNNEVVCYCCFILIYLWNFS 917
+R +WSR NN ++ CCFIL LWN S
Sbjct: 102 MRRVWSRREINNYIIL-CCFILQSLWNAS 19
>BE473768
Length = 431
Score = 33.9 bits (76), Expect = 0.64
Identities = 15/47 (31%), Positives = 28/47 (58%)
Frame = -3
Query: 1255 FVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLVLFTYSVTKYAW 1301
F + +++ F L R+ LC ++ A+ IF++F ++L +S T Y W
Sbjct: 147 FSVYILLYFCLYFCSRLSLLCLYS-ARFIFFIFLILLLCFSYTYYPW 10
>TC205389 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2O10_3
{Arabidopsis thaliana;} , partial (16%)
Length = 538
Score = 33.5 bits (75), Expect = 0.84
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +2
Query: 905 YCCFILIYLWNFSLLSVVYLAALFLYALCQN-TGPSYIFWVLILIYTEVCILLQYLYQI 962
YC F +W F +LS+ + F+Y N G Y +W+ + + VC+L+ + +I
Sbjct: 338 YCLF----MWQFDILSISSV*LPFVYLTFINCNGNKYHYWLRLQLIRSVCLLISHFNKI 502
>TC225702 UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (98%)
Length = 1431
Score = 33.1 bits (74), Expect = 1.1
Identities = 15/38 (39%), Positives = 23/38 (60%)
Frame = -2
Query: 1255 FVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLVLF 1292
F+ +L V FFL +L +L + A A F+LF +V+F
Sbjct: 788 FLFLLAVAFFLFLLTTAFFLLAAAAAAAAFFLFLIVVF 675
>TC225553 weakly similar to UP|Q06106 (Q06106) Ypr112cp, partial (6%)
Length = 1371
Score = 31.6 bits (70), Expect = 3.2
Identities = 23/81 (28%), Positives = 41/81 (50%)
Frame = +1
Query: 286 TKKLRQLHFWESSKDSLKWNRKRLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINM 345
+KK +Q+ F + K +K+ R + K+ + L++S KFW N F + INM
Sbjct: 1111 SKKRKQVDFLDEGK--MKFGRMAD*YSSKDL----NSILRMSWKFWDHNFVIGFQVYINM 1272
Query: 346 IALLLASFAVLNAISLLYIAS 366
L+S + ++SL + +S
Sbjct: 1273 PCCFLSSSVKITSLSLGFYSS 1335
>TC205388 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partial (86%)
Length = 1433
Score = 30.8 bits (68), Expect = 5.4
Identities = 16/53 (30%), Positives = 27/53 (50%), Gaps = 1/53 (1%)
Frame = +1
Query: 905 YCCFILIYLWNFSLLSVVYLAALFLYALCQN-TGPSYIFWVLILIYTEVCILL 956
YC F +W F +LS+ L F+Y N G Y W+ + ++ +C+L+
Sbjct: 1279 YCLF----MW*FDILSI*SLLLPFVYLTFINCNGNKYHSWLRLQLFRSICLLI 1425
>TC227740 similar to UP|Q8LBD3 (Q8LBD3) Cytochrome-b5 reductase-like protein,
partial (78%)
Length = 1368
Score = 30.4 bits (67), Expect = 7.1
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 1066 CRYWKSLTWGAETPPYFVQLSMEVNLWPKEGI 1097
C+ W S+ PPYF+ S+ VN P +G+
Sbjct: 1259 CQQWASVGVNLLLPPYFLTFSLRVNYSPLKGL 1354
>TC203960 UP|EF1A_SOYBN (P25698) Elongation factor 1-alpha (EF-1-alpha),
partial (31%)
Length = 699
Score = 30.4 bits (67), Expect = 7.1
Identities = 21/66 (31%), Positives = 30/66 (44%), Gaps = 3/66 (4%)
Frame = +1
Query: 891 HIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVY---LAALFLYALCQNTGPSYIFWVLIL 947
+I+ + + CCF L+ N + L VV+ L LFLY LC N G L+
Sbjct: 481 YIFVKRSEFQDFAALCCFGLL---NVNFLFVVFNFWLGLLFLYNLCSNFG-------LVC 630
Query: 948 IYTEVC 953
Y +C
Sbjct: 631 FYYNLC 648
>TC232133 weakly similar to UP|Q9V5S5 (Q9V5S5) CG33473-PB (Transcription
factor DKLF) (HL07808p), partial (5%)
Length = 461
Score = 30.4 bits (67), Expect = 7.1
Identities = 17/50 (34%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Frame = +1
Query: 817 QSLGNMAVNNLVNYLKIEREG--QEP---TDDSSEDEEYYEIENQNSGAE 861
+S+G + V+Y + E+ G ++P +D +DEEY E+E++N G E
Sbjct: 304 RSIGAIKSYKDVSYFQTEQVGSWEDPDTGSDSECDDEEYEEVEDENFGFE 453
>TC227793
Length = 444
Score = 30.4 bits (67), Expect = 7.1
Identities = 25/83 (30%), Positives = 38/83 (45%), Gaps = 5/83 (6%)
Frame = +3
Query: 895 RMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLI-----LIY 949
RM S + C C F I F +L +++L F+ YI+ +L+ +IY
Sbjct: 210 RMASQELLACQCGFYCITF--FFILDILFLFFNFV--------SWYIYLILV*NYCVIIY 359
Query: 950 TEVCILLQYLYQIIIQHTEFEFP 972
IL +LY+II + E FP
Sbjct: 360 IYYMILSFHLYEIIQVNKEIGFP 428
>TC233522
Length = 800
Score = 30.4 bits (67), Expect = 7.1
Identities = 15/44 (34%), Positives = 26/44 (59%), Gaps = 4/44 (9%)
Frame = +3
Query: 894 SRMRSNNEVVCYCCFILIYLWN----FSLLSVVYLAALFLYALC 933
S M + +EV C C ++++ LWN + S VY +L+L ++C
Sbjct: 492 SMMMNISEVECVCWYVVLCLWNKI*ISTASSYVYERSLWLCSIC 623
>TC227566 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (7%)
Length = 819
Score = 30.4 bits (67), Expect = 7.1
Identities = 19/47 (40%), Positives = 24/47 (50%)
Frame = +3
Query: 593 INELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFDYVS 639
+ E G ++ Y F SR L + Y S+ SYM FPEF YVS
Sbjct: 636 LRERGGRKR*AYCFHFRSRRLL*SLPYYKAYSIDSYMMLFPEF-YVS 773
>TC221950 similar to UP|Q9FL07 (Q9FL07) RING zinc finger protein-like,
partial (11%)
Length = 550
Score = 30.0 bits (66), Expect = 9.3
Identities = 23/59 (38%), Positives = 27/59 (44%), Gaps = 3/59 (5%)
Frame = +2
Query: 79 EVGPFPFTRRLLVRHTEK---ILFLALFYASLSPISAFGFLYLLGLINCSRLPKSSQIP 134
EVG FPF + TEK +L L +F+AS SP S F P SS IP
Sbjct: 311 EVGNFPFFGHNQILDTEKLSPLLTLLIFFASTSPSSTETF------------PSSSSIP 451
>TC214884 similar to UP|P93169 (P93169) Early light-induced protein, partial
(98%)
Length = 2021
Score = 30.0 bits (66), Expect = 9.3
Identities = 15/49 (30%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Frame = -3
Query: 1396 IHASLFLVKCDVDLNRASHQQGQKQTKMTKF--CNGICLFFVLMCVIWA 1442
+HA +FL+ C + L+R + +F G C FF+++ +IWA
Sbjct: 216 LHAYIFLLNCIISLSRCCVMS----LSLFQFV*AMGRCFFFIILILIWA 82
>TC228002
Length = 805
Score = 30.0 bits (66), Expect = 9.3
Identities = 17/64 (26%), Positives = 34/64 (52%)
Frame = +1
Query: 654 QEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKND 713
Q ++ W T + + ELK +++V+++++E +L+ L D ASL+N
Sbjct: 316 QSWRSKWDTQVKVYRDELSELKKTLNVEVDQLRAEFQDLRTTLQQQQEDV---TASLRNL 486
Query: 714 GLRE 717
GL++
Sbjct: 487 GLQD 498
>TC228095 similar to GB|AAO42974.1|28416887|BT004728 At3g53470 {Arabidopsis
thaliana;} , partial (50%)
Length = 918
Score = 30.0 bits (66), Expect = 9.3
Identities = 19/57 (33%), Positives = 27/57 (47%)
Frame = +1
Query: 294 FWESSKDSLKWNRKRLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLL 350
F + S + WN +R F E ++ K + +L FW FTL G I + LLL
Sbjct: 286 FSKFSDKYVSWNPRRSEFPLSEEVDPIKRTERSNLSFWNSPTFTLGGAIIIVTFLLL 456
>TC214824 UP|Q9MAZ9 (Q9MAZ9) Nonclathrin coat protein zeta1-COP, partial (68%)
Length = 1686
Score = 30.0 bits (66), Expect = 9.3
Identities = 19/80 (23%), Positives = 41/80 (50%), Gaps = 3/80 (3%)
Frame = -3
Query: 332 IENMFTLFGLEINMIALLLASFAVLNAISLLYIASLAACVLLH---RLLIKKLWPIFVFL 388
+ + + LEI M+A A +++ LY+AS +C + H R L+ + F F+
Sbjct: 1357 VARLLCIESLEIPMLAESCGEGATTHSVLCLYVAS--SCRIRHEGERELVSSFFFFFFFM 1184
Query: 389 FASVVTVEYLAIWMRLTFMN 408
+ ++ E ++ + +T++N
Sbjct: 1183 SSQILIDERISRYWIVTYLN 1124
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 83,979,105
Number of Sequences: 63676
Number of extensions: 1317497
Number of successful extensions: 10473
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 10209
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10465
length of query: 1753
length of database: 12,639,632
effective HSP length: 111
effective length of query: 1642
effective length of database: 5,571,596
effective search space: 9148560632
effective search space used: 9148560632
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0544.8


