

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.15
(322 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
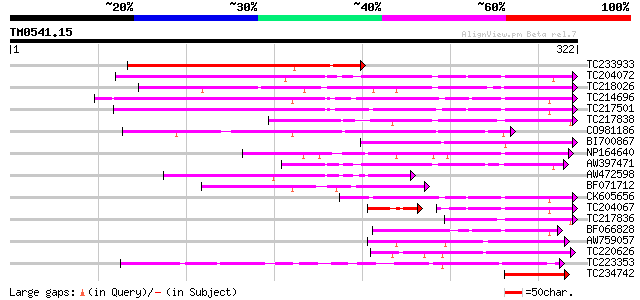
Score E
Sequences producing significant alignments: (bits) Value
TC233933 similar to UP|Q43563 (Q43563) Lectin precursor, partial... 166 1e-41
TC204072 UP|LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SB... 166 2e-41
TC218026 weakly similar to UP|Q9ZTA9 (Q9ZTA9) Mannose lectin, pa... 164 4e-41
TC214696 similar to UP|Q8L683 (Q8L683) Lectin precursor, partial... 144 7e-35
TC217501 similar to UP|Q703U4 (Q703U4) Lectin precursor, partial... 143 1e-34
TC217838 weakly similar to PIR|A03357|CVJBP concanavalin A precu... 117 7e-27
CO981186 96 2e-20
BI700867 similar to GP|1755066|gb|A lectin precursor {Sophora ja... 79 2e-15
NP164640 receptor-like protein kinase 72 3e-13
AW397471 similar to GP|3114258|pdb Soybean Agglutinin Complexed ... 69 4e-12
AW472598 similar to GP|3114258|pdb Soybean Agglutinin Complexed ... 67 1e-11
BF071712 weakly similar to GP|22208832|em lectin {Vigna linearis... 63 2e-10
CK605656 63 2e-10
TC204067 lectin [Glycine max] 49 4e-10
TC217836 similar to UP|Q9M7M4 (Q9M7M4) Mannose lectin FRIL (Frag... 57 9e-09
BF066828 weakly similar to PIR|A31972|A319 lectin DB58 precursor... 55 6e-08
AW759057 weakly similar to GP|6650223|gb| receptor-like protein ... 48 7e-06
TC220626 similar to UP|Q9M2S4 (Q9M2S4) Probable serine/threonine... 42 5e-04
TC223353 weakly similar to UP|Q9SEW3 (Q9SEW3) Receptor-like prot... 42 5e-04
TC234742 similar to UP|Q6UY58 (Q6UY58) Lectin-like receptor kina... 41 8e-04
>TC233933 similar to UP|Q43563 (Q43563) Lectin precursor, partial (7%)
Length = 422
Score = 166 bits (420), Expect = 1e-41
Identities = 83/137 (60%), Positives = 105/137 (76%), Gaps = 2/137 (1%)
Frame = +3
Query: 68 ATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPH 127
+ + + LL+ T+ KSDS SF+ P F+PD ILL+G A TT G LQLTK D+ GNP+ H
Sbjct: 15 SVFLMTFLLLITSAKSDSFSFNLPRFEPDALNILLDGSAKTTGGVLQLTKKDKRGNPTQH 194
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPEGGG 185
S G+SAFY A+HLSD KTGRVA+F TEF+FVVN + HGDGFTF+LASLDF+FP+
Sbjct: 195 SVGLSAFYAALHLSDAKTGRVANFATEFSFVVNTKGAPLHGDGFTFYLASLDFDFPD-NS 371
Query: 186 GGGFLGLFDEETAYNTS 202
GGFLGLF+++TA+NTS
Sbjct: 372 SGGFLGLFNKKTAFNTS 422
>TC204072 UP|LEC_SOYBN (P05046) Lectin precursor (Agglutinin) (SBA), complete
Length = 1086
Score = 166 bits (419), Expect = 2e-41
Identities = 110/268 (41%), Positives = 154/268 (57%), Gaps = 6/268 (2%)
Frame = +1
Query: 61 NITSELLATLIFSLLLITTTVKS-DSLSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKI 118
N+ L TL L+L+T+ S +++SFS+ F P ++L GDA T++G LQL K+
Sbjct: 151 NVVVSLSLTLTLVLVLLTSKANSAETVSFSWNKFVPKQPNMILQGDAIVTSSGKLQLNKV 330
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEF--TFVVNHSQPHGDGFTFFLASL 176
D+ G P P S G + + +H+ DK+TG VA F F TF ++ DG FFLA +
Sbjct: 331 DENGTPKPSSLGRALYSTPIHIWDKETGSVASFAASFNFTFYAPDTKRLADGLAFFLAPI 510
Query: 177 DFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAV 236
D + P+ G +LGLF+E N S + +VAVEFD+F NS+DP PHIGI++N+IRS
Sbjct: 511 DTK-PQTHAG--YLGLFNE----NESGDQVVAVEFDTFRNSWDPPNPHIGINVNSIRSIK 669
Query: 237 TIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWV 296
T W D + I+Y + S L V P Q T+NI + +D+++ L EWV
Sbjct: 670 TTSW---DLANNKVAKVLITYDA-STSLLVASLVYPSQRTSNI--LSDVVDLKTSLPEWV 831
Query: 297 VIGFSATTGEF--AETHSILSWSFNSTL 322
IGFSA TG E+H +LSWSF S L
Sbjct: 832 RIGFSAATGLDIPGESHDVLSWSFASNL 915
>TC218026 weakly similar to UP|Q9ZTA9 (Q9ZTA9) Mannose lectin, partial (72%)
Length = 1102
Score = 164 bits (416), Expect = 4e-41
Identities = 112/278 (40%), Positives = 150/278 (53%), Gaps = 29/278 (10%)
Frame = +3
Query: 74 LLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANT------TNGTLQLTKIDQYGNPSPH 127
L+L+ +DSLSFSF F D ++L GDA T LQLTK+D G P
Sbjct: 102 LMLLNRVNSADSLSFSFNNFSEDQEDLILQGDATTGASSENDKNVLQLTKLDDSGKPEFG 281
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG---DGFTFFLASLDFEFPEGG 184
S G ++ VHL K + V+ F T FTF ++ + P DG FF+AS D G
Sbjct: 282 SVGRVLYFAPVHLW-KSSQLVSTFETTFTFKISSASPDSVPADGLAFFIASPD---TTPG 449
Query: 185 GGGGFLGLFDEETAYNTSKNS------------------IVAVEFDSFSNSF--DPNFPH 224
GG LGLF T+ S +S +VAVEFD+F N+ DP + H
Sbjct: 450 AGGQDLGLFPHLTSLKNSSSSHHRKVTRITGVKDLASEPVVAVEFDTFINTDIGDPEYQH 629
Query: 225 IGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKY 284
IGIDIN+I S T W D G A+ISY+S S+ LTVV YP ++T + ++ Y
Sbjct: 630 IGIDINSITSVTTTKW---DWQNGKTVTAQISYNSASKRLTVVASYP---DSTPV-SLYY 788
Query: 285 PIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
ID+ + L EWV +GFSA+TG AE +++LSWSF+S+L
Sbjct: 789 DIDLFTILPEWVRVGFSASTGGAAEANTLLSWSFSSSL 902
>TC214696 similar to UP|Q8L683 (Q8L683) Lectin precursor, partial (80%)
Length = 1701
Score = 144 bits (362), Expect = 7e-35
Identities = 98/281 (34%), Positives = 157/281 (54%), Gaps = 7/281 (2%)
Frame = +3
Query: 49 QYSQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDAN- 107
++ Q+ T +N + L +L F L+L+T +D++SF+F F+P I+L DA+
Sbjct: 18 EFKQIKAMAT-SNFSIVLSVSLAFFLVLLTKAHSTDTVSFTFNKFNPVQPNIMLQKDASI 194
Query: 108 TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV--NHSQPH 165
+++G LQLTK+ G P+ S G + + + + D +TG+VA + T F F + +
Sbjct: 195 SSSGVLQLTKVGSNGVPTSGSLGRALYAAPIQIWDSETGKVASWATSFKFNIFAPNKSNS 374
Query: 166 GDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPH 224
DG FFLA + + P+ GFLGLF+ + VA+EFD+FSN +DP H
Sbjct: 375 ADGLAFFLAPVGSQ-PQ--SDDGFLGLFN--SPLKDKSLQTVAIEFDTFSNKKWDPANRH 539
Query: 225 IGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKY 284
IGID+N+I+S T W +++ G + ++Y++ + L V P + T+ I +
Sbjct: 540 IGIDVNSIKSVKTASWGLSN---GQVAEILVTYNAAT-SLLVASLIHPSKKTSYI--LSD 701
Query: 285 PIDIESYLSEWVVIGFSATTG---EFAETHSILSWSFNSTL 322
++++S L EWV +GFSATTG ETH ++SWSF S L
Sbjct: 702 TVNLKSNLPEWVSVGFSATTGLHEGSVETHDVISWSFASKL 824
>TC217501 similar to UP|Q703U4 (Q703U4) Lectin precursor, partial (79%)
Length = 1011
Score = 143 bits (360), Expect = 1e-34
Identities = 97/268 (36%), Positives = 147/268 (54%), Gaps = 5/268 (1%)
Frame = +2
Query: 60 ANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKI 118
+N + L +L F L+L+T ++++SF+ F P + ++ GDA + +G L+LTK+
Sbjct: 29 SNFSIVLSLSLAFFLVLLTKANSTNTVSFTVSKFSPRQQNLIFQGDAAISPSGVLRLTKV 208
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHGDGFTFFLASLDF 178
D P+ S G + + + + D +TG+VA + T F F V DG FFLA +
Sbjct: 209 DSIDVPTTGSLGRALYATPIQIWDSETGKVASWATSFKFKVFSPNKTADGLAFFLAPVGS 388
Query: 179 EFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNS-FDPNFPHIGIDINTIRSAVT 237
+ P+ GGFLGLF+ ++ + + VAVEFD++ N+ +DP HIGID+N+I+S T
Sbjct: 389 K-PQ--SKGGFLGLFNSDSKNKSVQT--VAVEFDTYYNAKWDPANRHIGIDVNSIKSVKT 553
Query: 238 IPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVV 297
W + G I I+Y + L V P + T+ I + + ++S L EWV
Sbjct: 554 ASWGL---ANGQIAQILITYDA-DTSLLVASLIHPSRKTSYI--LSETVSLKSNLPEWVN 715
Query: 298 IGFSATTG---EFAETHSILSWSFNSTL 322
IGFSATTG F ETH + SWSF S L
Sbjct: 716 IGFSATTGLNKGFVETHDVFSWSFASKL 799
>TC217838 weakly similar to PIR|A03357|CVJBP concanavalin A precursor - jack
bean {Canavalia ensiformis;} , partial (52%)
Length = 1202
Score = 117 bits (293), Expect = 7e-27
Identities = 72/180 (40%), Positives = 102/180 (56%), Gaps = 5/180 (2%)
Frame = +3
Query: 148 VADFRTEFTFVVNH-SQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSI 206
VA F T+FTF ++ S GDG FF+A D + P GG LGLF + ++
Sbjct: 18 VASFETDFTFSISSDSTTPGDGLAFFIAPFDTKIPPNSGGSN-LGLFPSD--------NV 170
Query: 207 VAVEFDSFSN--SFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDL 264
VAVEFD++ N DP++ HIGID+N+I S T W + G I ISY+S S+ L
Sbjct: 171 VAVEFDTYPNRDKGDPDYRHIGIDVNSIVSKATARW---EWQNGKIATVHISYNSASKRL 341
Query: 265 TVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWS--FNSTL 322
TV YP Q T + + I++ L EWV +G SA+TG+ +T++I SWS FN+++
Sbjct: 342 TVAAFYPGTQTVT----LSHDIELNKVLPEWVRVGLSASTGQQKQTNTIHSWSLAFNNSI 509
>CO981186
Length = 801
Score = 96.3 bits (238), Expect = 2e-20
Identities = 71/233 (30%), Positives = 120/233 (51%), Gaps = 10/233 (4%)
Frame = +1
Query: 65 ELLATLIFSLLLITTTVKS-DSLSFSFPTF--DPDVRVILLNGD-ANTTNGTLQLTKIDQ 120
E++ T+ +L I + +K+ +SL+F+ F + +L GD A NG+++L +D
Sbjct: 109 EMVITVFLLVLAIPSPLKTAESLNFNITNFANSESAKNMLYVGDGAVNKNGSIELNIVDY 288
Query: 121 YGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV----NHSQPHGDGFTFFLASL 176
G + + + L D +G V DF T FTF + N S + DGF F++A
Sbjct: 289 -----DFRVGRALYGQPLRLWDSSSGVVTDFSTRFTFTIDRGNNKSASYADGFAFYIAPH 453
Query: 177 DFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAV 236
++ P GG F LF+ + +N ++AVEFD+F+ + DP F H+GID N+++S
Sbjct: 454 GYQIPPNAAGGTF-ALFNVTSNPFIPRNHVLAVEFDTFNGTIDPPFQHVGIDDNSLKSVA 630
Query: 237 TIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNI--DAIKYPID 287
T + + D+ NA ++Y++ +R L V + G T N ++ Y ID
Sbjct: 631 TAKFDI-DKNLXXKCNALVNYNASNRTLFVSWSF-NGAATPNXXNSSVSYQID 783
>BI700867 similar to GP|1755066|gb|A lectin precursor {Sophora japonica},
partial (19%)
Length = 430
Score = 79.3 bits (194), Expect = 2e-15
Identities = 47/127 (37%), Positives = 73/127 (57%), Gaps = 4/127 (3%)
Frame = +3
Query: 200 NTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSS 259
N +N +VAVEFD+F S DP H+G+D N++ SA + ++D G I+Y++
Sbjct: 3 NLPENHVVAVEFDTFIGSTDPPTKHVGVDDNSLTSAAFGNFDIDDN-LGKKCYTLITYTA 179
Query: 260 VSRDLTVVVHYPPGQNTTNID----AIKYPIDIESYLSEWVVIGFSATTGEFAETHSILS 315
++ L V + +TN + + Y ID++ L EWV IGFSA+TG E ++I S
Sbjct: 180 STQTLFVSWSFKAKPASTNHNDNSSSFSYQIDLKKILPEWVNIGFSASTGLSTERNTIYS 359
Query: 316 WSFNSTL 322
W F+S+L
Sbjct: 360 WEFSSSL 380
>NP164640 receptor-like protein kinase
Length = 828
Score = 72.4 bits (176), Expect = 3e-13
Identities = 59/203 (29%), Positives = 95/203 (46%), Gaps = 15/203 (7%)
Frame = +1
Query: 133 AFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH--GDGFTFFLA---SLDFEFPEGGGGG 187
AFY K +V F T F F + P G GF F ++ SL +P
Sbjct: 79 AFYPTQFQFKHKNAKVVSFSTAFAFAIIPQYPKLGGHGFAFTISRSTSLKDAYPSQ---- 246
Query: 188 GFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIP---WSV 242
+LGL + N S N + AVEFD+ + D N H+GI++N + S ++ +S
Sbjct: 247 -YLGLLNPNDVGNFS-NHLFAVEFDTVQDFEFGDINDNHVGINLNNMASNKSVEAAFFSR 420
Query: 243 NDQPQ-----GAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVV 297
N++ G + A + Y + +L V + + T+ I + Y +D+ L + +
Sbjct: 421 NNKQNLNLKSGEVTQAWVDYDXLKNNLEVRLSTTSSKPTSPI--LSYKVDLSPILQDSMY 594
Query: 298 IGFSATTGEFAETHSILSWSFNS 320
+GFS++TG A +H IL WSF +
Sbjct: 595 VGFSSSTGLLASSHYILGWSFKT 663
>AW397471 similar to GP|3114258|pdb Soybean Agglutinin Complexed With 2
4-Pentasaccharide, partial (71%)
Length = 562
Score = 68.6 bits (166), Expect = 4e-12
Identities = 57/165 (34%), Positives = 81/165 (48%), Gaps = 2/165 (1%)
Frame = +2
Query: 155 FTFVVNHSQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF 214
FTF ++ DG FLA D + P+ LGLF+E N S + + VE D+
Sbjct: 29 FTFYAPDTKRLADGLACFLAPNDTK-PQTHARD--LGLFNE----NESGDQVDPVEIDTS 187
Query: 215 SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQ 274
N +DP PHIGI+ N I S T W D + I+Y + + L + Y P Q
Sbjct: 188 RNCWDPPNPHIGINGNCIISIKTTSW---DLANNKVSKVLITYDASTSLLDASLVY-P*Q 355
Query: 275 NTTNIDAIKYPIDIESYLSEWVVIGFSATTGEF--AETHSILSWS 317
T+N + +D + +EWV+IGFSA G E+H +LSW+
Sbjct: 356 RTSN--TLFDAVDQTTSRAEWVMIGFSAADGLDIPEESHDVLSWT 484
>AW472598 similar to GP|3114258|pdb Soybean Agglutinin Complexed With 2
4-Pentasaccharide, partial (49%)
Length = 421
Score = 66.6 bits (161), Expect = 1e-11
Identities = 50/146 (34%), Positives = 81/146 (55%), Gaps = 3/146 (2%)
Frame = +2
Query: 88 FSFPTFDPDVRVILLNGDAN-TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTG 146
F++ F P ++L GDA T++G LQL+K+D+ G P P S G + + +H+ DK+
Sbjct: 5 FNWNKFLPKQASMILQGDAIVTSSGKLQLSKVDENGTPKPLSLGRALYSTPIHIWDKEFV 184
Query: 147 RV--ADFRTEFTFVVNHSQPHGDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKN 204
V + F + T +V ++ G FFLA +D + P+ +GL++E Y +
Sbjct: 185 SVTASAFPSC*TLLVPDTRRLAYGLAFFLAPIDTK-PQTHASN--VGLYNE---YEYG-D 343
Query: 205 SIVAVEFDSFSNSFDPNFPHIGIDIN 230
+VA E D+ NS+DP HIGI++N
Sbjct: 344 QVVADEIDTIRNSWDPPDTHIGINVN 421
>BF071712 weakly similar to GP|22208832|em lectin {Vigna linearis}, partial
(43%)
Length = 421
Score = 62.8 bits (151), Expect = 2e-10
Identities = 49/135 (36%), Positives = 66/135 (48%), Gaps = 6/135 (4%)
Frame = +2
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV--NHSQPHGD 167
+G LQ+TK+ G P+ S G + F + + D +TG VA + T F F + H D
Sbjct: 2 SGVLQITKVCISGVPTSGSFGRALFPAPIPIWDHETGNVAHWGTFFKFNIFSPHKSDSAD 181
Query: 168 GFTFFLASLDFEFPEGG---GGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFP 223
G FF A P G GFLGLF+ +A VA+EFD+FSN +DP
Sbjct: 182 GLAFFFA------PGGSKPQSDHGFLGLFN--SALKDKSLPTVAIEFDTFSNKKWDPAHR 337
Query: 224 HIGIDINTIRSAVTI 238
IG +N I +A I
Sbjct: 338 LIGNGVNIINTASII 382
>CK605656
Length = 628
Score = 62.8 bits (151), Expect = 2e-10
Identities = 46/139 (33%), Positives = 71/139 (50%), Gaps = 4/139 (2%)
Frame = -2
Query: 188 GFLGLFDEETAYNTSKNSIVAVEFDSFSNS-FDPNFPHIGIDINTIRSAVTIPWSVNDQP 246
GF GLF+ + V +EFD+FSN +DP HIGI++N+I+S T W +++
Sbjct: 588 GFFGLFNSPLKDKFFQT--VGIEFDTFSNKKWDPANRHIGIEVNSIKSVKTASWGLSN-- 421
Query: 247 QGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTG- 305
G + +SY++ L V PP + T+ I + P++++ EWV +GF A G
Sbjct: 420 -GQVGGIFVSYNA-PPGLLVGFLIPPSKKTSYI--LFDPVNLKGNFPEWVSVGFFAPPGL 253
Query: 306 --EFAETHSILSWSFNSTL 322
ET + W F S L
Sbjct: 252 HEGSVETPEGIFWFFASKL 196
>TC204067 lectin [Glycine max]
Length = 720
Score = 49.3 bits (116), Expect(2) = 4e-10
Identities = 34/82 (41%), Positives = 48/82 (58%), Gaps = 2/82 (2%)
Frame = +3
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
NDQ + N I+Y++ S +L V P Q ++ I + +D++ L EWV IGFSA
Sbjct: 330 NDQ----VTNVLITYNA-STNLLVASLVHPSQRSSYI--LSDVLDLKVALPEWVRIGFSA 488
Query: 303 TTG--EFAETHSILSWSFNSTL 322
TTG +ETH + SWSF+S L
Sbjct: 489 TTGLNVASETHDVHSWSFSSNL 554
Score = 32.3 bits (72), Expect(2) = 4e-10
Identities = 16/31 (51%), Positives = 25/31 (80%)
Frame = +2
Query: 204 NSIVAVEFDSFSNSFDPNFPHIGIDINTIRS 234
N+I++VEFD++ + PN HIGI++N+IRS
Sbjct: 224 NNIISVEFDTWDS---PNL-HIGINVNSIRS 304
>TC217836 similar to UP|Q9M7M4 (Q9M7M4) Mannose lectin FRIL (Fragment),
partial (25%)
Length = 437
Score = 57.4 bits (137), Expect = 9e-09
Identities = 31/77 (40%), Positives = 46/77 (59%), Gaps = 2/77 (2%)
Frame = +3
Query: 248 GAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEF 307
G I ISY+S S+ LTV YP Q T + + I++ L EWV +G SA+TG+
Sbjct: 72 GKIATVHISYNSASKRLTVAAFYPGTQTVT----LSHDIELNKVLPEWVRVGLSASTGQQ 239
Query: 308 AETHSILSWS--FNSTL 322
+T++I SWS FN+++
Sbjct: 240 KQTNTIHSWSLAFNNSI 290
>BF066828 weakly similar to PIR|A31972|A319 lectin DB58 precursor - horse
gram, partial (26%)
Length = 556
Score = 54.7 bits (130), Expect = 6e-08
Identities = 39/113 (34%), Positives = 64/113 (56%), Gaps = 5/113 (4%)
Frame = +2
Query: 207 VAVEFDSFSNS-FDPNFPHIGIDINTIRSAVTIP-WSVNDQPQGAIQNAKISYSSVSRDL 264
VA+EFD+FSN +DP HIGID+N+I+S T W N Q + + ++Y++ + L
Sbjct: 218 VAIEFDTFSNKKWDPANRHIGIDVNSIKSVKTASRWLCNGQQEDSF----VTYNAAT-SL 382
Query: 265 TVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTG---EFAETHSIL 314
V P + T+ I + +++S L EW+ + FS+TTG ET+ ++
Sbjct: 383 *VGNLIRPSKKTSFI--LSGHSELDSNLPEWLSVAFSSTTGLH*RIVETNDVI 535
>AW759057 weakly similar to GP|6650223|gb| receptor-like protein kinase
{Glycine max}, partial (44%)
Length = 397
Score = 47.8 bits (112), Expect = 7e-06
Identities = 36/119 (30%), Positives = 57/119 (47%), Gaps = 4/119 (3%)
Frame = +3
Query: 204 NSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWSVNDQP--QGAIQNAKISYSS 259
N I AVEFD+ + D N H+GIDIN+++S + S+ G A + Y S
Sbjct: 12 NHIFAVEFDTVQDFEFGDINDNHVGIDINSMQSNTSANVSLVGLTLKSGKPILAWVDYDS 191
Query: 260 VSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
++V + P + + + +D+ + + +GFSA+TG A H I WSF
Sbjct: 192 RLNLISVALS--PNSSKPKTPLLTFNVDLSPVFHDTMYVGFSASTGLLASFHYI*GWSF 362
>TC220626 similar to UP|Q9M2S4 (Q9M2S4) Probable serine/threonine-specific
protein kinase, partial (7%)
Length = 543
Score = 41.6 bits (96), Expect = 5e-04
Identities = 39/130 (30%), Positives = 60/130 (46%), Gaps = 14/130 (10%)
Frame = +2
Query: 206 IVAVEFDSFSN----SFDPNFPHIGIDINTIRS--AVTIPWSVND-------QPQGAIQN 252
+VAVEFD+ N D N HIGID+N I S A T + + G +
Sbjct: 11 LVAVEFDTGRNPEFNDIDDN--HIGIDLNNIESINATTAGYFNSSGAFVPVRMRTGQNIH 184
Query: 253 AKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPID-IESYLSEWVVIGFSATTGEFAETH 311
A I + + + V V P G + ++Y I Y+S + +GFSA+ + E
Sbjct: 185 AWIDFDGENLEFNVTVA-PIGVSRPTKPTLRYQNPAIADYVSSNMYVGFSASKTNWIEAQ 361
Query: 312 SILSWSFNST 321
+L+WSF+ +
Sbjct: 362 RVLAWSFSDS 391
>TC223353 weakly similar to UP|Q9SEW3 (Q9SEW3) Receptor-like protein kinase
(Fragment), partial (17%)
Length = 714
Score = 41.6 bits (96), Expect = 5e-04
Identities = 68/258 (26%), Positives = 107/258 (41%), Gaps = 6/258 (2%)
Frame = -1
Query: 64 SELLATLIFSLL-LITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYG 122
S ++A +F L LI S S + P DPD I L GDA+ + + +
Sbjct: 657 SGMVAFSLFILCSLILLVQPSSSSTQQPPNLDPDS--IYLLGDAHVVTAADADSHV-RLT 487
Query: 123 NPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHGDGFTFFLASLDFEFPE 182
P+P SSGI + +D + TEF+F V HG G LA+
Sbjct: 486 RPAPSSSGILRRREPLAFADPTS-----LSTEFSFSVTG---HGHGLLLVLAA------- 352
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSV 242
GG ++G+ ++TSK+ V DPN H+GID+ + S +V
Sbjct: 351 GGNLSNYVGV-----EFDTSKDDNVG----------DPNANHVGIDVGSHVSVAVA--NV 223
Query: 243 ND----QPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIES-YLSEWVV 297
+D G NA + Y + S+ L V + Q ++ + + ID + + V+
Sbjct: 222 SDVHLVLNNGEKLNAWVDYEASSKVLEVRLSKWGAQKPSD-PIVSHDIDFSKIWGANPVI 46
Query: 298 IGFSATTGEFAETHSILS 315
G S++ G HS+ S
Sbjct: 45 AGISSSNG----AHSVPS 4
>TC234742 similar to UP|Q6UY58 (Q6UY58) Lectin-like receptor kinase 7;3,
partial (12%)
Length = 349
Score = 40.8 bits (94), Expect = 8e-04
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = +3
Query: 282 IKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
+ Y +D+ L E + +GFSA+TG A +H IL WSF
Sbjct: 66 LSYHVDLSPILKESMYVGFSASTGLLASSHYILGWSF 176
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,076,256
Number of Sequences: 63676
Number of extensions: 208867
Number of successful extensions: 1504
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 1422
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1462
length of query: 322
length of database: 12,639,632
effective HSP length: 97
effective length of query: 225
effective length of database: 6,463,060
effective search space: 1454188500
effective search space used: 1454188500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0541.15


