

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0487.10
(289 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
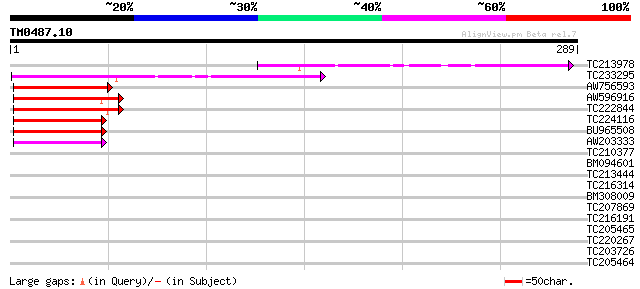
Sequences producing significant alignments: (bits) Value
TC213978 104 4e-23
TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%) 92 2e-19
AW756593 57 7e-09
AW596916 56 2e-08
TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat... 54 1e-07
TC224116 47 8e-06
BU965508 47 8e-06
AW203333 42 2e-04
TC210377 similar to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), partial... 36 0.018
BM094601 33 0.20
TC213444 33 0.20
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 32 0.26
BM308009 similar to PIR|D86241|D862 protein T16B5.8 [imported] -... 31 0.75
TC207869 similar to UP|Q94KS0 (Q94KS0) Histidine-containing phos... 31 0.75
TC216191 homologue to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A ... 31 0.75
TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 30 1.7
TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, parti... 30 1.7
TC203726 homologue to UP|Q8S4X2 (Q8S4X2) Ribulose-5-phosphate-3-... 29 2.8
TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 28 3.7
>TC213978
Length = 716
Score = 104 bits (260), Expect = 4e-23
Identities = 74/165 (44%), Positives = 96/165 (57%), Gaps = 4/165 (2%)
Frame = +1
Query: 127 SIFSCKTLVVLKL-EGLDLNT---TVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVL 182
++FSCKTLVVLKL GL+ N SVDLP L LHLQ L +RR +A+LL G P L
Sbjct: 22 ALFSCKTLVVLKLIGGLNRNPFPLDFKSVDLPLLTTLHLQSFIL-ERRDMAELLRGSPNL 198
Query: 183 EDLKAKAIYFKKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIP 242
E L +YF E FE L KL+RA I+ L EV+NNV FLRI ++
Sbjct: 199 EYLFVGHMYFSG-PEARFE--RLPKLLRATIAFGHVPL---EVVNNVQFLRID--WMEHK 354
Query: 243 HDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVIN 287
+ F NLTH+ELGY + +W++V+E + CP LQ+L I+
Sbjct: 355 EEANLIPEFQNLTHLELGYSECTRDWVDVLEVIQRCPNLQILDID 489
>TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%)
Length = 557
Score = 92.4 bits (228), Expect = 2e-19
Identities = 64/164 (39%), Positives = 93/164 (56%), Gaps = 4/164 (2%)
Frame = +3
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFR--RTKGWH 59
+DRIS LPD +L ILS LPTK+AV T ILSKRW+PLW +V+ LDFD+ + G
Sbjct: 63 DDRISHLPDVLLLQILSLLPTKQAVITGILSKRWRPLWPAVSVLDFDDESSPEFHHPGGL 242
Query: 60 SRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYS 119
+ F + V+ V+ D I++ R+ + + ++ W+ V R E V++ +
Sbjct: 243 TGFAEFVYSVLLLHD-APAIERFRLRCANPNYSA-RDIATWL-CHVARRRAERVELSLSL 413
Query: 120 LSPLNL-SSIFSCKTLVVLKLEGLDLNTTVS-SVDLPFLKVLHL 161
+ L +F C T+ V+KL G+ LN S SV LP LKVLH+
Sbjct: 414 SRYVALPRCLFHCDTVSVMKLNGVFLNALASFSVSLPLLKVLHV 545
>AW756593
Length = 370
Score = 57.4 bits (137), Expect = 7e-09
Identities = 24/50 (48%), Positives = 37/50 (74%)
Frame = +3
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANF 52
D+IS LPD IL +++ F+ T++AV T +LSKRW LW+ ++TL F+ + F
Sbjct: 63 DKISELPDNILLHMMDFMDTREAVQTCVLSKRWNNLWKRLSTLLFNTSKF 212
>AW596916
Length = 455
Score = 56.2 bits (134), Expect = 2e-08
Identities = 29/62 (46%), Positives = 39/62 (62%), Gaps = 6/62 (9%)
Frame = +3
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTL-----DF-DEANFRRTK 56
DR+S LPD +L +I+ F+ K AV T +LSKRWK LW+ +T L DF D A+F +
Sbjct: 264 DRLSDLPDFVLLHIMKFMSMKHAVQTCVLSKRWKELWKRLTNLALHSSDFADLAHFSKFL 443
Query: 57 GW 58
W
Sbjct: 444 SW 449
>TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat shock
transcription factor HSF30, partial (6%)
Length = 746
Score = 53.5 bits (127), Expect = 1e-07
Identities = 25/64 (39%), Positives = 41/64 (64%), Gaps = 8/64 (12%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFD--------EANFRR 54
DRIS+LPD +L ++++F+ TK AV T +LSKRW L + +T L F+ + +F++
Sbjct: 496 DRISALPDSLLFHMINFMDTKSAVQTCVLSKRWNDLSKCLTNLTFNSDLPSEGLDRSFKK 675
Query: 55 TKGW 58
+ W
Sbjct: 676 FESW 687
>TC224116
Length = 520
Score = 47.4 bits (111), Expect = 8e-06
Identities = 24/47 (51%), Positives = 30/47 (63%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDE 49
D IS LP I+ IL LP + AV TSILS +W+ W S+T L FD+
Sbjct: 287 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITQLVFDD 427
>BU965508
Length = 447
Score = 47.4 bits (111), Expect = 8e-06
Identities = 24/47 (51%), Positives = 30/47 (63%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDE 49
D IS LP I+ IL LP + AV TSILS +W+ W S+T L FD+
Sbjct: 296 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITRLVFDD 436
>AW203333
Length = 418
Score = 42.4 bits (98), Expect = 2e-04
Identities = 22/47 (46%), Positives = 28/47 (58%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDE 49
D IS+LP IL ILS +P + AV TS+LSK W W + L F +
Sbjct: 85 DMISTLPKTILHDILSRMPEEDAVRTSVLSKSWAEAWSTYPILYFSD 225
>TC210377 similar to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), partial (10%)
Length = 1077
Score = 36.2 bits (82), Expect = 0.018
Identities = 19/41 (46%), Positives = 28/41 (67%), Gaps = 1/41 (2%)
Frame = +3
Query: 8 LPDEILCYILSFLPTKKAVATSIL-SKRWKPLWRSVTTLDF 47
LP+ +L I+ F+ TK +V T +L SKRWK L + +T+L F
Sbjct: 294 LPNCVLLQIMEFMNTKYSVQTCVLLSKRWKDL*KYLTSLSF 416
>BM094601
Length = 429
Score = 32.7 bits (73), Expect = 0.20
Identities = 17/39 (43%), Positives = 24/39 (60%)
Frame = +3
Query: 6 SSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTT 44
+SL DE+L I LP+ + + S++ KRW L RS TT
Sbjct: 15 NSLYDELLQEIFQKLPSSSSSSVSLVCKRWLRLHRSSTT 131
>TC213444
Length = 530
Score = 32.7 bits (73), Expect = 0.20
Identities = 14/25 (56%), Positives = 20/25 (80%)
Frame = +2
Query: 2 EDRISSLPDEILCYILSFLPTKKAV 26
EDR+S +PD I+ +ILSF+ TK A+
Sbjct: 455 EDRLSDMPDCIIHHILSFMETKDAI 529
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 32.3 bits (72), Expect = 0.26
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +3
Query: 54 RTKGWHSRFVQSVHRVIHTPDFLQPIQ 80
R KGWHS + + + ++ H PDF P++
Sbjct: 309 RVKGWHSSWWKDLKKLYHQPDFQVPLE 389
>BM308009 similar to PIR|D86241|D862 protein T16B5.8 [imported] -
Arabidopsis thaliana, partial (16%)
Length = 428
Score = 30.8 bits (68), Expect = 0.75
Identities = 18/49 (36%), Positives = 29/49 (58%), Gaps = 1/49 (2%)
Frame = +3
Query: 5 ISSLPDEILCYILSFLPTKKAVAT-SILSKRWKPLWRSVTTLDFDEANF 52
+ S+PD IL ILS + + V++ + +SKRWK + TL F ++F
Sbjct: 30 MDSMPDAILQCILSRITNARDVSSCNCVSKRWKDSTPYIRTLYFPRSSF 176
>TC207869 similar to UP|Q94KS0 (Q94KS0) Histidine-containing phosphotransfer
protein, partial (93%)
Length = 802
Score = 30.8 bits (68), Expect = 0.75
Identities = 19/63 (30%), Positives = 31/63 (49%)
Frame = +3
Query: 18 SFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQSVHRVIHTPDFLQ 77
S P++ V + L K R ++ + +A+ + + GW +RF V VI D LQ
Sbjct: 378 SAFPSETLVRSKTLKGVLKTCNR*NRSIPW*KASLKLSSGWSNRFWLQVGSVIRGRDILQ 557
Query: 78 PIQ 80
P+Q
Sbjct: 558 PLQ 566
>TC216191 homologue to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A ligase ,
partial (21%)
Length = 606
Score = 30.8 bits (68), Expect = 0.75
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 6/60 (10%)
Frame = -1
Query: 121 SPLNLSSIFSCKTLVVLKLEGL------DLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQ 174
SP +FSCK V+K L + T+ + L FL LHL LQ ++CL +
Sbjct: 261 SP*RKLYLFSCKIPPVVKCMSLFHPR*SQNHLTLQQMQLEFLLQLHLSYLQRQHQKCLGE 82
>TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (15%)
Length = 490
Score = 29.6 bits (65), Expect = 1.7
Identities = 22/65 (33%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Frame = +3
Query: 6 SSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDE--ANFRRTKGWHSRFV 63
S LP E+L ILS L V S++ KRW + SV ++ F + W+ F
Sbjct: 180 SDLPTELLELILSRLSLDDNVRASVVCKRWHSVATSVCVVNQSPWLMYFPKFGDWY-EFY 356
Query: 64 QSVHR 68
VHR
Sbjct: 357 DPVHR 371
>TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, partial (7%)
Length = 586
Score = 29.6 bits (65), Expect = 1.7
Identities = 17/58 (29%), Positives = 29/58 (49%)
Frame = +2
Query: 8 LPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQS 65
LPD++L IL++LP +SKRW + S + ++ ++ K W+ F S
Sbjct: 338 LPDDLLERILAYLPIASIFRAGCVSKRWHEIVNSERFV-WNLSHVLPQKPWYFMFTSS 508
>TC203726 homologue to UP|Q8S4X2 (Q8S4X2) Ribulose-5-phosphate-3-epimerase,
complete
Length = 1748
Score = 28.9 bits (63), Expect = 2.8
Identities = 19/47 (40%), Positives = 26/47 (54%), Gaps = 2/47 (4%)
Frame = +3
Query: 114 DICIYSLSPLNLSSIFS--CKTLVVLKLEGLDLNTTVSSVDLPFLKV 158
D +YS+SP NLS IF K L L ++GL + + + L F KV
Sbjct: 1551 DKALYSVSPCNLSIIFPHIVKQLSFLSIQGLKKTS*IHYMLLFFFKV 1691
>TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (17%)
Length = 756
Score = 28.5 bits (62), Expect = 3.7
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = +1
Query: 6 SSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLD 46
S LP E+L ILS L V S++ KRW + SV ++
Sbjct: 184 SDLPTELLELILSRLSLDDNVRASVVCKRWHSVATSVCVVN 306
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,810,247
Number of Sequences: 63676
Number of extensions: 192853
Number of successful extensions: 1432
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1423
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1427
length of query: 289
length of database: 12,639,632
effective HSP length: 96
effective length of query: 193
effective length of database: 6,526,736
effective search space: 1259660048
effective search space used: 1259660048
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0487.10


