

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0476a.1
(57 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
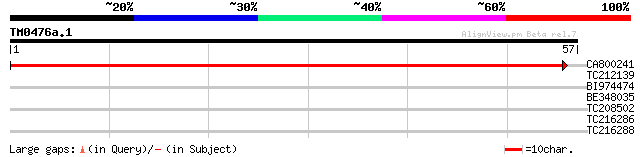
CA800241 84 1e-17
TC212139 similar to UP|Q94EN5 (Q94EN5) Beta-1,3-glucanase (Frag... 27 2.8
BI974474 26 4.8
BE348035 weakly similar to GP|5882738|gb|A Contains 3 PF|01535 D... 25 6.3
TC208502 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F2... 25 8.2
TC216286 similar to UP|Q8GTR0 (Q8GTR0) Sugar transporter, partia... 25 8.2
TC216288 homologue to UP|Q8GTR0 (Q8GTR0) Sugar transporter, part... 25 8.2
>CA800241
Length = 433
Score = 84.3 bits (207), Expect = 1e-17
Identities = 41/56 (73%), Positives = 47/56 (83%)
Frame = +3
Query: 1 RRRRVIYRSTHELDANDVVSISMWVESHLNCVFFYEDFSNSDPFILGIQTE*QLQR 56
R+ R I RST+ELD +D VSISMWVESH N VFFYEDFS+S+PF LGIQTE QLQ+
Sbjct: 12 RQERAIRRSTYELDDDDAVSISMWVESHQNLVFFYEDFSDSNPFTLGIQTEWQLQQ 179
>TC212139 similar to UP|Q94EN5 (Q94EN5) Beta-1,3-glucanase (Fragment) ,
partial (29%)
Length = 560
Score = 26.6 bits (57), Expect = 2.8
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +3
Query: 8 RSTHELDANDVVSISMWVESHLNCVF 33
+ST D ND+ +IS SH +CVF
Sbjct: 156 KSTQSCDFNDMATISTTNPSHGSCVF 233
>BI974474
Length = 343
Score = 25.8 bits (55), Expect = 4.8
Identities = 11/22 (50%), Positives = 12/22 (54%)
Frame = +3
Query: 26 ESHLNCVFFYEDFSNSDPFILG 47
+ H N VFFY D PFI G
Sbjct: 135 QCHFNVVFFYHDKMPK*PFIYG 200
>BE348035 weakly similar to GP|5882738|gb|A Contains 3 PF|01535 DUF17
domains. {Arabidopsis thaliana}, partial (5%)
Length = 469
Score = 25.4 bits (54), Expect = 6.3
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = -3
Query: 3 RRVIYRSTHELDANDVVSISMWVESHLNCVFFYE 36
R + + H A+D + +WVE L C+ FY+
Sbjct: 323 RNKVVKIVHNRFASDPS*L*IWVEGALICISFYQ 222
>TC208502 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F28L1_7
{Arabidopsis thaliana;} , partial (19%)
Length = 867
Score = 25.0 bits (53), Expect = 8.2
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -2
Query: 1 RRRRVIYRSTHELDANDVVSISMWVES 27
RRR I R H +D++ SI W+ES
Sbjct: 590 RRRVAIRRRVHVIDSHRRPSIHHWLES 510
>TC216286 similar to UP|Q8GTR0 (Q8GTR0) Sugar transporter, partial (45%)
Length = 1342
Score = 25.0 bits (53), Expect = 8.2
Identities = 10/41 (24%), Positives = 19/41 (45%)
Frame = +1
Query: 5 VIYRSTHELDANDVVSISMWVESHLNCVFFYEDFSNSDPFI 45
+IY + ++ N + + W+ S L C F + S F+
Sbjct: 1069 IIYVTVRDIIFNRLNFVPFWIRSSLACKFCLQSLSMGGSFV 1191
>TC216288 homologue to UP|Q8GTR0 (Q8GTR0) Sugar transporter, partial (16%)
Length = 886
Score = 25.0 bits (53), Expect = 8.2
Identities = 10/41 (24%), Positives = 19/41 (45%)
Frame = +1
Query: 5 VIYRSTHELDANDVVSISMWVESHLNCVFFYEDFSNSDPFI 45
+IY + ++ N + + W+ S L C F + S F+
Sbjct: 562 IIYVTVRDIIFNRLNFVPFWIRSSLACKFCLQSLSMGGSFV 684
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.144 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,938,234
Number of Sequences: 63676
Number of extensions: 31425
Number of successful extensions: 300
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 300
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 300
length of query: 57
length of database: 12,639,632
effective HSP length: 33
effective length of query: 24
effective length of database: 10,538,324
effective search space: 252919776
effective search space used: 252919776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0476a.1


