

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0437b.11
(528 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
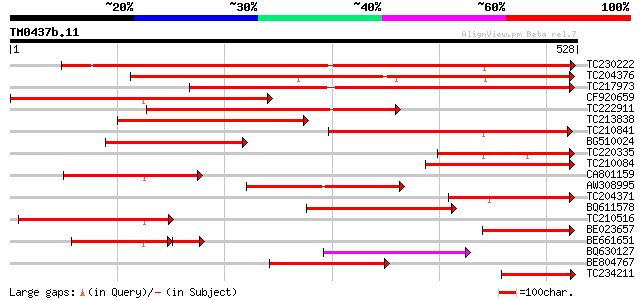
Score E
Sequences producing significant alignments: (bits) Value
TC230222 weakly similar to UP|Q9SUC6 (Q9SUC6) Berberine bridge e... 543 e-155
TC204376 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like p... 407 e-114
TC217973 weakly similar to UP|Q93ZA3 (Q93ZA3) At1g30760/T5I8_22,... 380 e-105
CF920659 345 3e-95
TC222911 similar to UP|Q94KD7 (Q94KD7) AT4g20860/T13K14_20, part... 270 2e-72
TC213838 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidop... 239 2e-63
TC210841 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidop... 219 2e-57
BG510024 similar to PIR|T10626|T10 reticuline oxidase homolog F2... 182 3e-46
TC220335 similar to UP|Q9AYM8 (Q9AYM8) CPRD2 protein, partial (25%) 177 1e-44
TC210084 weakly similar to UP|Q9FKU8 (Q9FKU8) Berberine bridge e... 170 1e-42
CA801159 similar to GP|16323167|gb At1g30760/T5I8_22 {Arabidopsi... 147 1e-35
AW308995 145 6e-35
TC204371 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like p... 134 8e-32
BQ611578 similar to GP|16323167|gb At1g30760/T5I8_22 {Arabidopsi... 134 1e-31
TC210516 weakly similar to UP|Q93Y11 (Q93Y11) Berberine bridge e... 123 2e-28
BE023657 similar to GP|16323167|gb| At1g30760/T5I8_22 {Arabidops... 118 6e-27
BE661651 86 5e-25
BQ630127 108 4e-24
BE804767 104 8e-23
TC234211 similar to UP|Q84N20 (Q84N20) Nectarin 5 (Fragment), pa... 102 3e-22
>TC230222 weakly similar to UP|Q9SUC6 (Q9SUC6) Berberine bridge enzyme-like
protein, partial (68%)
Length = 1872
Score = 543 bits (1400), Expect = e-155
Identities = 267/484 (55%), Positives = 353/484 (72%), Gaps = 5/484 (1%)
Frame = +1
Query: 49 EIVLTQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRV 108
+I T +SS Y +L +N R+++S+ KP +I+T S +QAAI CS + GLQ+RV
Sbjct: 1 KITFTSSSSLYPQVLDLLEQNPRWVNST-RKPLIILTSFHESEIQAAILCSKQLGLQLRV 177
Query: 109 RSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVH 168
RSGGHDYEGLSY+S VPF+++DL N RSI I+++DE+AWVQ+GA+LGELYY I+ S VH
Sbjct: 178 RSGGHDYEGLSYLSKVPFVMVDLINIRSIEINLDDETAWVQAGASLGELYYKISKASKVH 357
Query: 169 AFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFWA 228
FPAG CP++GIGGH SGGG + R+HGLAADHV+DA +IDVNGKI +R MGED+FWA
Sbjct: 358 GFPAGICPSIGIGGHISGGGQGMMMRRHGLAADHVVDAYLIDVNGKIHDRKSMGEDVFWA 537
Query: 229 IKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHKLPGELFLH 288
I+GG +SFGVI WK++LV VPP VT F++P+ ++ + + +WQ IAH+L +LF+
Sbjct: 538 IRGGSATSFGVILGWKIRLVRVPPIVTGFNIPRTPEEGATNLIHRWQHIAHELHEDLFIR 717
Query: 289 SVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLY 348
+ S K +F ++LG ++L+PLM +F ELGL+ DCTEM+WIQSVL+
Sbjct: 718 VIAQNSGDK---SKKFQATFNSVFLGGIDSLIPLMNESFPELGLQAKDCTEMSWIQSVLF 888
Query: 349 LAGYPINASLNVLLQRNQTFGS-FKAKSDYVNEPIPRAGLEGLWKMLLEEDS-PLLILTP 406
+AGY + L +LL R TF S FKAKSD+V EPIP++GL+G WKMLLEE++ +LIL P
Sbjct: 889 IAGYKKDDPLELLLDRITTFKSFFKAKSDFVKEPIPKSGLDGAWKMLLEEETLAMLILEP 1068
Query: 407 YGGRMSEISESETPFPHRNKSIFGIQYLVNWDKN--EETKMHVEWMRRLYAYMKPYVSKN 464
YGGRM EISES+ PFPHR +++ IQYLV W+ N EE++ H+ W + +Y YM PYVSK+
Sbjct: 1069YGGRMDEISESDIPFPHRKGNLYNIQYLVKWEVNSDEESRRHLHWAKMVYKYMTPYVSKS 1248
Query: 465 PRGAYLNYRDLDIGVNR-GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSI 523
PR AY NY+DLD+G N+ NTSY +A WG KYFK NF RL +K DP NFFR+EQSI
Sbjct: 1249PRAAYFNYKDLDLGKNKHENTSYSKASVWGEKYFKGNFRRLVHIKTTFDPQNFFRNEQSI 1428
Query: 524 PPLS 527
P L+
Sbjct: 1429PLLN 1440
>TC204376 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like protein,
partial (63%)
Length = 1418
Score = 407 bits (1046), Expect = e-114
Identities = 212/427 (49%), Positives = 286/427 (66%), Gaps = 13/427 (3%)
Frame = +1
Query: 113 HDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPA 172
HDYEG+SYIS PF+I+D+ N+R I++D+++E A V++GATLGE+YY I +S V FPA
Sbjct: 1 HDYEGISYISEEPFVILDMFNYRRITVDVKNEVAVVEAGATLGEVYYRIWEKSKVLGFPA 180
Query: 173 GSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFWAIKGG 232
G CPTVG+GGHFSGGG+ + RK+GL+ D+VIDAQI+DV G +LNR MGEDLFWAI+GG
Sbjct: 181 GVCPTVGVGGHFSGGGYGNMLRKYGLSVDNVIDAQIVDVKGNLLNRKTMGEDLFWAIRGG 360
Query: 233 GGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNV--SEIFQKWQTIAHKLPGELFLHSV 290
GG+SFGVI S+ +KLV VP VT F + K L+ NV +++ +WQ +A LF+ +
Sbjct: 361 GGASFGVILSFTIKLVPVPETVTGFRVEKTLETNVTATDLVVQWQQVAPNTDDRLFMRLL 540
Query: 291 M-GVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYL 349
+ VS+ G ++V S L+LG A ++ ++ F LGL++++CTE++WI SVL+
Sbjct: 541 LQPVSSKVVKGTRTVRASVVALFLGGANEVVSILAKEFTLLGLKKENCTEVSWIDSVLW- 717
Query: 350 AGYPINASL------NVLLQRN-QTFGSFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLL 402
+ + SL LL RN G K KSDYV I R GLE L+K ++E L
Sbjct: 718 --WNDDNSLKNGDKPETLLDRNLNNAGFLKRKSDYVQNAISRDGLEWLFKRMIELGKTGL 891
Query: 403 ILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNWDKNE--ETKMHVEWMRRLYAYMKPY 460
+ PYGG+M+EI TPFPHR +++ IQY VNWD +RL++YM P+
Sbjct: 892 VFNPYGGKMAEIPSDATPFPHRKGNLYKIQYSVNWDDRSPGAALNFTNQAKRLFSYMTPF 1071
Query: 461 VSKNPRGAYLNYRDLDIGVNR-GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRH 519
VSKNPR A+LNYRDLDIGVN G S++E +G KYF NF+RL ++K VDP NFFR+
Sbjct: 1072VSKNPRSAFLNYRDLDIGVNSFGENSFQEGVVYGTKYFNDNFQRLVKIKTIVDPENFFRN 1251
Query: 520 EQSIPPL 526
EQSIP L
Sbjct: 1252EQSIPVL 1272
>TC217973 weakly similar to UP|Q93ZA3 (Q93ZA3) At1g30760/T5I8_22, partial
(55%)
Length = 1432
Score = 380 bits (975), Expect = e-105
Identities = 182/362 (50%), Positives = 253/362 (69%), Gaps = 3/362 (0%)
Frame = +1
Query: 168 HAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFW 227
H FPAG+C ++GIGGH +GG + ++ RK+GL AD+V+DA+I+D NG+IL+R MGEDLFW
Sbjct: 1 HGFPAGTCTSLGIGGHITGGAYGSMVRKYGLGADNVLDAKIVDANGRILDRKAMGEDLFW 180
Query: 228 AIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHKLPGELFL 287
AI+GGGG SFG++ WKVKLV VPP VT+F + K L+Q +++ +WQ +A L LF+
Sbjct: 181 AIRGGGGGSFGILLWWKVKLVPVPPTVTVFTVKKTLEQGATKLLHRWQEVAPFLDENLFI 360
Query: 288 HSVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVL 347
+ + S +V S+ GL+LG A LL +M+++F ELG+ R DC E +WI+SVL
Sbjct: 361 RVRIQRAQS------TVTTSYEGLFLGGARKLLKIMKTSFPELGVTRKDCMETSWIKSVL 522
Query: 348 YLAGYPINASLNVLLQRNQTFG-SFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLLILTP 406
Y+AG+P VLL+ FK KSD+V +PIP GLEGL + LL EDSPL++ +P
Sbjct: 523 YIAGFPSGTPPEVLLKGKPIAKFFFKGKSDFVRKPIPETGLEGLRQRLLVEDSPLILWSP 702
Query: 407 YGGRMSEISESETPFPHRNKSIFGIQYLVNWDKNEE-TKMHVEWMRRLYAYMKPYVSKNP 465
YGGRM++ SES+TPFP+RN ++F Y+ W + E+ H++W+ L+ YM YV P
Sbjct: 703 YGGRMNQFSESDTPFPYRNGTLFISLYISLWQEGEKNVAKHIDWIGNLHNYMGAYVPSFP 882
Query: 466 RGAYLNYRDLDIGVN-RGNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSIP 524
RG Y+NYRDLD+G+N + NT + WG +YFK+NF+RL ++K +VDP N FRHEQSIP
Sbjct: 883 RGQYVNYRDLDLGINTKNNTGNIQESAWGYRYFKNNFDRLVKIKTKVDPQNVFRHEQSIP 1062
Query: 525 PL 526
PL
Sbjct: 1063PL 1068
>CF920659
Length = 766
Score = 345 bits (885), Expect = 3e-95
Identities = 173/247 (70%), Positives = 204/247 (82%), Gaps = 3/247 (1%)
Frame = +2
Query: 1 MAFTYTKLALLSITIFISIFLETSISFDIEKPFLHCFSSIVRNSNPTEEIVLTQNSSSYE 60
MA + KL LS+T+FISIF TS EK FL CF +I+ N T +++ T++SSSYE
Sbjct: 23 MANSMVKLVFLSVTVFISIFPATSTFAGHEKGFLQCFQTILGADNTTWQVIFTKSSSSYE 202
Query: 61 SLLQSSIRNTRFLDS-SVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLS 119
LL+SSIRN RFL+S SV KPNLIVTP L H+Q A+ CS K GLQVRVRSGGHDYEGLS
Sbjct: 203 PLLESSIRNARFLNSTSVPKPNLIVTPHSLFHIQVALFCSKKSGLQVRVRSGGHDYEGLS 382
Query: 120 YISN--VPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPT 177
Y+S+ +PFLIIDL N RSI+I++++ESAWVQSGAT+GELYYAIA +S VH FPAGSC T
Sbjct: 383 YVSHSHIPFLIIDLFNLRSITINMDEESAWVQSGATVGELYYAIAKKSKVHGFPAGSCST 562
Query: 178 VGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFWAIKGGGGSSF 237
+G+GGHFSGGGF TIFRK+GLA+D+VIDAQIIDVNG ILNR LMGEDLFWAI+GGGGSSF
Sbjct: 563 IGVGGHFSGGGFGTIFRKYGLASDNVIDAQIIDVNGMILNRTLMGEDLFWAIRGGGGSSF 742
Query: 238 GVITSWK 244
GVIT+WK
Sbjct: 743 GVITAWK 763
>TC222911 similar to UP|Q94KD7 (Q94KD7) AT4g20860/T13K14_20, partial (32%)
Length = 709
Score = 270 bits (689), Expect = 2e-72
Identities = 127/237 (53%), Positives = 172/237 (71%)
Frame = +3
Query: 128 IIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGG 187
++DL N RSI I++ DE+AWVQ+GA++GELYY I+ S VH FPAG+CP+VGIGGH SGG
Sbjct: 3 MVDLINIRSIEINLADETAWVQAGASIGELYYKISKASKVHGFPAGTCPSVGIGGHISGG 182
Query: 188 GFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLMGEDLFWAIKGGGGSSFGVITSWKVKL 247
G + RKHGLAAD+V+DA +ID NGKI +R MGED+FWAI+GG SSFGVI +WK+KL
Sbjct: 183 GQGLMLRKHGLAADNVVDAYLIDANGKIHDRKSMGEDVFWAIRGGDASSFGVILAWKIKL 362
Query: 248 VHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHKLPGELFLHSVMGVSNSPKHGGKSVIVS 307
V VPP VT F++P+ ++ V+++ +WQ IAH L +L + + +S K K +
Sbjct: 363 VRVPPIVTGFNVPRTPEEGVTDLIHRWQYIAHDLHEDLVIRVIAQISGHDK--SKKFRAT 536
Query: 308 FTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYLAGYPINASLNVLLQR 364
F ++LG + L+PLM +F ELGL+ DCTEM+WIQSV+++AGY I L +LL R
Sbjct: 537 FNSIFLGGVDRLIPLMNESFPELGLQAKDCTEMSWIQSVMFIAGYNIEDPLELLLNR 707
>TC213838 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (33%)
Length = 543
Score = 239 bits (610), Expect = 2e-63
Identities = 105/178 (58%), Positives = 142/178 (78%)
Frame = +3
Query: 101 KQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYA 160
K LQ+++RSGGHDYEG+SY++ VPF I+D+ N R+I +DI E+AWVQ+GATLGE+YY
Sbjct: 3 KHNLQMKIRSGGHDYEGVSYVAEVPFFILDMFNLRTIEVDIGTETAWVQAGATLGEVYYR 182
Query: 161 IANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNL 220
IA +S HAFPAG C TVG+GGH SGGG+ + RK+GL+ D+VIDAQ++DV G++L+R
Sbjct: 183 IAEKSKTHAFPAGVCHTVGVGGHISGGGYGNMMRKYGLSVDNVIDAQMVDVQGRLLDRKS 362
Query: 221 MGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIA 278
MGEDLFWAI GGGG+SFGV+ ++K+KLV VP VT+F + + L+QN ++I WQ +A
Sbjct: 363 MGEDLFWAITGGGGASFGVVLAYKIKLVRVPEIVTVFQVGRTLEQNATDIVYNWQHVA 536
>TC210841 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (28%)
Length = 698
Score = 219 bits (558), Expect = 2e-57
Identities = 104/231 (45%), Positives = 156/231 (67%), Gaps = 4/231 (1%)
Frame = +2
Query: 298 KHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYLAGYPINAS 357
++G K+V F L+LG +++L+ L+ F +LGL++ DC E W++SVL+ I +S
Sbjct: 2 RNGTKTVRARFIALFLGDSKSLVSLLNDKFPQLGLKQSDCIETGWLRSVLFWDNIDIASS 181
Query: 358 LNVLLQRN-QTFGSFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLLILTPYGGRMSEISE 416
L++LL+R ++ K KSDYV +PI G EG+WK ++E + L PYGGRM+EI
Sbjct: 182 LDILLERQPRSLNYLKRKSDYVKKPISIEGFEGIWKKMIELEDTLFQFNPYGGRMAEIPS 361
Query: 417 SETPFPHRNKSIFGIQYLVNWDK--NEETKMHVEWMRRLYAYMKPYVSKNPRGAYLNYRD 474
+ +PFPHR +++ IQY NW+K E ++ R+L+ +M P+VSKNPR A+ NY+D
Sbjct: 362 TASPFPHRAGNLWKIQYQANWNKPGKEVADHYINLTRKLHKFMTPFVSKNPREAFYNYKD 541
Query: 475 LDIGVN-RGNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSIP 524
LD+G+N G SY E + +G++YFK NF+RL ++K +VDP NFFR+EQSIP
Sbjct: 542 LDLGINHNGKNSYAEGRVYGVEYFKDNFDRLVQIKTKVDPHNFFRNEQSIP 694
>BG510024 similar to PIR|T10626|T10 reticuline oxidase homolog F21C20.190 -
Arabidopsis thaliana, partial (24%)
Length = 402
Score = 182 bits (462), Expect = 3e-46
Identities = 81/132 (61%), Positives = 107/132 (80%)
Frame = +1
Query: 90 SHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNFRSISIDIEDESAWVQ 149
SHVQA + C+ +Q+++RSGGHDYEG+SYIS+ PF+I+D+ NFR I++DI++E A VQ
Sbjct: 7 SHVQATVICAKSVNIQLKIRSGGHDYEGISYISDEPFIILDMFNFRRITVDIKNEVAVVQ 186
Query: 150 SGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQII 209
+GATLGE+YY I +S VH FPAG CPTVG+GGHFSGGG+ + RK+GL+ D+VIDAQI+
Sbjct: 187 AGATLGEVYYRIWKKSKVHGFPAGVCPTVGVGGHFSGGGYGNMLRKYGLSVDNVIDAQIV 366
Query: 210 DVNGKILNRNLM 221
DV G +LNR M
Sbjct: 367 DVKGNLLNRKTM 402
>TC220335 similar to UP|Q9AYM8 (Q9AYM8) CPRD2 protein, partial (25%)
Length = 535
Score = 177 bits (449), Expect = 1e-44
Identities = 86/132 (65%), Positives = 102/132 (77%), Gaps = 4/132 (3%)
Frame = +3
Query: 399 SPLLILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNWDK--NEETKMHVEWMRRLYAY 456
S + TPYG RM EISESE PFPHR +IF IQY V+W + +EE + H+ W+RR+Y+Y
Sbjct: 6 SSFVQFTPYGSRMDEISESEIPFPHRAGNIFHIQYGVSWQEEGDEEAQRHINWIRRMYSY 185
Query: 457 MKPYVSKNPRGAYLNYRDLDIGVN--RGNTSYEEAKCWGLKYFKSNFERLARVKAQVDPS 514
M+ YVSK+PR AYLNYRDLDIGVN +G TSY +A WGLKYFK+NF RLARVK VDP
Sbjct: 186 METYVSKSPRAAYLNYRDLDIGVNNNKGYTSYSQASVWGLKYFKNNFNRLARVKTNVDPL 365
Query: 515 NFFRHEQSIPPL 526
NFFR+EQSIP L
Sbjct: 366 NFFRNEQSIPSL 401
>TC210084 weakly similar to UP|Q9FKU8 (Q9FKU8) Berberine bridge enzyme,
partial (22%)
Length = 692
Score = 170 bits (431), Expect = 1e-42
Identities = 79/141 (56%), Positives = 100/141 (70%), Gaps = 2/141 (1%)
Frame = +3
Query: 388 EGLWKMLLEEDSPLLILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNW-DKNEETKMH 446
+ LWK+ +++D +I PYGG+MS I+ES TPFPHR ++ IQY+ W D + H
Sbjct: 3 DALWKIFVQDDGXXMIWNPYGGKMSRIAESATPFPHRKGVLYKIQYVTGWLDGEKSMAKH 182
Query: 447 VEWMRRLYAYMKPYVSKNPRGAYLNYRDLDIGVN-RGNTSYEEAKCWGLKYFKSNFERLA 505
+ WMR+ Y YM PYVSK PR Y+NYRDLDIG+N + NTS +A WG +YFK NF RL
Sbjct: 183 MNWMRKFYFYMAPYVSKYPRETYVNYRDLDIGMNQKNNTSLLKAWSWGYRYFKGNFNRLV 362
Query: 506 RVKAQVDPSNFFRHEQSIPPL 526
+VK +VDPSNFFRHEQSIP L
Sbjct: 363 KVKTKVDPSNFFRHEQSIPLL 425
>CA801159 similar to GP|16323167|gb At1g30760/T5I8_22 {Arabidopsis thaliana},
partial (23%)
Length = 409
Score = 147 bits (370), Expect = 1e-35
Identities = 68/131 (51%), Positives = 93/131 (70%), Gaps = 2/131 (1%)
Frame = +3
Query: 51 VLTQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRS 110
+ T ++ S+ S+L SS +N R L SV KP I TP SHVQAA+ CS K G+ +RVRS
Sbjct: 15 IYTASNPSFTSILDSSAQNLRLLVPSVPKPEFIFTPSRDSHVQAAVICSKKLGIHIRVRS 194
Query: 111 GGHDYEGLSYISNV--PFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAIANQSNVH 168
GGHDYEG+SY+S + PF+++DL R I +D++ +AWVQ+GAT GE+YY I +S+VH
Sbjct: 195 GGHDYEGISYVSEIESPFIVVDLVKLRGIDVDVKSNTAWVQAGATTGEVYYRIYEKSSVH 374
Query: 169 AFPAGSCPTVG 179
FPAG C ++G
Sbjct: 375 GFPAGLCTSLG 407
>AW308995
Length = 458
Score = 145 bits (365), Expect = 6e-35
Identities = 69/147 (46%), Positives = 100/147 (67%)
Frame = +2
Query: 221 MGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHK 280
MGEDLFWAI+GGGG+SFGV+ SWK++LV VP VT+F + + L+Q +++ KWQ +A K
Sbjct: 2 MGEDLFWAIRGGGGASFGVVVSWKIRLVPVPEVVTVFRVERTLEQGATDVVHKWQYVADK 181
Query: 281 LPGELFLHSVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEM 340
L LF+ V+ S+ + G K++ F L+LG ++ LL +M +F ELGL + C EM
Sbjct: 182 LHDGLFIRVVL--SSVKRKGVKTIRAKFNALFLGNSQELLGVMNKSFPELGLVAEQCIEM 355
Query: 341 NWIQSVLYLAGYPINASLNVLLQRNQT 367
+WI SVL+ YP+ S++VLL R+ T
Sbjct: 356 SWIDSVLFWDNYPVGTSVDVLLXRHNT 436
>TC204371 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like protein,
partial (20%)
Length = 598
Score = 134 bits (338), Expect = 8e-32
Identities = 65/121 (53%), Positives = 82/121 (67%), Gaps = 3/121 (2%)
Frame = +3
Query: 409 GRMSEISESETPFPHRNKSIFGIQYLVNWDKNEETKM--HVEWMRRLYAYMKPYVSKNPR 466
G+M+EI TPFPHR +++ IQY VNWD +RL++YM P+VSKNPR
Sbjct: 15 GKMAEIPSDATPFPHRKGNLYKIQYSVNWDDPSPGAALNFTNQAKRLFSYMTPFVSKNPR 194
Query: 467 GAYLNYRDLDIGVNR-GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSIPP 525
A+LNYRDLDIGVN G S++E +G KYF NF+RL ++K VDP NFFR+EQSIP
Sbjct: 195 SAFLNYRDLDIGVNSFGENSFQEGLVYGTKYFNDNFQRLVKIKTTVDPENFFRNEQSIPV 374
Query: 526 L 526
L
Sbjct: 375 L 377
>BQ611578 similar to GP|16323167|gb At1g30760/T5I8_22 {Arabidopsis thaliana},
partial (24%)
Length = 428
Score = 134 bits (337), Expect = 1e-31
Identities = 67/141 (47%), Positives = 94/141 (66%), Gaps = 1/141 (0%)
Frame = +2
Query: 277 IAHKLPGELFLHSVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDD 336
+A + +LF+ ++ + +++ S+ +LG A+ LL +M+ +F ELGL + D
Sbjct: 5 VAPYIDEDLFIRVIIQPATVGNKTERTITTSYNAQFLGGADRLLQVMKESFPELGLTKKD 184
Query: 337 CTEMNWIQSVLYLAGYPINASLNVLLQRNQTFGS-FKAKSDYVNEPIPRAGLEGLWKMLL 395
C E +WI+SVLY+AGYP + VLLQ TF + FKAKSD+V +PIP GLEGLW+ LL
Sbjct: 185 CLETSWIKSVLYIAGYPNDTPPEVLLQGKSTFKNYFKAKSDFVRDPIPETGLEGLWQRLL 364
Query: 396 EEDSPLLILTPYGGRMSEISE 416
EEDSPL+I PYGG MS+ SE
Sbjct: 365 EEDSPLMIWNPYGGMMSKFSE 427
>TC210516 weakly similar to UP|Q93Y11 (Q93Y11) Berberine bridge enzyme-like
protein, partial (18%)
Length = 446
Score = 123 bits (308), Expect = 2e-28
Identities = 59/146 (40%), Positives = 96/146 (65%), Gaps = 2/146 (1%)
Frame = +3
Query: 9 ALLSITIFISIFLETSISFDIEKPFLHCFSSIVRNSNPTEEIVLTQNSSSYESLLQSSIR 68
++L+ + + + + + S IE+ F HC + + N + T + S+ S+L+S+ +
Sbjct: 9 SILASFVVLLLSISFTASLPIEEAFNHCLTQHSQTPNQFSSSIYTSTNGSFTSILESTAQ 188
Query: 69 NTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNV--PF 126
N R+L SV KP+ I TP + S VQAA+ C+ K G+ +RVRSGGHDYEGLSY+S + PF
Sbjct: 189 NLRYLLPSVPKPDFIFTPLDDSQVQAAVICAKKLGIHMRVRSGGHDYEGLSYVSLIEKPF 368
Query: 127 LIIDLSNFRSISIDIEDESAWVQSGA 152
+I+DL+ R++++DI +AW+Q+GA
Sbjct: 369 MILDLAKLRAVNVDIARNTAWIQAGA 446
>BE023657 similar to GP|16323167|gb| At1g30760/T5I8_22 {Arabidopsis
thaliana}, partial (13%)
Length = 400
Score = 118 bits (296), Expect = 6e-27
Identities = 55/87 (63%), Positives = 64/87 (73%), Gaps = 1/87 (1%)
Frame = +2
Query: 441 EETKMHVEWMRRLYAYMKPYVSKNPRGAYLNYRDLDIGVN-RGNTSYEEAKCWGLKYFKS 499
E H+ WMR+ Y YM PYVSK PR Y+NYRDLDIG+N + NTS +A WG +YFK
Sbjct: 5 ESMAKHMNWMRKFYFYMAPYVSKYPRETYVNYRDLDIGMNQKNNTSLLKASSWGYRYFKG 184
Query: 500 NFERLARVKAQVDPSNFFRHEQSIPPL 526
NF RL +VK +VDPSNFFRHEQSIP L
Sbjct: 185 NFNRLVKVKTKVDPSNFFRHEQSIPLL 265
>BE661651
Length = 603
Score = 86.3 bits (212), Expect(2) = 5e-25
Identities = 39/96 (40%), Positives = 62/96 (63%), Gaps = 2/96 (2%)
Frame = +2
Query: 58 SYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEG 117
+Y +L SI+N RF + + KP IV P+ L +Q ++ C + +++RVR GGH YEG
Sbjct: 227 NYYKILNFSIQNLRFAEPVIPKPIAIVLPESLEQLQKSVACCREGFMEIRVRCGGHSYEG 406
Query: 118 LSYISN--VPFLIIDLSNFRSISIDIEDESAWVQSG 151
SY+++ PF+IID+ N + +D+E E+AWV+ G
Sbjct: 407 TSYVADDGTPFVIIDMMNLNHVWVDMETETAWVEGG 514
Score = 46.6 bits (109), Expect(2) = 5e-25
Identities = 20/30 (66%), Positives = 22/30 (72%)
Frame = +1
Query: 152 ATLGELYYAIANQSNVHAFPAGSCPTVGIG 181
ATLGE YYAI+ SN H F GSCPT G+G
Sbjct: 514 ATLGETYYAISQASNEHGFSXGSCPTGGVG 603
>BQ630127
Length = 421
Score = 108 bits (271), Expect = 4e-24
Identities = 55/138 (39%), Positives = 83/138 (59%), Gaps = 1/138 (0%)
Frame = +3
Query: 293 VSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYLAGY 352
VS++ G K++ S L+LG A+ L+ L+ F LGL+++ C EM WI SV++ A Y
Sbjct: 3 VSSNFVKGKKTIRASVEALFLGEADELVKLLGQEFPLLGLKKELCHEMRWIDSVVWWANY 182
Query: 353 PINASLNVLLQRNQ-TFGSFKAKSDYVNEPIPRAGLEGLWKMLLEEDSPLLILTPYGGRM 411
+S+N LL RN + S K KSDYV PI + G +WK ++E ++ PYGG+M
Sbjct: 183 NDGSSVNALLDRNHYSVHSNKRKSDYVQTPISKDGFTWIWKKMIELGKVSIVFNPYGGKM 362
Query: 412 SEISESETPFPHRNKSIF 429
+E+ TPFPHR +++
Sbjct: 363 NEVPSDATPFPHRAGNLY 416
>BE804767
Length = 357
Score = 104 bits (260), Expect = 8e-23
Identities = 48/111 (43%), Positives = 72/111 (64%)
Frame = -1
Query: 243 WKVKLVHVPPKVTIFDLPKKLDQNVSEIFQKWQTIAHKLPGELFLHSVMGVSNSPKHGGK 302
WK+KLV VP VT+F + K L+Q +++ Q+WQ +A K+ LF+ ++ N G +
Sbjct: 357 WKIKLVPVPQTVTVFTVTKTLEQGGNKLLQRWQQVAPKIDENLFIRVIIQPGNGTVPGKR 178
Query: 303 SVIVSFTGLYLGIAENLLPLMQSNFAELGLRRDDCTEMNWIQSVLYLAGYP 353
++ S+ L+LG A+ LL +M+ F ELGL DC E +WI+SVLY+AGYP
Sbjct: 177 TLTTSYNALFLGGADRLLQVMKHGFPELGLTIKDCVETSWIKSVLYIAGYP 25
>TC234211 similar to UP|Q84N20 (Q84N20) Nectarin 5 (Fragment), partial (16%)
Length = 431
Score = 102 bits (255), Expect = 3e-22
Identities = 47/70 (67%), Positives = 57/70 (81%), Gaps = 1/70 (1%)
Frame = +2
Query: 459 PYVSKNPRGAYLNYRDLDIGVNR-GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFF 517
PYVS +PR +Y+NYRD+DIGVN GN SY EA+ WG KYFK N++RL VK +VDPSNFF
Sbjct: 11 PYVSSSPRSSYINYRDVDIGVNGPGNASYAEARVWGEKYFKRNYDRLVEVKTKVDPSNFF 190
Query: 518 RHEQSIPPLS 527
R+EQSIP L+
Sbjct: 191 RYEQSIPSLA 220
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,785,805
Number of Sequences: 63676
Number of extensions: 328243
Number of successful extensions: 1691
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 1638
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1655
length of query: 528
length of database: 12,639,632
effective HSP length: 102
effective length of query: 426
effective length of database: 6,144,680
effective search space: 2617633680
effective search space used: 2617633680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0437b.11


