

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
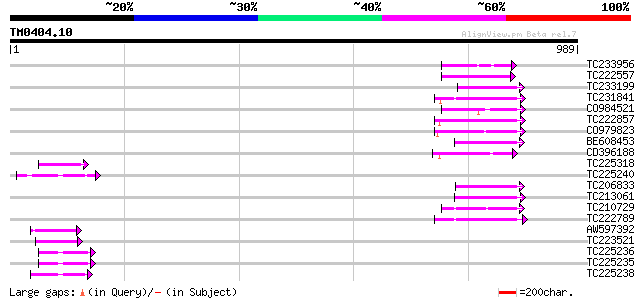
Query= TM0404.10
(989 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC233956 59 8e-09
TC222557 57 3e-08
TC233199 57 3e-08
TC231841 57 5e-08
CO984521 56 7e-08
TC222857 56 7e-08
CO979823 56 9e-08
BE608453 56 9e-08
CD396188 weakly similar to GP|6175788|gb|A ORF144 {Xestia c-nigr... 55 2e-07
TC225318 similar to UP|P93486 (P93486) Glycine-rich RNA-binding ... 55 2e-07
TC225240 similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protein 45, p... 54 3e-07
TC206833 homologue to UP|Q6PDS7 (Q6PDS7) Hist1h4h protein (Fragm... 54 3e-07
TC213061 54 4e-07
TC210729 similar to UP|Q7QHL7 (Q7QHL7) AgCP7056 (Fragment), part... 54 4e-07
TC222789 54 4e-07
AW597392 weakly similar to GP|7630030|emb| RNA binding protein-l... 53 6e-07
TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition mo... 53 8e-07
TC225236 similar to UP|Q84LL7 (Q84LL7) Salt tolerance protein 6,... 52 1e-06
TC225235 similar to GB|AAP37853.1|30725662|BT008494 At1g11650 {A... 52 1e-06
TC225238 similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protein 45, p... 52 1e-06
>TC233956
Length = 757
Score = 59.3 bits (142), Expect = 8e-09
Identities = 43/137 (31%), Positives = 68/137 (49%), Gaps = 6/137 (4%)
Frame = +2
Query: 753 QLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILA---RE 809
+L+ + K +LI+R I +E+ LC C E+ SHL C KI LW L+ +
Sbjct: 215 RLLWDRLPTKDNLIKRQIQVEDD-LCPFCHSQSETASHLFFTCGKIMPLWWEFLSWVKED 391
Query: 810 EVYCCFPSSIQGLLLEWSSLRA-ISDP--IMWEIIPYAMFWSIWRARNDLVFNQKDFSAD 866
+V+ C P + L +SS + +S+ MW I A+ SIWR RND++F +
Sbjct: 392 KVFHCRP--MDNFLQHYSSAASKVSNTRRTMWWI---AVTNSIWRLRNDIIFQNQTVDIT 556
Query: 867 YIWDLHLIRVMWWIKSW 883
+ D L + W++ W
Sbjct: 557 RLTDSTLFLLWTWLRGW 607
>TC222557
Length = 1002
Score = 57.4 bits (137), Expect = 3e-08
Identities = 39/130 (30%), Positives = 58/130 (44%), Gaps = 1/130 (0%)
Frame = +1
Query: 753 QLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVY 812
+L++ I K +L RR + + N +C C E SHL +CP+I LW LA +
Sbjct: 484 RLIKERIPTKGNLWRRRVQLNNL-MCPFCNRQEEEASHLFFNCPRILPLWWESLAWTKTV 660
Query: 813 CCF-PSSIQGLLLEWSSLRAISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDL 871
F P Q + + W+ A+ WSIWR RN +VF K F+ + D
Sbjct: 661 GVFSPIPRQNYMQHTLISKGKHLQARWKCWWVALTWSIWRHRNRIVFLNKAFNGSKLMDD 840
Query: 872 HLIRVMWWIK 881
+ V W +
Sbjct: 841 AIFLVWSWFR 870
>TC233199
Length = 1105
Score = 57.4 bits (137), Expect = 3e-08
Identities = 33/119 (27%), Positives = 57/119 (47%), Gaps = 1/119 (0%)
Frame = +1
Query: 781 CGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIM-WE 839
CG E SHL HC KI +W +A + FP+++ L+ S ++ I W
Sbjct: 580 CGCSEEEASHLFFHCLKIQPIWWETMAWVNIKSVFPTNLIHHFLQHSYVQVDGIRIKRWR 759
Query: 840 IIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHF 898
A+ WSIW+ RN ++F+ F+ + + + + + W+++ K DF+ HF
Sbjct: 760 SW*MAVVWSIWQMRNSIIFSNASFNVNKLLEDAMFLIWSWLRNLEK-------DFNIHF 915
>TC231841
Length = 791
Score = 56.6 bits (135), Expect = 5e-08
Identities = 42/164 (25%), Positives = 75/164 (45%), Gaps = 5/164 (3%)
Frame = +3
Query: 742 LWKTSLFQ----VSQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPK 797
LWK L + +L+ + +++L R+ I +++ C C ES +HL HC +
Sbjct: 174 LWKLKLPSKITIFAWRLIRDRLPTRSNLRRKQIEVDDPR-CPFCRSAEESAAHLFFHCSR 350
Query: 798 IWLLWTTILAREEVYCCFPS-SIQGLLLEWSSLRAISDPIMWEIIPYAMFWSIWRARNDL 856
I +W L+ + FP+ Q L + A W+ A+ W+IW+ RN++
Sbjct: 351 IAPVWWESLSWVNLLGVFPNHPRQHFLQHIYGVTAGMRASRWKWWWLALTWTIWKQRNNM 530
Query: 857 VFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEK 900
+F+ F+A+ I D + + W+ K DFS H+ +
Sbjct: 531 IFSNGTFNANKILDEAIFLIWTWLTHLEK-------DFSIHYNQ 641
>CO984521
Length = 716
Score = 56.2 bits (134), Expect = 7e-08
Identities = 39/156 (25%), Positives = 72/156 (46%), Gaps = 8/156 (5%)
Frame = -2
Query: 753 QLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVY 812
+L++ + KA+L ++ + ++ LC +C + E+ SHL HC K+ LW + +
Sbjct: 490 RLIQDKLPTKANLRKKRVELQEY-LCPLCRSVEETASHLFFHCSKVSPLWWESQSWVNMM 314
Query: 813 CCFP--------SSIQGLLLEWSSLRAISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFS 864
FP I G + R W+ +A+ +SIW+ RN ++F+ +F
Sbjct: 313 GVFPYQPDQHFSQHIFGASVGLQGKR-------WQWWWFALTYSIWKHRNSIIFSNANFD 155
Query: 865 ADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEK 900
A + D + + W++ + K DFS HF +
Sbjct: 154 AHKLMDDAVFILWTWLRCFEK-------DFSLHFNQ 68
>TC222857
Length = 657
Score = 56.2 bits (134), Expect = 7e-08
Identities = 42/164 (25%), Positives = 73/164 (43%), Gaps = 5/164 (3%)
Frame = -3
Query: 742 LWKTSL----FQVSQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPK 797
LWK + + +L+ + KA+L R + I + C C + E+ SH+ +HC K
Sbjct: 613 LWKIKIPSKFLMFAWRLLWDRLPTKANLRARQVQISDL-TCPFCRRVEETASHMFIHCIK 437
Query: 798 IWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAIS-DPIMWEIIPYAMFWSIWRARNDL 856
+W + + P SI +++SSL+ W+ A+ WSIW+ RN +
Sbjct: 436 TQPIWWETMNWINMQGPLPWSITDHFMQFSSLKEAGIRSRRWQWWWMAVTWSIWQLRNSI 257
Query: 857 VFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEK 900
VF+ F + + + + W+ + K DFS HF +
Sbjct: 256 VFSNATFDGNKLVEDASFLLWTWLHNLEK-------DFSLHFNQ 146
>CO979823
Length = 853
Score = 55.8 bits (133), Expect = 9e-08
Identities = 40/166 (24%), Positives = 78/166 (46%), Gaps = 7/166 (4%)
Frame = -2
Query: 742 LWK----TSLFQVSQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPK 797
+WK T + +L+ + K++L RR + +++ +C +C I E +HL +C K
Sbjct: 591 IWKLKIPTKAAVFAWRLVRDRLPTKSNLRRRQVMVQDM-VCPLCNNIEEGAAHLFFNCTK 415
Query: 798 IWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIM---WEIIPYAMFWSIWRARN 854
LW ++ + P + + L++ + I+D + W+ A+ W+IW+ RN
Sbjct: 414 TLPLWWESMSWVNLKTAMPQTPRQHFLQYGT--DIADGLKSKRWKCWWIALTWTIWQHRN 241
Query: 855 DLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEK 900
+VF F + + + L+ + W K+ K DF+ HF +
Sbjct: 240 KVVFQNATFHGNKVLEDALLLLWSWFKALEK-------DFNLHFNQ 124
>BE608453
Length = 378
Score = 55.8 bits (133), Expect = 9e-08
Identities = 33/123 (26%), Positives = 57/123 (45%), Gaps = 1/123 (0%)
Frame = +1
Query: 777 LCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPI 836
LC +C E SHL HC K+ +W ++ +V FP S + L ++
Sbjct: 13 LCPLCRTQQEDASHLFFHCSKLQPIWWETMSWLQVKGAFPLSPKQHFLHHLGVQPAGVRY 192
Query: 837 -MWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFS 895
W A+ WSIW+ RN +V + +F A+ +++ + + W++ + K DF+
Sbjct: 193 NRWHCWWLALTWSIWKLRNSIVCSNANFDANKLFEEAIFLLWSWLRGFEK-------DFT 351
Query: 896 QHF 898
HF
Sbjct: 352 VHF 360
>CD396188 weakly similar to GP|6175788|gb|A ORF144 {Xestia c-nigrum
granulovirus}, partial (5%)
Length = 701
Score = 55.1 bits (131), Expect = 2e-07
Identities = 45/158 (28%), Positives = 66/158 (41%), Gaps = 9/158 (5%)
Frame = -1
Query: 738 GLVTLWKTSL----FQVSQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLML 793
G LW+ + + +L+ + K +LIRR I ++N LC C ES SHL
Sbjct: 638 GFRQLWEIKIPPTALSFAWRLLWDRLPSKENLIRRQIVLQND-LCPFCQSQVESASHLFF 462
Query: 794 HCPKIWLLW---TTILAREEVYCCFPSS--IQGLLLEWSSLRAISDPIMWEIIPYAMFWS 848
C K+ LW T + + V P +Q L S I W A S
Sbjct: 461 TCHKVMPLWWEFNTWVREDRVLHSKPMDNFLQHCSLAGSRNSNRRRKIWW----IAATRS 294
Query: 849 IWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKE 886
IW RND++FN + F + D + W++ W K+
Sbjct: 293 IWNLRNDMIFNNQPFDISKLVDKAIFLTWSWLRGWEKD 180
>TC225318 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (86%)
Length = 749
Score = 54.7 bits (130), Expect = 2e-07
Identities = 33/87 (37%), Positives = 46/87 (51%)
Frame = +2
Query: 51 LFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRM 110
LF+ GL+ +K+AF+ FG ++ V + R FGFV FS D A A+ M
Sbjct: 176 LFIGGLSYGVDDQSLKDAFSGFGDVVDAKVITDRDSGRSRGFGFVNFSNDESASSALSAM 355
Query: 111 NGSILNGATINVSLARLPQDQHTRPRP 137
+G LNG +I VS A D+ + PRP
Sbjct: 356 DGKDLNGRSIRVSYA---NDKPSAPRP 427
>TC225240 similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protein 45, partial (48%)
Length = 928
Score = 54.3 bits (129), Expect = 3e-07
Identities = 41/146 (28%), Positives = 65/146 (44%)
Frame = +1
Query: 12 AWNKRHNTSAQSTKSRANKWSGLGNNRRADGIHRRIGKSLFVDGLTDQTTHLQVKNAFAS 71
A NK +T +Q S N + A H ++FV L T ++ F
Sbjct: 175 ASNKNPSTQSQPKASYQNP-------QGAQNEHDPNNTTIFVGNLDPNVTDDHLRQVFGQ 333
Query: 72 FGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMNGSILNGATINVSLARLPQDQ 131
+G ++ V + KR GFV+F+ S AE A+R +NG++L G + +S R P ++
Sbjct: 334 YGELVHVKIPAGKR------CGFVQFADRSCAEEALRVLNGTLLGGQNVRLSWGRSPSNK 495
Query: 132 HTRPRPTDNQWKGVSEWGEHSKPNKG 157
+ +P NQW G G + G
Sbjct: 496 --QAQPDANQWNGSGGGGYYGYAQGG 567
>TC206833 homologue to UP|Q6PDS7 (Q6PDS7) Hist1h4h protein (Fragment),
complete
Length = 1119
Score = 53.9 bits (128), Expect = 3e-07
Identities = 33/122 (27%), Positives = 55/122 (45%), Gaps = 1/122 (0%)
Frame = -3
Query: 778 CDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFP-SSIQGLLLEWSSLRAISDPI 836
C +C + ES HL C +I +W L + FP +S Q L L
Sbjct: 466 CPLCRNMEESACHLFFQCSRILPVWWESLNWTNIVGAFPLNSKQHFLYHIGGLEGGVRAN 287
Query: 837 MWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQ 896
W++ A+ W+IW+ RN+++F+ F A+ + D L + W+++ K FS
Sbjct: 286 RWKVWWLALTWTIWKHRNNVIFSNATFDANKVLDDALFLLWTWLRNLEK-------GFST 128
Query: 897 HF 898
H+
Sbjct: 127 HY 122
>TC213061
Length = 823
Score = 53.5 bits (127), Expect = 4e-07
Identities = 32/125 (25%), Positives = 57/125 (45%), Gaps = 1/125 (0%)
Frame = +2
Query: 777 LCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWS-SLRAISDP 835
+C +C + ES SHL HC KI +W L+ ++ FP + + S +
Sbjct: 353 VCPLCRSMDESASHLFFHCSKILPIWWESLSWVKLVGAFPHHPRHHFHQHSHEVYQGLQG 532
Query: 836 IMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFS 895
W+ A+ W+IW+ RND++F+ F+A + D + + W + DF+
Sbjct: 533 NRWKWWWLALTWTIWKHRNDIIFSNATFNAHKVMDDAVFLIWTWFRHLEN-------DFT 691
Query: 896 QHFEK 900
H+ +
Sbjct: 692 SHYNQ 706
>TC210729 similar to UP|Q7QHL7 (Q7QHL7) AgCP7056 (Fragment), partial (5%)
Length = 727
Score = 53.5 bits (127), Expect = 4e-07
Identities = 36/149 (24%), Positives = 70/149 (46%), Gaps = 3/149 (2%)
Frame = +2
Query: 753 QLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVY 812
+L+ + + L RR + + + C C + E+ HL HC KI LW ++ +
Sbjct: 101 RLIRDRLPTRHKLQRRQVQVADMS-CPFCRVKEENAGHLFFHCTKIQPLWWEAMSWINLK 277
Query: 813 CCFPSSIQGLLLEWSSLRAISDPIM---WEIIPYAMFWSIWRARNDLVFNQKDFSADYIW 869
P + + + L+A D + W+ A+ WSIW+ RN +VF+ F A+ ++
Sbjct: 278 GAAPLTPKMNFNQHMGLQA--DGVRSNRWQCWWMALTWSIWKTRNSIVFSNASFDANQLF 451
Query: 870 DLHLIRVMWWIKSWWKECPFTVLDFSQHF 898
+ + + W++++ K F++HF
Sbjct: 452 EDAVFILWTWLRNYEK-------GFTEHF 517
>TC222789
Length = 954
Score = 53.5 bits (127), Expect = 4e-07
Identities = 43/166 (25%), Positives = 75/166 (44%), Gaps = 5/166 (3%)
Frame = +2
Query: 742 LWKTSL---FQV-SQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPK 797
LWK + ++V + +L+ + K +L RR I + + C C + E HL HC K
Sbjct: 362 LWKLKVPIKYEVFAWRLLRDRLPTKVNLHRRQIQVMDRS-CPFCRNVEEDAGHLFFHCSK 538
Query: 798 IWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAIS-DPIMWEIIPYAMFWSIWRARNDL 856
I +W L+ + P + L+ + A W+ A+ W+IW+ RN +
Sbjct: 539 IIPIWWESLSWVNISGALPKDPRQHFLQHDLIMAGGIRTTRWKCWWLAVTWTIWQQRNKI 718
Query: 857 VFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIR 902
+F F A+ + D + W+ + K+ F+V +F+Q IR
Sbjct: 719 IFFNDSFDANKLIDEAAFLLWTWLSNLEKD--FSV-NFNQWSSNIR 847
>AW597392 weakly similar to GP|7630030|emb| RNA binding protein-like
{Arabidopsis thaliana}, partial (78%)
Length = 392
Score = 53.1 bits (126), Expect = 6e-07
Identities = 34/89 (38%), Positives = 48/89 (53%)
Frame = +2
Query: 37 NRRADGIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVR 96
NR+ + + +RIG LFV L+ TT Q+K F+ FG + + +R FGFV
Sbjct: 44 NRKVE-MSQRIGTKLFVHRLSFYTTQEQLKKLFSPFGLVTQADLALDPITKRPKGFGFVS 220
Query: 97 FSFDSDAEVAMRRMNGSILNGATINVSLA 125
F + +AE A + MNG I+NG I V A
Sbjct: 221 FKSEIEAEKACKAMNGRIVNGRLILVEPA 307
>TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (57%)
Length = 651
Score = 52.8 bits (125), Expect = 8e-07
Identities = 29/82 (35%), Positives = 43/82 (52%)
Frame = +1
Query: 45 RRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAE 104
R I LFV GL+ TT + AF+++G+++ + + R FGFV F+ +AE
Sbjct: 67 RGIAYKLFVGGLSFYTTENALSEAFSNYGQVIEAKIVTDRVSDRSKGFGFVTFASQDEAE 246
Query: 105 VAMRRMNGSILNGATINVSLAR 126
A+ M G LNG I V A+
Sbjct: 247 NAIEDMKGKTLNGRVIFVDYAK 312
>TC225236 similar to UP|Q84LL7 (Q84LL7) Salt tolerance protein 6, partial
(39%)
Length = 825
Score = 52.4 bits (124), Expect = 1e-06
Identities = 31/100 (31%), Positives = 52/100 (52%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV L T ++ F+ +G ++ V + KR GFV+F+ S AE A+R
Sbjct: 198 TIFVGNLDPNVTDDHLRQVFSQYGELVHVKIPAGKR------CGFVQFADRSCAEEALRV 359
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWG 149
+NG++L G + +S R P ++ + P NQW G + G
Sbjct: 360 LNGTLLGGQNVRLSWGRSPSNKQAQADP--NQWNGAAGAG 473
>TC225235 similar to GB|AAP37853.1|30725662|BT008494 At1g11650 {Arabidopsis
thaliana;} , partial (64%)
Length = 1154
Score = 52.4 bits (124), Expect = 1e-06
Identities = 31/100 (31%), Positives = 52/100 (52%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV L T ++ F+ +G ++ V + KR GFV+F+ S AE A+R
Sbjct: 852 TIFVGNLDPNVTDDHLRQVFSQYGELVHVKIPAGKR------CGFVQFADRSCAEEALRV 1013
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWG 149
+NG++L G + +S R P ++ + P NQW G + G
Sbjct: 1014LNGTLLGGQNVRLSWGRSPSNKQAQADP--NQWNGAAGAG 1127
Score = 30.4 bits (67), Expect = 4.0
Identities = 21/77 (27%), Positives = 35/77 (45%), Gaps = 1/77 (1%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAF-ASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++FV L T ++ F A + + G V + R +GFVRFS +S+ AM
Sbjct: 525 TIFVGDLAADVTDYLLQETFRARYNSVKGAKVVIDRLTGRTKGYGFVRFSEESEQMRAMT 704
Query: 109 RMNGSILNGATINVSLA 125
M G + + + + A
Sbjct: 705 EMQGVLCSTRPMRIGPA 755
>TC225238 similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protein 45, partial (31%)
Length = 1258
Score = 52.4 bits (124), Expect = 1e-06
Identities = 33/109 (30%), Positives = 54/109 (49%)
Frame = +1
Query: 36 NNRRADGIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFV 95
N + A H ++FV L T ++ F +G ++ V + KR GFV
Sbjct: 592 NPQGAQNEHDPNNTTIFVGNLDPNVTDDHLRQVFGHYGELVHVKIPAGKR------CGFV 753
Query: 96 RFSFDSDAEVAMRRMNGSILNGATINVSLARLPQDQHTRPRPTDNQWKG 144
+F+ S AE A+R +NG++L G + +S R P ++ + +P NQW G
Sbjct: 754 QFADRSCAEEALRVLNGTLLGGQNVRLSWGRSPSNK--QAQPDANQWNG 894
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,094,338
Number of Sequences: 63676
Number of extensions: 768748
Number of successful extensions: 5755
Number of sequences better than 10.0: 176
Number of HSP's better than 10.0 without gapping: 5133
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5698
length of query: 989
length of database: 12,639,632
effective HSP length: 106
effective length of query: 883
effective length of database: 5,889,976
effective search space: 5200848808
effective search space used: 5200848808
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0404.10


