

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0403.8
(117 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
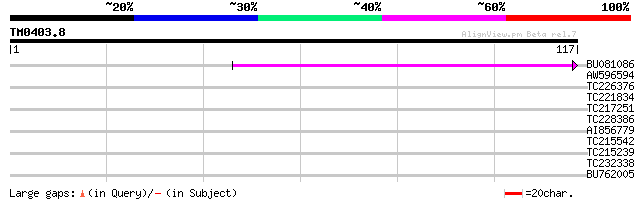
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 62 4e-11
AW596594 30 0.21
TC226376 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase... 30 0.21
TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 28 0.81
TC217251 weakly similar to UP|DJB1_HUMAN (P25685) DnaJ homolog s... 25 4.0
TC228386 25 4.0
AI856779 25 5.3
TC215542 25 5.3
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 25 5.3
TC232338 similar to UP|Q9XH61 (Q9XH61) Serine carboxypeptidase, ... 25 6.9
BU762005 similar to GP|9294061|dbj WD40-repeat protein {Arabidop... 24 9.0
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 62.0 bits (149), Expect = 4e-11
Identities = 31/71 (43%), Positives = 43/71 (59%)
Frame = +2
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
+T +I A + G G T YIPR++ + S S PFK RR+F + + +AMTIN +Q
Sbjct: 191 ITMFADHIIEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQ 370
Query: 107 GQSLSHVSLYL 117
GQ L+ V LYL
Sbjct: 371 GQLLASVGLYL 403
>AW596594
Length = 295
Score = 29.6 bits (65), Expect = 0.21
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -1
Query: 88 SRRQFLVTLCFAMTINNNQGQSLSHVSLYL 117
S+ QF++TLC A + QS+SH+ +++
Sbjct: 295 SK*QFIITLCRAALFSKGMSQSMSHLQVHI 206
>TC226376 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase subunit C
(V-ATPase C subunit) (Vacuolar proton pump C subunit) ,
partial (6%)
Length = 636
Score = 29.6 bits (65), Expect = 0.21
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -1
Query: 88 SRRQFLVTLCFAMTINNNQGQSLSHVSLYL 117
S+ QF++TLC A + QS+SH+ +++
Sbjct: 561 SK*QFIITLCRAALFSKGMSQSMSHLQVHI 472
>TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, partial (45%)
Length = 3472
Score = 27.7 bits (60), Expect = 0.81
Identities = 26/105 (24%), Positives = 53/105 (49%), Gaps = 5/105 (4%)
Frame = -3
Query: 3 *VLKRYAILLRNSKP*TNIKS*SFDHASSKY----RLSSWFV**NPMRVTHLTQ*MIVAT 58
*V RY +L+++ + +K+ F H++S+ + S+ ++ P ++ +++ T
Sbjct: 1178 *VSARYILLMKSGYTLSVLKA--FQHSNSRASPSAKFSTSYIFSTP*LSSNCLSRVVMMT 1005
Query: 59 VFPG-NKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTI 102
+ PG N+ G +P + SDS + + R F +LCF+ I
Sbjct: 1004 LLPGPNQPG----LPTSAFNCSDSLISSRTKRTLFSWSLCFSSVI 882
>TC217251 weakly similar to UP|DJB1_HUMAN (P25685) DnaJ homolog subfamily B
member 1 (Heat shock 40 kDa protein 1) (Heat shock
protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1),
partial (21%)
Length = 1669
Score = 25.4 bits (54), Expect = 4.0
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = -2
Query: 85 FKFSRRQFLVTLCFAMTINN 104
+KF RR+ ++ CF INN
Sbjct: 1641 YKFQRRKLFISACFQGQINN 1582
>TC228386
Length = 1517
Score = 25.4 bits (54), Expect = 4.0
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +3
Query: 84 PFKFSRRQFLVTLCFAMTINNN 105
PF +S Q V LC+A IN N
Sbjct: 693 PFSYSEDQKQVCLCWAFQINGN 758
>AI856779
Length = 486
Score = 25.0 bits (53), Expect = 5.3
Identities = 15/40 (37%), Positives = 24/40 (59%), Gaps = 1/40 (2%)
Frame = -2
Query: 78 SSDSGLP-FKFSRRQFLVTLCFAMTINNNQGQSLSHVSLY 116
+SD GL ++ + + LCF +TIN + QSLS ++ Y
Sbjct: 446 ASDFGLQNIRYKK*YVIYILCFGITINQSY-QSLSFLTRY 330
>TC215542
Length = 1695
Score = 25.0 bits (53), Expect = 5.3
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -1
Query: 97 CFAMTINNNQGQSLSHVSLY 116
C+ M + N QGQS H+S++
Sbjct: 372 CYTMRVINEQGQSK*HLSIH 313
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 25.0 bits (53), Expect = 5.3
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = -3
Query: 92 FLVTLCFAMTINNNQGQS 109
F + +CFAMT N ++GQ+
Sbjct: 326 FPLIVCFAMTTNKSEGQT 273
>TC232338 similar to UP|Q9XH61 (Q9XH61) Serine carboxypeptidase, partial
(57%)
Length = 1257
Score = 24.6 bits (52), Expect = 6.9
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = -2
Query: 57 ATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTL 96
+T+FPG KL K+ P I LT S +R +FLV++
Sbjct: 158 STIFPGTKLSKS---PLIMLTGSFG------NRLRFLVSI 66
>BU762005 similar to GP|9294061|dbj WD40-repeat protein {Arabidopsis
thaliana}, partial (24%)
Length = 466
Score = 24.3 bits (51), Expect = 9.0
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -2
Query: 72 PRISLTSSDSGLPFKFS 88
PRI S+D GLPF F+
Sbjct: 201 PRIPHKSADEGLPFAFA 151
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.353 0.153 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,691,199
Number of Sequences: 63676
Number of extensions: 51329
Number of successful extensions: 511
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 510
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 511
length of query: 117
length of database: 12,639,632
effective HSP length: 93
effective length of query: 24
effective length of database: 6,717,764
effective search space: 161226336
effective search space used: 161226336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0403.8


