

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
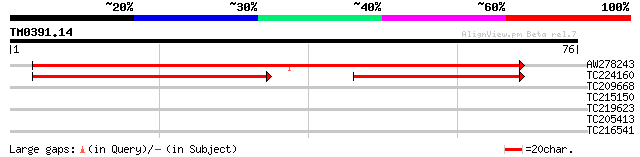
Query= TM0391.14
(76 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW278243 64 1e-11
TC224160 39 2e-09
TC209668 homologue to UP|PAL2_CICAR (Q9SMK9) Phenylalanine ammon... 27 1.5
TC215150 homologue to UP|Q9XGX6 (Q9XGX6) Cellulose synthase cata... 26 4.3
TC219623 25 5.6
TC205413 homologue to GB|AAP12863.1|30017259|BT006214 At4g19050 ... 25 9.5
TC216541 25 9.5
>AW278243
Length = 372
Score = 64.3 bits (155), Expect = 1e-11
Identities = 26/68 (38%), Positives = 46/68 (67%), Gaps = 2/68 (2%)
Frame = +1
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
F+F+YFLQI+P++ +R +++ ++ Y WH F+ N N + V +W P V IYL+DI +
Sbjct: 28 FAFAYFLQIRPLVDPTRAIIKEDNINYSWHDFVSKNNHNALTVVSVWAPVVAIYLLDIYV 207
Query: 62 WYAIYSSL 69
+Y + S++
Sbjct: 208FYTLVSAV 231
>TC224160
Length = 932
Score = 38.5 bits (88), Expect(2) = 2e-09
Identities = 13/23 (56%), Positives = 19/23 (82%)
Frame = +3
Query: 47 LWIPAVLIYLMDIQIWYAIYSSL 69
LW P +L+Y MD QIWYA++++L
Sbjct: 582 LWAPVLLVYFMDTQIWYALFTTL 650
Score = 37.7 bits (86), Expect(2) = 2e-09
Identities = 13/32 (40%), Positives = 23/32 (71%)
Frame = +1
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAF 35
F FS+F+QI+P++ ++ ++ + V Y WHAF
Sbjct: 448 FLFSFFVQIKPLVRPTKDIMSIPRVNYGWHAF 543
>TC209668 homologue to UP|PAL2_CICAR (Q9SMK9) Phenylalanine ammonia-lyase 2
, partial (43%)
Length = 940
Score = 27.3 bits (59), Expect = 1.5
Identities = 11/33 (33%), Positives = 21/33 (63%)
Frame = +1
Query: 10 LQIQPIIASSRVVLELKDVEYQWHAFIKNGNTV 42
LQI ++A +++ K ++ QWH ++N NT+
Sbjct: 223 LQI*QLVAIRA*IMDSKVLKLQWHPIVQNFNTL 321
>TC215150 homologue to UP|Q9XGX6 (Q9XGX6) Cellulose synthase catalytic
subunit, partial (12%)
Length = 712
Score = 25.8 bits (55), Expect = 4.3
Identities = 14/36 (38%), Positives = 16/36 (43%)
Frame = +3
Query: 32 WHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYS 67
WH FI N NT +LW VL + I Y S
Sbjct: 537 WHCFICNTNTNVPPVLWCSYVL*ANSEAHILYVYIS 644
>TC219623
Length = 626
Score = 25.4 bits (54), Expect = 5.6
Identities = 7/26 (26%), Positives = 18/26 (68%)
Frame = +2
Query: 44 VGLLWIPAVLIYLMDIQIWYAIYSSL 69
+G+LW+ +L+ L ++ +W +++ L
Sbjct: 254 IGILWVGLLLLGL*ELMVWEILFAPL 331
>TC205413 homologue to GB|AAP12863.1|30017259|BT006214 At4g19050 {Arabidopsis
thaliana;} , complete
Length = 1073
Score = 24.6 bits (52), Expect = 9.5
Identities = 16/55 (29%), Positives = 26/55 (47%)
Frame = -2
Query: 2 IFFSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYL 56
+FF FL+++ + S + + +EY H F+K G LLW +I L
Sbjct: 640 LFFEAHNFLKVRMVYVS---IYTEQPLEYCLHNFLKVGRKW*TKLLWEYGFIIKL 485
>TC216541
Length = 726
Score = 24.6 bits (52), Expect = 9.5
Identities = 22/68 (32%), Positives = 33/68 (48%), Gaps = 6/68 (8%)
Frame = +3
Query: 12 IQPIIASSRVVLELK-DVEYQWHAFIKNGNTVYVGL-----LWIPAVLIYLMDIQIWYAI 65
I P +ASS L + V Y + +G+ + + L L + VL+ LMD+Q Y
Sbjct: 156 ISPSVASSTPHLPVHLAVSYFLGVLLLDGHLLLLALELELDLHMQNVLVCLMDLQQIYPY 335
Query: 66 YSSLKMKL 73
SLK+ L
Sbjct: 336 LRSLKLLL 359
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,322,501
Number of Sequences: 63676
Number of extensions: 55880
Number of successful extensions: 509
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 506
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 508
length of query: 76
length of database: 12,639,632
effective HSP length: 52
effective length of query: 24
effective length of database: 9,328,480
effective search space: 223883520
effective search space used: 223883520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0391.14


