

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.11
(490 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
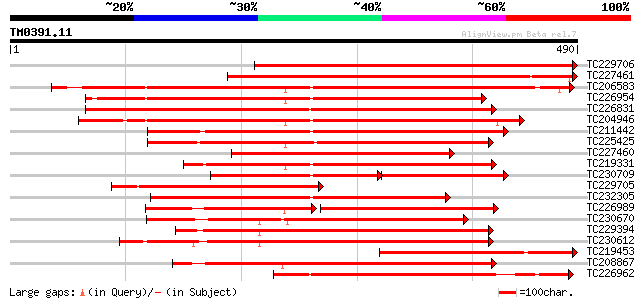
Sequences producing significant alignments: (bits) Value
TC229706 homologue to UP|Q84SA6 (Q84SA6) Serine/threonine protei... 526 e-149
TC227461 similar to UP|Q84SA6 (Q84SA6) Serine/threonine protein ... 489 e-139
TC206583 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 436 e-123
TC226954 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 431 e-121
TC226831 similar to PIR|T52285|T52285 serine/threonine-specific ... 404 e-113
TC204946 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 397 e-111
TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicite... 384 e-107
TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T2... 360 e-100
TC227460 homologue to UP|Q84SA6 (Q84SA6) Serine/threonine protei... 359 2e-99
TC219331 similar to UP|Q9FM85 (Q9FM85) Protein kinase-like prote... 350 8e-97
TC230709 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicite... 228 5e-94
TC229705 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific... 324 6e-89
TC232305 weakly similar to UP|KPCE_HUMAN (Q02156) Protein kinase... 317 1e-86
TC226989 similar to UP|PBS1_ARATH (Q9FE20) Serine/threonine-prot... 189 1e-86
TC230670 weakly similar to UP|KPEL_DROME (Q05652) Probable serin... 310 9e-85
TC229394 similar to GB|BAA98102.1|8978074|AB025609 protein serin... 306 2e-83
TC230612 similar to GB|BAA98102.1|8978074|AB025609 protein serin... 295 3e-80
TC219453 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific... 283 2e-76
TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific... 274 6e-74
TC226962 weakly similar to UP|Q9Y616 (Q9Y616) IL-1 receptor-asso... 253 2e-67
>TC229706 homologue to UP|Q84SA6 (Q84SA6) Serine/threonine protein kinase,
partial (50%)
Length = 1162
Score = 526 bits (1354), Expect = e-149
Identities = 256/279 (91%), Positives = 270/279 (96%)
Frame = +2
Query: 212 EDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDF 271
EDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHE++QRPIIYRDF
Sbjct: 2 EDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEEAQRPIIYRDF 181
Query: 272 KTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVY 331
KTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHL+SKSDVY
Sbjct: 182 KTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLTSKSDVY 361
Query: 332 SFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQK 391
SFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLG RRMF++IIDPRLEGHFSVKGAQK
Sbjct: 362 SFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGDRRMFYRIIDPRLEGHFSVKGAQK 541
Query: 392 AAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGI 451
AA LAAQCLSRDPK+RPLMSEVV LKPLP+LKDMAISSYHFQ ARVDRTMSMPNHKNGI
Sbjct: 542 AALLAAQCLSRDPKSRPLMSEVVRALKPLPSLKDMAISSYHFQIARVDRTMSMPNHKNGI 721
Query: 452 RTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKPVG 490
RTQLVS+P+KGQP+R LSSPN P+GSPY ++KSPKP+G
Sbjct: 722 RTQLVSVPRKGQPVRTLSSPNVPHGSPYLHHTKSPKPIG 838
>TC227461 similar to UP|Q84SA6 (Q84SA6) Serine/threonine protein kinase,
partial (59%)
Length = 1204
Score = 489 bits (1260), Expect = e-139
Identities = 244/304 (80%), Positives = 267/304 (87%), Gaps = 2/304 (0%)
Frame = +2
Query: 189 LAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIA 248
LAE+NYLGDL+HP+LVKLIG+CIEDDQRLLVYEFMPRGSLENHLFRR LPLPWSIRMKIA
Sbjct: 2 LAEVNYLGDLVHPHLVKLIGYCIEDDQRLLVYEFMPRGSLENHLFRRSLPLPWSIRMKIA 181
Query: 249 LGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVM 308
LGAAKGLAFLHE+++RP+IYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG+KTHVSTRVM
Sbjct: 182 LGAAKGLAFLHEEAERPVIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGDKTHVSTRVM 361
Query: 309 GTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGH 368
GTYGYAAPEYVMTGHL+S+SDVYSFGVVLLEMLTGRRS+DK RPNGEHNLVEWARP LG
Sbjct: 362 GTYGYAAPEYVMTGHLTSRSDVYSFGVVLLEMLTGRRSMDKNRPNGEHNLVEWARPHLGE 541
Query: 369 RRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAI 428
RR F+++IDPRLEGHFS+KGAQKAA LAA CLSRDPKARPLMSEVV LKPLPNLKDMA
Sbjct: 542 RRRFYKLIDPRLEGHFSIKGAQKAAHLAAHCLSRDPKARPLMSEVVEALKPLPNLKDMAS 721
Query: 429 SSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRY--SKSP 486
SSY+FQT + DR + PN +NG RTQ L + GQ R LS P+ + SPY SP
Sbjct: 722 SSYYFQTMQADRFSASPNTRNG-RTQGALLTRNGQQQRSLSIPHGTHASPYHHQFPQPSP 898
Query: 487 KPVG 490
KP G
Sbjct: 899 KPNG 910
>TC206583 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(84%)
Length = 1629
Score = 436 bits (1122), Expect = e-123
Identities = 236/460 (51%), Positives = 306/460 (66%), Gaps = 8/460 (1%)
Frame = +1
Query: 37 WFRFSFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESS 96
+F SF +CI R + G + + I +GF T+ S
Sbjct: 16 FFGVSFSANCIQDR------------NMGVCLSAQIKAESPYNTGFNSKYVSTDGNDLGS 159
Query: 97 TTTS-NEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGW 155
T + SI TP+ E L+ +S+L+ FT + LK ATRNFRP+S+LGEGGFG VFKGW
Sbjct: 160 TNDKVSANSIPQTPRSEGEILQ-SSNLKSFTLSELKTATRNFRPDSVLGEGGFGSVFKGW 336
Query: 156 IEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQ 215
I+EN KPGTG+ +AVK LN +G GH+EWLAE+NYLG H +LV+LIGFC+ED+
Sbjct: 337 IDENSLTATKPGTGIVIAVKRLNQDGIPGHREWLAEVNYLGQHFHSHLVRLIGFCLEDEH 516
Query: 216 RLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFK 272
RLLVYEFMPRGS ENHLFRR PL WS+R+K+AL AAKGLAFLH ++ +IYRDFK
Sbjct: 517 RLLVYEFMPRGSWENHLFRRGSYFQPLSWSLRLKVALDAAKGLAFLHS-AEAKVIYRDFK 693
Query: 273 TSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYS 332
TSN+LLD++YNAKLSDFGLAKDGP G+K+HVSTRVMGTYGYAAPEY+ TGHL++KSDVYS
Sbjct: 694 TSNVLLDSKYNAKLSDFGLAKDGPTGDKSHVSTRVMGTYGYAAPEYLATGHLTAKSDVYS 873
Query: 333 FGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKA 392
FGVVLLEML+G+R++DK RP+G+HNLVEWA+P + ++R F+++D RL+G +S A K
Sbjct: 874 FGVVLLEMLSGKRAVDKNRPSGQHNLVEWAKPFMANKRKIFRVLDTRLQGQYSTDDAYKL 1053
Query: 393 AQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIR 452
A LA +CLS + K RP M +VV TL+ L ++A V+R P+ NG R
Sbjct: 1054ATLALRCLSIESKFRPNMDQVVTTLEQLQLSNVNGGPRVRRRSADVNRGHQNPSSVNGSR 1233
Query: 453 TQLVSLPKKGQPLRILSSPNC---PNGSP-YSRYSKSPKP 488
+ + + L +PN P+ SP Y+ + K P
Sbjct: 1234VR----RRSADDISRLETPNAYPRPSASPLYT*HCKHTHP 1341
>TC226954 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(80%)
Length = 1175
Score = 431 bits (1109), Expect = e-121
Identities = 218/350 (62%), Positives = 266/350 (75%), Gaps = 3/350 (0%)
Frame = +2
Query: 66 NSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKF 125
N IK+V + +G + SS + S+ SI T + E L+ +S+L+ F
Sbjct: 143 NRIKAV----SPSNTGITSRSVSRSGHDISSNSRSSSASIPVTSRSEGEILQ-SSNLKSF 307
Query: 126 TFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGH 185
+++ L+ ATRNFRP+S+LGEGGFG VFKGWI+E+ A KPG G VAVK LN +G QGH
Sbjct: 308 SYHELRAATRNFRPDSVLGEGGFGSVFKGWIDEHSLAATKPGIGKIVAVKKLNQDGLQGH 487
Query: 186 KEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPWS 242
+EWLAE+NYLG L HPNLVKLIG+C ED+ RLLVYEFMP+GS+ENHLFRR P WS
Sbjct: 488 REWLAEINYLGQLQHPNLVKLIGYCFEDEHRLLVYEFMPKGSMENHLFRRGSYFQPFSWS 667
Query: 243 IRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTH 302
+RMKIALGAAKGLAFLH ++ +IYRDFKTSNILLD YNAKLSDFGLA+DGP G+K+H
Sbjct: 668 LRMKIALGAAKGLAFLHS-TEHKVIYRDFKTSNILLDTNYNAKLSDFGLARDGPTGDKSH 844
Query: 303 VSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWA 362
VSTRVMGT GYAAPEY+ TGHL++KSDVYSFGVVLLEM++GRR+IDK +P GEHNLVEWA
Sbjct: 845 VSTRVMGTRGYAAPEYLATGHLTTKSDVYSFGVVLLEMISGRRAIDKNQPTGEHNLVEWA 1024
Query: 363 RPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSE 412
+P L ++R F+++DPRLEG +S AQ AA LA QC S +PK RP M E
Sbjct: 1025KPYLSNKRRVFRVMDPRLEGQYSQNRAQAAAALAMQCFSVEPKCRPNMDE 1174
>TC226831 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (82%)
Length = 1693
Score = 404 bits (1038), Expect = e-113
Identities = 206/357 (57%), Positives = 263/357 (72%), Gaps = 2/357 (0%)
Frame = +1
Query: 66 NSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSE-ELKVASSLRK 124
+S K S SG K + + S + S ++ P SE E+ + +L+
Sbjct: 259 SSAKVEAAHSSRTPSGISKTSPSSVPSNLSILSYSEASDFSNLPTPRSEGEILSSPNLKA 438
Query: 125 FTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQG 184
FTFN LK ATRNFRP+SLLGEGGFG V+KGWI+E+ KPG+G+ VAVK L G QG
Sbjct: 439 FTFNELKNATRNFRPDSLLGEGGFGYVYKGWIDEHTFTASKPGSGMVVAVKKLKPEGLQG 618
Query: 185 HKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR-PLPLPWSI 243
HKEWL E++YLG L H NLVKLIG+C + + RLLVYEFM +GSLENHLFRR P PL WS+
Sbjct: 619 HKEWLTEVDYLGQLHHQNLVKLIGYCADGENRLLVYEFMSKGSLENHLFRRGPQPLSWSV 798
Query: 244 RMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHV 303
RMK+A+GAA+GL+FLH +++ +IYRDFK SNILLDAE+NAKLSDFGLAK GP G++THV
Sbjct: 799 RMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDAEFNAKLSDFGLAKAGPTGDRTHV 975
Query: 304 STRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWAR 363
ST+VMGT GYAAPEYV TG L++KSDVYSFGVVLLE+L+GRR++D+ + E NLVEWA+
Sbjct: 976 STQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLSGRRAVDRSKAGVEQNLVEWAK 1155
Query: 364 PVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
P LG +R F+I+D +L G + KGA AA LA +CL+R+ K RP ++EV+ TL+ +
Sbjct: 1156PYLGDKRRLFRIMDTKLGGQYPQKGAYMAATLALKCLNREAKGRPPITEVLQTLEQI 1326
>TC204946 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(78%)
Length = 1607
Score = 397 bits (1020), Expect = e-111
Identities = 211/393 (53%), Positives = 275/393 (69%), Gaps = 7/393 (1%)
Frame = +3
Query: 60 TSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVA 119
T G + K V ++++ S + + P S + E S+ TP++ E L+ +
Sbjct: 123 TQIKAGLNSKHVSADAKDHSSPISNKITKDVSTPISKVS---EVSVPQTPRIEGEILQ-S 290
Query: 120 SSLRKFTFNGLKVATRNFRPESLLG-EGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILN 178
S+L+ F+ L ATRNFR +S+LG EG FG VFKGWI+ A KPGTG+ VAVK L+
Sbjct: 291 SNLKNFSLTELTAATRNFRKDSVLGGEGDFGSVFKGWIDNQSLAAAKPGTGVVVAVKRLS 470
Query: 179 HNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL- 237
+ QGHK+ LAE+NYLG L HP+LVKLIG+C ED RLLVYEFMPRGSLENHLF R
Sbjct: 471 LDSFQGHKDRLAEVNYLGQLSHPHLVKLIGYCFEDKDRLLVYEFMPRGSLENHLFMRGSY 650
Query: 238 --PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDG 295
PL W +R+K+ALGAAKGLAFLH ++ +IYRDFKTSN+LLD+ YNAKL+D GLAKDG
Sbjct: 651 FQPLSWGLRLKVALGAAKGLAFLHS-AETKVIYRDFKTSNVLLDSNYNAKLADLGLAKDG 827
Query: 296 PEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGE 355
P EK+H STRVMGTYGYAAPEY+ TG+LS+KSDV+SFGVVLLEML+GRR++DK RP+G+
Sbjct: 828 PTREKSHASTRVMGTYGYAAPEYLATGNLSAKSDVFSFGVVLLEMLSGRRAVDKNRPSGQ 1007
Query: 356 HNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVH 415
HNLVEWA+P L ++R +++D RLEG + + A K A L+ +CL+ + K RP M EV
Sbjct: 1008HNLVEWAKPYLSNKRKLLRVLDNRLEGQYELDEACKVATLSLRCLAIESKLRPTMDEVAT 1187
Query: 416 TLKPL--PNLK-DMAISSYHFQTARVDRTMSMP 445
L+ L P++K + S+ HF R+ + P
Sbjct: 1188DLEQLQVPHVKQNRRKSADHFTHGRIATASASP 1286
>TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicited protein
264, partial (73%)
Length = 1047
Score = 384 bits (985), Expect = e-107
Identities = 193/313 (61%), Positives = 242/313 (76%), Gaps = 1/313 (0%)
Frame = +2
Query: 120 SSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNH 179
S L FT L+ AT +F ++LGEGGFG V+KG++++ + +K TVAVK L+
Sbjct: 20 SKLYAFTLEELREATNSFSWSNMLGEGGFGPVYKGFVDDKLRSGLK---AQTVAVKRLDL 190
Query: 180 NGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR-PLP 238
+G QGH+EWLAE+ +LG L HP+LVKLIG+C ED+ RLL+YE+MPRGSLEN LFRR
Sbjct: 191 DGLQGHREWLAEIIFLGQLRHPHLVKLIGYCYEDEHRLLMYEYMPRGSLENQLFRRYSAA 370
Query: 239 LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG 298
+PWS RMKIALGAAKGLAFLHE + +P+IYRDFK SNILLD+++ AKLSDFGLAKDGPEG
Sbjct: 371 MPWSTRMKIALGAAKGLAFLHE-ADKPVIYRDFKASNILLDSDFTAKLSDFGLAKDGPEG 547
Query: 299 EKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNL 358
E THV+TR+MGT GYAAPEY+MTGHL++KSDVYS+GVVLLE+LTGRR +DK R N +L
Sbjct: 548 EDTHVTTRIMGTQGYAAPEYIMTGHLTTKSDVYSYGVVLLELLTGRRVVDKSRSNEGKSL 727
Query: 359 VEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLK 418
VEWARP+L ++ + IID RLEG F +KGA K A LA +CLS P ARP MS+V+ L+
Sbjct: 728 VEWARPLLRDQKKVYNIIDRRLEGQFPMKGAMKVAMLAFKCLSHHPNARPTMSDVIKVLE 907
Query: 419 PLPNLKDMAISSY 431
PL + D+ I +
Sbjct: 908 PLQDYDDVFIGPF 946
>TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T23A1.8
{Arabidopsis thaliana;} , partial (80%)
Length = 1386
Score = 360 bits (925), Expect = e-100
Identities = 181/302 (59%), Positives = 226/302 (73%), Gaps = 3/302 (0%)
Frame = +1
Query: 120 SSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNH 179
S+LR FTF LK ATRNF+ +++LGEGGFG VFKGW+EE T+ K G+G +AVK LN
Sbjct: 97 SNLRIFTFAELKAATRNFKADTVLGEGGFGKVFKGWLEEKATS--KGGSGTVIAVKKLNS 270
Query: 180 NGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL-- 237
QG +EW +E+N+LG L H NLVKL+G+C+E+ + LLVYEFM +GSLENHLF R
Sbjct: 271 ESLQGLEEWQSEVNFLGRLSHTNLVKLLGYCLEESELLLVYEFMQKGSLENHLFGRGSAV 450
Query: 238 -PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGP 296
PLPW IR+KIA+GAA+GLAFLH + +IYRDFK SNILLD YNAK+SDFGLAK GP
Sbjct: 451 QPLPWDIRLKIAIGAARGLAFLHTSEK--VIYRDFKASNILLDGSYNAKISDFGLAKLGP 624
Query: 297 EGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEH 356
++HV+TRVMGT+GYAAPEYV TGHL KSDVY FGVVL+E+LTG R++D RP+G+H
Sbjct: 625 SASQSHVTTRVMGTHGYAAPEYVATGHLYVKSDVYGFGVVLVEILTGLRALDSNRPSGQH 804
Query: 357 NLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHT 416
L EW +P L RR I+D RLEG F K A + AQL+ +CL+ +PK RP M +V+
Sbjct: 805 KLTEWVKPYLHDRRKLKGIMDSRLEGKFPSKAAFRIAQLSMKCLASEPKHRPSMKDVLEN 984
Query: 417 LK 418
L+
Sbjct: 985 LE 990
>TC227460 homologue to UP|Q84SA6 (Q84SA6) Serine/threonine protein kinase,
partial (44%)
Length = 581
Score = 359 bits (921), Expect = 2e-99
Identities = 168/193 (87%), Positives = 186/193 (96%)
Frame = +3
Query: 192 LNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGA 251
+N+LGDL+HPNLVKL+G+CIE+DQRLLVYEFMPRGSLENHLFRR +PLPWSIRMKIALGA
Sbjct: 3 VNFLGDLVHPNLVKLVGYCIEEDQRLLVYEFMPRGSLENHLFRRSIPLPWSIRMKIALGA 182
Query: 252 AKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTY 311
AKGLAFLHE+++RP+IYRDFKTSNILLDAEYNAKLSDFGLAKDGPEG+KTHVSTRVMGTY
Sbjct: 183 AKGLAFLHEEAERPVIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGDKTHVSTRVMGTY 362
Query: 312 GYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRM 371
GYAAPEYVMTGHL+SKSDVYSFGVVLLEMLTGRRS+DK RPNGEHNLVEWA+P LG RR
Sbjct: 363 GYAAPEYVMTGHLTSKSDVYSFGVVLLEMLTGRRSMDKHRPNGEHNLVEWAQPHLGERRR 542
Query: 372 FFQIIDPRLEGHF 384
F+++IDP LE HF
Sbjct: 543 FYKLIDPHLESHF 581
>TC219331 similar to UP|Q9FM85 (Q9FM85) Protein kinase-like protein
(At5g56460), partial (61%)
Length = 1183
Score = 350 bits (898), Expect = 8e-97
Identities = 173/274 (63%), Positives = 217/274 (79%), Gaps = 4/274 (1%)
Frame = +1
Query: 151 VFKGWIEENGTAPVKPGTGLTVAVKILN-HNGHQGHKEWLAELNYLGDLLHPNLVKLIGF 209
V+KG+I E P L VAVK+ + N HQGH+EWL+++ + G L HPNLVK+IG+
Sbjct: 7 VYKGFISEELIRKGLPT--LDVAVKVHDGDNSHQGHREWLSQVEFWGQLSHPNLVKVIGY 180
Query: 210 CIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPI 266
C ED+ R+L+YE+M RG L+N+LF+ PL WS+RMKIA GAAKGLAFLHE +++P+
Sbjct: 181 CCEDNHRVLIYEYMSRGGLDNYLFKYAPAIPPLSWSMRMKIAFGAAKGLAFLHE-AEKPV 357
Query: 267 IYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSS 326
IYRDFKTSNILLD EYN+KLSDFGLAKDGP G+K+HVSTRVMGTYGYAAPEY+MTGHL+
Sbjct: 358 IYRDFKTSNILLDQEYNSKLSDFGLAKDGPVGDKSHVSTRVMGTYGYAAPEYIMTGHLTP 537
Query: 327 KSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSV 386
+SDVYSFGVVLLE+LTGR+S+DK RP E NL EWA P+L ++ F IIDPRL+G + +
Sbjct: 538 RSDVYSFGVVLLELLTGRKSLDKLRPAREQNLAEWALPLLKEKKKFLNIIDPRLDGDYPI 717
Query: 387 KGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
K KAA LA CL+R+PKARPLM ++V +L+PL
Sbjct: 718 KAVHKAAMLAYHCLNRNPKARPLMRDIVDSLEPL 819
>TC230709 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicited protein
264, partial (61%)
Length = 1245
Score = 228 bits (582), Expect(2) = 5e-94
Identities = 111/150 (74%), Positives = 130/150 (86%), Gaps = 1/150 (0%)
Frame = +3
Query: 174 VKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF 233
VK+L+ +G QGHKEWL E+ +LG L HP+LVKLIG+C E++ RLLVYE++PRGSLEN LF
Sbjct: 3 VKLLDLDGSQGHKEWLTEVVFLGQLRHPHLVKLIGYCCEEEHRLLVYEYLPRGSLENQLF 182
Query: 234 RR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLA 292
R LPWS RMKIA GAAKGLAFLHE +++P+IYRDFK SNILLD++YNAKLSDFGLA
Sbjct: 183 RIFTASLPWSTRMKIAAGAAKGLAFLHE-AKKPVIYRDFKASNILLDSDYNAKLSDFGLA 359
Query: 293 KDGPEGEKTHVSTRVMGTYGYAAPEYVMTG 322
KDGPEG+ THVSTRVMGT GYAAPEY+MTG
Sbjct: 360 KDGPEGDDTHVSTRVMGTQGYAAPEYIMTG 449
Score = 134 bits (338), Expect(2) = 5e-94
Identities = 67/110 (60%), Positives = 82/110 (73%)
Frame = +2
Query: 322 GHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLE 381
GHL++ SDVYSFGVVLLE+LTGRRS+DK RP E NLVEWAR L R +I+DPRLE
Sbjct: 563 GHLTAMSDVYSFGVVLLELLTGRRSVDKGRPQREQNLVEWARSALNDSRKLSRIMDPRLE 742
Query: 382 GHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSY 431
G +S GA+KAA LA QCLS P++RPLMS VV+ L+PL + D+ I +
Sbjct: 743 GQYSEVGARKAAALAYQCLSHRPRSRPLMSTVVNVLEPLQDFDDVPIGPF 892
>TC229705 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific protein
kinase-like, partial (35%)
Length = 550
Score = 324 bits (830), Expect = 6e-89
Identities = 160/184 (86%), Positives = 171/184 (91%), Gaps = 1/184 (0%)
Frame = +1
Query: 89 TEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGF 148
T APP SSTTTSN ES+ STPK FSEELKV+S LRKFTFN LK+ATRNFRPESLLGEGGF
Sbjct: 1 TNAPPGSSTTTSNAESVPSTPK-FSEELKVSSRLRKFTFNELKLATRNFRPESLLGEGGF 177
Query: 149 GCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIG 208
GCVFKGWIEENGTAPVKPGTGLTVAVK LNH+G QGHKEWLAEL+ LGDL+HPNLVKL+G
Sbjct: 178 GCVFKGWIEENGTAPVKPGTGLTVAVKTLNHDGLQGHKEWLAELDILGDLVHPNLVKLVG 357
Query: 209 FCIEDDQRLLVYEFMPRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPII 267
FCIEDDQRLLVYE MPRGSLENHLFR+ LPLPWSIRMKIALGAAKGLAFLHE++QRP+I
Sbjct: 358 FCIEDDQRLLVYECMPRGSLENHLFRKGSLPLPWSIRMKIALGAAKGLAFLHEEAQRPVI 537
Query: 268 YRDF 271
YRDF
Sbjct: 538 YRDF 549
>TC232305 weakly similar to UP|KPCE_HUMAN (Q02156) Protein kinase C, epsilon
type (nPKC-epsilon) , partial (5%)
Length = 908
Score = 317 bits (811), Expect = 1e-86
Identities = 158/261 (60%), Positives = 199/261 (75%), Gaps = 1/261 (0%)
Frame = +3
Query: 122 LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNG 181
L K+T + L+ ATRNFRP+++LGEGGFG VFKGWI++N P + G G+ VAVK N +
Sbjct: 129 LIKYTLDELRSATRNFRPDTVLGEGGFGRVFKGWIDKNTFKPSRVGVGIPVAVKKSNPDS 308
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR-PLPLP 240
QG +EW +E+ LG HPNL KLIG+C E+ Q LLVYE+M +GSLE+HLFRR P PL
Sbjct: 309 LQGLEEWQSEVQLLGKFSHPNLGKLIGYCWEESQFLLVYEYMQKGSLESHLFRRGPKPLS 488
Query: 241 WSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEK 300
W IR+KIA+GAA+GLAFLH S++ +IYRDFK+SNILLD ++NAKLSDFGLAK GP K
Sbjct: 489 WDIRLKIAIGAARGLAFLHT-SEKSVIYRDFKSSNILLDGDFNAKLSDFGLAKFGPVNGK 665
Query: 301 THVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVE 360
+HV+TR+MGTYGYAAPEY+ TGHL KSDVY FGVVLLEMLTGR ++D +P G NLVE
Sbjct: 666 SHVTTRIMGTYGYAAPEYMATGHLYIKSDVYGFGVVLLEMLTGRAALDTNQPTGMQNLVE 845
Query: 361 WARPVLGHRRMFFQIIDPRLE 381
L ++ +++DP +E
Sbjct: 846 CTMSSLHAKKRLKEVMDPNME 908
>TC226989 similar to UP|PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase
PBS1 (AvrPphB susceptible protein 1) , partial (86%)
Length = 1730
Score = 189 bits (480), Expect(2) = 1e-86
Identities = 91/154 (59%), Positives = 113/154 (73%)
Frame = +3
Query: 269 RDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKS 328
RD K+SNILLD Y+ KLSDFGLAK GP G+K+HVSTRVMGTYGY APEY MTG L+ KS
Sbjct: 660 RDCKSSNILLDEGYHPKLSDFGLAKLGPVGDKSHVSTRVMGTYGYCAPEYAMTGQLTVKS 839
Query: 329 DVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKG 388
DVYSFGVV LE++TGR++ID +P GE NLV WARP+ RR F ++ DPRL+G F ++G
Sbjct: 840 DVYSFGVVFLELITGRKAIDSTQPQGEQNLVTWARPLFNDRRKFSKLADPRLQGRFPMRG 1019
Query: 389 AQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPN 422
+A +A+ C+ RPL+ +VV L L N
Sbjct: 1020LYQALAVASMCIQESAATRPLIGDVVTALSYLAN 1121
Score = 149 bits (376), Expect(2) = 1e-86
Identities = 82/151 (54%), Positives = 101/151 (66%), Gaps = 3/151 (1%)
Frame = +2
Query: 118 VASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL 177
V + + FTF L AT+NFRPES +GEGGFG V+KG +E T VAVK L
Sbjct: 224 VQIAAQTFTFRELAAATKNFRPESFVGEGGFGRVYKGRLET---------TAQIVAVKQL 376
Query: 178 NHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP- 236
+ NG QG++E+L E+ L L HPNLV LIG+C + DQRLLVYEFMP GSLE+HL P
Sbjct: 377 DKNGLQGNREFLVEVLMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPP 556
Query: 237 --LPLPWSIRMKIALGAAKGLAFLHEDSQRP 265
PL W+ RMKIA+GAAKGL +LH+ + P
Sbjct: 557 DKEPLDWNTRMKIAVGAAKGLEYLHDKANPP 649
>TC230670 weakly similar to UP|KPEL_DROME (Q05652) Probable
serine/threonine-protein kinase pelle , partial (10%)
Length = 1094
Score = 310 bits (794), Expect = 9e-85
Identities = 159/285 (55%), Positives = 202/285 (70%), Gaps = 7/285 (2%)
Frame = +2
Query: 119 ASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILN 178
A+ LR F+F+ LK ATR F L+GEGGFG V++G +++N VA+K LN
Sbjct: 269 ANHLRLFSFSDLKSATRAFSRALLVGEGGFGSVYRGLLDQND-----------VAIKQLN 415
Query: 179 HNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD----QRLLVYEFMPRGSLENHLFR 234
NGHQGHKEW+ ELN LG + HPNLVKL+G+C EDD QRLLVYEFMP SLE+HL
Sbjct: 416 RNGHQGHKEWINELNLLGVVKHPNLVKLVGYCAEDDERGIQRLLVYEFMPNKSLEDHLLA 595
Query: 235 RPLP---LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGL 291
R +P +PW R++IA AA+GLA+LHE+ +I+RDFKTSNILLD +NAKLSDFGL
Sbjct: 596 R-VPSTIIPWGTRLRIAQDAARGLAYLHEEMDFQLIFRDFKTSNILLDENFNAKLSDFGL 772
Query: 292 AKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKR 351
A+ GP +VST V+GT GYAAPEYV TG L++KSDV+SFGVVL E++TGRR++D+
Sbjct: 773 ARQGPSEGSGYVSTAVVGTIGYAAPEYVQTGKLTAKSDVWSFGVVLYELITGRRAVDRNL 952
Query: 352 PNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLA 396
P E L++W RP + R F I+DPRL+G + +K A K A LA
Sbjct: 953 PRNEQKLLDWVRPYVSDPRKFHHILDPRLKGQYCIKSAHKLAILA 1087
>TC229394 similar to GB|BAA98102.1|8978074|AB025609 protein serine/threonine
kinase-like {Arabidopsis thaliana;} , partial (63%)
Length = 1170
Score = 306 bits (783), Expect = 2e-83
Identities = 153/280 (54%), Positives = 203/280 (71%), Gaps = 5/280 (1%)
Frame = +3
Query: 144 GEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNL 203
GEGGFG V+KG I + G + VA+K LN G QGHKEWLAE+ +LG + HPNL
Sbjct: 3 GEGGFGSVYKGSIAQLDGQ----GDPIPVAIKRLNTRGFQGHKEWLAEVQFLGIVNHPNL 170
Query: 204 VKLIGFCIEDD----QRLLVYEFMPRGSLENHLFRRPLP-LPWSIRMKIALGAAKGLAFL 258
VKL+G+C D QRLLVYEFMP SLE+HLF + LP LPW R++I LGAA+GLA+L
Sbjct: 171 VKLLGYCSVDGERGIQRLLVYEFMPNRSLEDHLFNKKLPTLPWKTRLEIMLGAAQGLAYL 350
Query: 259 HEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEY 318
HE + +IYRDFK+SN+LLDA+++ KLSD GLA++GP+G++THVST V+GT GYAAPEY
Sbjct: 351 HEGLEIQVIYRDFKSSNVLLDADFHPKLSDXGLAREGPQGDQTHVSTAVVGTQGYAAPEY 530
Query: 319 VMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDP 378
+ TGHL +SD++SFGVVL E+LTGRRS+++ RP E L++W + F I+D
Sbjct: 531 IETGHLKVQSDMWSFGVVLYEILTGRRSLERNRPTAEQKLLDWVKQYPADTSRFVIIMDA 710
Query: 379 RLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLK 418
RL +S+ A+K A+LA CL ++P+ RP MS++V +LK
Sbjct: 711 RLRNQYSLPAARKIAKLADSCLKKNPEDRPSMSQIVESLK 830
>TC230612 similar to GB|BAA98102.1|8978074|AB025609 protein serine/threonine
kinase-like {Arabidopsis thaliana;} , partial (70%)
Length = 1606
Score = 295 bits (755), Expect = 3e-80
Identities = 164/332 (49%), Positives = 220/332 (65%), Gaps = 9/332 (2%)
Frame = +3
Query: 96 STTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGW 155
S+T+S S S +L+ E + R FT L AT F +GEGGFG V++G
Sbjct: 135 SSTSSPLPSPRSIKELYKEN---EHNFRIFTLQELVDATHGFNRMLKIGEGGFGKVYRGT 305
Query: 156 IE---ENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIE 212
I+ E+G P+ VA+K LN G QGHKEWLAE+ +L + HPNLVKL+G+C
Sbjct: 306 IKPHPEDGADPI------LVAIKKLNTRGLQGHKEWLAEVQFLSVVNHPNLVKLLGYCSV 467
Query: 213 DD----QRLLVYEFMP-RGSLENHLFRRPLP-LPWSIRMKIALGAAKGLAFLHEDSQRPI 266
D QRLLVYEFM R S + F LP L W R++I LGAA+GL +LH + +
Sbjct: 468 DSEKGIQRLLVYEFMSNRESGKITFFSLSLPHLTWKTRLQIMLGAAQGLHYLHNGLEVKV 647
Query: 267 IYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSS 326
IYRDFK+SN+LLD +++ KLSDFGLA++GP G++THVST V+GT GYAAPEYV TGHL
Sbjct: 648 IYRDFKSSNVLLDKKFHPKLSDFGLAREGPTGDQTHVSTAVVGTQGYAAPEYVETGHLKI 827
Query: 327 KSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSV 386
+SD++SFGVVL E+LTGRR +++ RP GE L+EW + + F +IIDPRL+ +S+
Sbjct: 828 QSDIWSFGVVLYEILTGRRVLNRNRPIGEQKLIEWVKNYPANSSRFSKIIDPRLKNQYSL 1007
Query: 387 KGAQKAAQLAAQCLSRDPKARPLMSEVVHTLK 418
A+K A+LA CL ++P+ RP MS++V +LK
Sbjct: 1008GAARKVAKLADNCLKKNPEDRPSMSQIVESLK 1103
>TC219453 similar to UP|Q9LFP7 (Q9LFP7) Serine/threonine specific protein
kinase-like, partial (23%)
Length = 882
Score = 283 bits (723), Expect = 2e-76
Identities = 141/171 (82%), Positives = 152/171 (88%)
Frame = +1
Query: 320 MTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPR 379
MTGHL+SKSDVYSFGVVLLEMLTGRRSIDK RPNGEHNLVEWARPVLG RRM +IIDPR
Sbjct: 7 MTGHLTSKSDVYSFGVVLLEMLTGRRSIDKNRPNGEHNLVEWARPVLGDRRMLLRIIDPR 186
Query: 380 LEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVD 439
LEGHFSVKG+QKAAQLAAQCL+RDPK+RP+MSEVV LKPL NLKDMAISSYHFQ ARVD
Sbjct: 187 LEGHFSVKGSQKAAQLAAQCLNRDPKSRPMMSEVVQALKPLQNLKDMAISSYHFQVARVD 366
Query: 440 RTMSMPNHKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKPVG 490
RTMSMP KNG++ QL SL +KGQP+RILSSP P+GSPY Y KSPKP G
Sbjct: 367 RTMSMP--KNGMQAQLASLSRKGQPVRILSSPKGPHGSPYHHYIKSPKPNG 513
>TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific protein
kinase-like protein, partial (72%)
Length = 1118
Score = 274 bits (701), Expect = 6e-74
Identities = 140/286 (48%), Positives = 192/286 (66%), Gaps = 6/286 (2%)
Frame = +1
Query: 141 SLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLH 200
+++G GGFG V++G + + G VA+K ++ G QG +E+ E+ L L
Sbjct: 10 NVIGHGGFGLVYRGVLND----------GRKVAIKFMDQAGKQGEEEFKVEVELLSRLHS 159
Query: 201 PNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFR------RPLPLPWSIRMKIALGAAKG 254
P L+ L+G+C + + +LLVYEFM G L+ HL+ P+ L W R++IAL AAKG
Sbjct: 160 PYLLALLGYCSDSNHKLLVYEFMANGGLQEHLYPVSNSIITPVKLDWETRLRIALEAAKG 339
Query: 255 LAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYA 314
L +LHE P+I+RDFK+SNILLD +++AK+SDFGLAK GP+ HVSTRV+GT GY
Sbjct: 340 LEYLHEHVSPPVIHRDFKSSNILLDKKFHAKVSDFGLAKLGPDRAGGHVSTRVLGTQGYV 519
Query: 315 APEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQ 374
APEY +TGHL++KSDVYS+GVVLLE+LTGR +D KRP GE LV WA P+L R +
Sbjct: 520 APEYALTGHLTTKSDVYSYGVVLLELLTGRVPVDMKRPPGEGVLVSWALPLLTDREKVVK 699
Query: 375 IIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
I+DP LEG +S+K + A +AA C+ + RPLM++VV +L PL
Sbjct: 700 IMDPSLEGQYSMKEVVQVAAIAAMCVQPEADYRPLMADVVQSLVPL 837
>TC226962 weakly similar to UP|Q9Y616 (Q9Y616) IL-1
receptor-associated-kinase-M (Interleukin-1
receptor-associated kinase 3), partial (8%)
Length = 1084
Score = 253 bits (645), Expect = 2e-67
Identities = 138/261 (52%), Positives = 177/261 (66%), Gaps = 2/261 (0%)
Frame = +1
Query: 229 ENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLS 287
ENHLFRR P PL WS+RMK+A+GAA+GL+FLH +++ +IYRDFK SNILLDAE+N+KLS
Sbjct: 13 ENHLFRRGPQPLSWSVRMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDAEFNSKLS 189
Query: 288 DFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSI 347
DFGLAK GP G++THVST+VMGT GYAAPEYV TG L++KSDVYSFGVVLLE+L+GRR++
Sbjct: 190 DFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLSGRRAV 369
Query: 348 DKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKAR 407
DK E NLV+WA+P L +R F+I+D +LEG + KGA AA LA QCL+ + KAR
Sbjct: 370 DKTITGMEQNLVDWAKPYLSDKRRLFRIMDTKLEGQYPQKGAFTAATLALQCLNSEAKAR 549
Query: 408 PLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKGQPLRI 467
P M+EV+ TL+ + +T +H R Q P + P R
Sbjct: 550 PPMTEVLATLEQI----------------EAPKTAGRNSHSEHHRLQ---TPVRKSPARN 672
Query: 468 LSSPN-CPNGSPYSRYSKSPK 487
S N P SP + +SP+
Sbjct: 673 RSPLNLTPTASPLPAHRQSPR 735
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,778,407
Number of Sequences: 63676
Number of extensions: 316496
Number of successful extensions: 3566
Number of sequences better than 10.0: 974
Number of HSP's better than 10.0 without gapping: 2793
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2846
length of query: 490
length of database: 12,639,632
effective HSP length: 101
effective length of query: 389
effective length of database: 6,208,356
effective search space: 2415050484
effective search space used: 2415050484
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0391.11


