

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
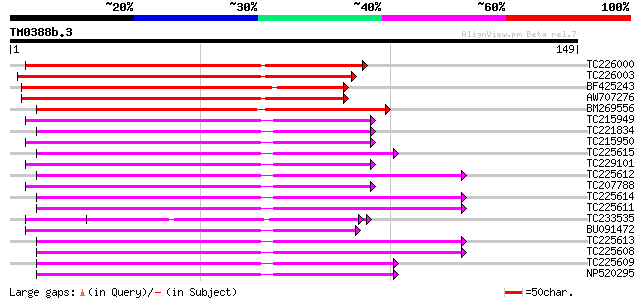
TC226000 similar to UP|O49469 (O49469) TMV resistance protein N-... 72 7e-14
TC226003 70 5e-13
BF425243 69 6e-13
AW707276 similar to GP|9858483|gb|A resistance protein MG55 {Gly... 68 2e-12
BM269556 similar to PIR|T06143|T06 disease resistence protein ho... 66 5e-12
TC215949 similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resi... 62 8e-11
TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 61 2e-10
TC215950 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 61 2e-10
TC225615 UP|Q84ZU7 (Q84ZU7) R 5 protein, complete 59 9e-10
TC229101 disease resistance-like protein 59 9e-10
TC225612 homologue to UP|Q84ZU8 (Q84ZU8) R 10 protein, complete 58 2e-09
TC207788 homologue to UP|Q8W2C0 (Q8W2C0) Functional candidate re... 56 6e-09
TC225614 UP|Q84ZV7 (Q84ZV7) R 12 protein, partial (88%) 56 6e-09
TC225611 homologue to UP|Q84ZU6 (Q84ZU6) R 1 protein, complete 55 1e-08
TC233535 weakly similar to UP|Q6XZH4 (Q6XZH4) Nematode resistanc... 54 2e-08
BU091472 similar to GP|9965103|gb|A resistance protein LM6 {Glyc... 54 3e-08
TC225613 UP|Q9FVK5 (Q9FVK5) Resistance protein LM6 (Fragment), p... 54 3e-08
TC225608 UP|Q84ZU5 (Q84ZU5) R 8 protein, complete 54 3e-08
TC225609 UP|Q84ZV3 (Q84ZV3) R 4 protein, complete 54 4e-08
NP520295 disease resistance-like protein GS5-3 [Glycine max] 52 1e-07
>TC226000 similar to UP|O49469 (O49469) TMV resistance protein N-like,
partial (4%)
Length = 938
Score = 72.4 bits (176), Expect = 7e-14
Identities = 40/90 (44%), Positives = 57/90 (62%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L L+L N K L++ P F L+ LD GC L + PSIGLL +L +L+L+ C +LV
Sbjct: 150 LTSLNLRNCKSLIKLPRFGEDLILKNLDLEGCKKLRHIDPSIGLLKKLEYLNLKNCKNLV 329
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNF 94
SL ++ LNSL+ L+LSGC++L +T F
Sbjct: 330 SL-PNSILGLNSLQYLNLSGCSKLYNTELF 416
>TC226003
Length = 557
Score = 69.7 bits (169), Expect = 5e-13
Identities = 41/89 (46%), Positives = 55/89 (61%)
Frame = +1
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
P L L+L N K L++ P F LE L GC L + PSIGLL +L +L+LE C +
Sbjct: 292 PKLTSLNLRNCKSLIKLPRFGEDLILENLYLEGCHKLRHIDPSIGLL*KLKYLNLENC*N 471
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLEST 91
LVSL ++ LNSL L+LSGC++L +T
Sbjct: 472 LVSL-PNSILGLNSLEYLYLSGCSKLYNT 555
>BF425243
Length = 418
Score = 69.3 bits (168), Expect = 6e-13
Identities = 39/86 (45%), Positives = 56/86 (64%)
Frame = +1
Query: 4 FLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
+LK L+LS+SKYL ETPNF LE+L C L +VH SIG L L ++L+ C +L
Sbjct: 13 WLKILNLSHSKYLTETPNFSKLPNLEKLILKDCPRLCKVHKSIGDLCNLHLINLKDCKTL 192
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLE 89
+L G V +L S++ L LSGC++++
Sbjct: 193GNLPRG-VYKLKSVKTLILSGCSKID 267
>AW707276 similar to GP|9858483|gb|A resistance protein MG55 {Glycine max},
partial (36%)
Length = 571
Score = 67.8 bits (164), Expect = 2e-12
Identities = 36/86 (41%), Positives = 57/86 (65%)
Frame = +2
Query: 4 FLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
+LK L+LS+SKYL TPNF G LE+L C ++ +VH SIG L +L ++++ C+SL
Sbjct: 188 WLKILNLSHSKYLTATPNFSGLPGLEKLILKDCPSMSKVHKSIGDLHKLVLINMKDCTSL 367
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLE 89
+L + L S++ L+LSGC+ ++
Sbjct: 368 TNL-PREMYHLKSVKTLNLSGCSEID 442
>BM269556 similar to PIR|T06143|T06 disease resistence protein homolog
F24J7.60 - Arabidopsis thaliana, partial (2%)
Length = 427
Score = 66.2 bits (160), Expect = 5e-12
Identities = 36/93 (38%), Positives = 58/93 (61%)
Frame = +3
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
++LS SK L ++P+F G+ LE L GCT+L +VHPS+ +LA ++L+ C L +
Sbjct: 3 INLSFSKNLKQSPDFGGAPNLESLVLEGCTSLTEVHPSLVRHKKLAMMNLKDCKRLKT-- 176
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSH 100
L + E++SL+ L+LSGC+ + P F + H
Sbjct: 177 LPSKMEMSSLKDLNLSGCSEFKYLPEFGESMEH 275
>TC215949 similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, partial (30%)
Length = 1805
Score = 62.4 bits (150), Expect = 8e-11
Identities = 37/92 (40%), Positives = 47/92 (50%)
Frame = +2
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L L L L E P+ +LE+L F C NL +H S+GLL +L FL+ E C L
Sbjct: 746 LTSLTLDECDSLTEIPDVSCLSKLEKLSFGNCRNLFTIHHSVGLLEKLKFLNAESCPELK 925
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
S +L SL L LS C+ LES P +G
Sbjct: 926 SF---PPLKLTSLESLKLSYCSSLESFPEILG 1012
Score = 30.0 bits (66), Expect = 0.42
Identities = 32/119 (26%), Positives = 50/119 (41%), Gaps = 4/119 (3%)
Frame = +2
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC----SSL 63
L+LS SK+ + + L L+ C L ++ G+ L S GC SS
Sbjct: 1343 LELSRSKFTVIPECIKECRFLTILNVDRCDRLQEIR---GIPPNLEEFSATGCPALTSSS 1513
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQF 122
+S++L EL+ R ++ P + P F C PGN I +WF ++F
Sbjct: 1514 ISMFLNQ--ELHEARDIYFH-------LPR-VKIPEWFECQS----PGNSISFWFRNEF 1648
>TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, partial (45%)
Length = 3472
Score = 61.2 bits (147), Expect = 2e-10
Identities = 31/89 (34%), Positives = 49/89 (54%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L K+L + P+ G+ L+ L F C NL+++H S+G L +L ++ EGCS L +
Sbjct: 2164 LSFDKFKFLTQIPDLSGTPNLKELSFVLCENLVEIHESVGFLDKLKSMNFEGCSKLETF- 2340
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIG 96
+L SL ++LS C+ L S P +G
Sbjct: 2341 --PPIKLTSLESINLSYCSSLVSFPEILG 2421
>TC215950 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (23%)
Length = 1891
Score = 60.8 bits (146), Expect = 2e-10
Identities = 37/92 (40%), Positives = 47/92 (50%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L L L L E P+ LE L F+ C NL ++H S+GLL +L L+ EGC L
Sbjct: 508 LTSLILDECDSLTEIPDVSCLSNLENLSFSECLNLFRIHHSVGLLGKLKILNAEGCPELK 687
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
S +L SL L LS C+ LES P +G
Sbjct: 688 SF---PPLKLTSLESLDLSYCSSLESFPEILG 774
Score = 26.2 bits (56), Expect = 6.1
Identities = 28/95 (29%), Positives = 36/95 (37%), Gaps = 23/95 (24%)
Frame = +1
Query: 55 LSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLEST----PNF---------------- 94
L LEG S ++ + E L IL LSGC RL+ PN
Sbjct: 1114 LRLEG--SKCTVIPECIKECRFLSILILSGCYRLQEIRGIPPNLERFAATESPDLTSSSI 1287
Query: 95 ---IGNPSHFRCGFDIVVPGNKIPYWFDSQFKGGS 126
+ H D +P KIP WF+ Q +G S
Sbjct: 1288 SMLLNQELHEAGHTDFSLPILKIPEWFECQSRGPS 1392
>TC225615 UP|Q84ZU7 (Q84ZU7) R 5 protein, complete
Length = 2724
Score = 58.9 bits (141), Expect = 9e-10
Identities = 34/95 (35%), Positives = 45/95 (46%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L N K+L + P+ L L F GC +L+ V SIG L +L L+ GC L S
Sbjct: 1885 LKFDNCKFLTQIPDVSDLPNLRELSFKGCESLVAVDDSIGFLNKLKKLNAYGCRKLTSF- 2061
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L SL L LSGC+ LE P +G + +
Sbjct: 2062 --PPLNLTSLETLQLSGCSSLEYFPEILGEMENIK 2160
>TC229101 disease resistance-like protein
Length = 2755
Score = 58.9 bits (141), Expect = 9e-10
Identities = 30/92 (32%), Positives = 50/92 (53%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
++ L+ K+L TP+ G+ L+ L C NL+++H S+G L +L ++ E CS L
Sbjct: 1213 MRVLNFDRCKFLTRTPDLSGAPILKELSSVFCENLVEIHDSVGFLDKLEIMNFESCSKLE 1392
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
+ +L SL ++LS C+ L S P +G
Sbjct: 1393 TF---PPIKLTSLESINLSHCSSLVSFPEILG 1479
Score = 28.5 bits (62), Expect = 1.2
Identities = 38/149 (25%), Positives = 54/149 (35%), Gaps = 27/149 (18%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K LDLS + + + + L +L CT+L ++ G+ L LS C+SL
Sbjct: 1801 VKSLDLSANNFTILPSCIQECRLLRKLYLDYCTHLHEIR---GIPPNLETLSAIRCTSLK 1971
Query: 65 SLYLGTVCELNS----LRILHLSGCTRLEST---PNFIGNPSHFRC-------------- 103
L L E LR L L C L+ P I S C
Sbjct: 1972 DLDLAVPLESTKEGCCLRQLILDDCENLQEIRGIPPSIEFLSATNCRSLTASCRRMLLKQ 2151
Query: 104 ------GFDIVVPGNKIPYWFDSQFKGGS 126
+PG +IP WF+ +G S
Sbjct: 2152 ELHEAGNKRYSLPGTRIPEWFEHCSRGQS 2238
>TC225612 homologue to UP|Q84ZU8 (Q84ZU8) R 10 protein, complete
Length = 2706
Score = 57.8 bits (138), Expect = 2e-09
Identities = 38/113 (33%), Positives = 53/113 (46%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L N K+L + P+ L L F C +L+ V SIG L +L LS GCS L S
Sbjct: 1897 LKFDNCKFLTQIPDVSDLPNLRELSFEECESLVAVDDSIGFLNKLKKLSAYGCSKLKSF- 2073
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL+ L LS C+ LE P IG + + F +P ++ + F +
Sbjct: 2074 --PPLNLTSLQTLELSQCSSLEYFPEIIGEMENIKHLFLYGLPIKELSFSFQN 2226
>TC207788 homologue to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, complete
Length = 3949
Score = 56.2 bits (134), Expect = 6e-09
Identities = 32/92 (34%), Positives = 46/92 (49%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L L+ + ++L P+ L++L F C NL +HPS+G L +L L EGCS L
Sbjct: 2005 LTSLNFDSCQHLTLIPDVSCVPHLQKLSFKDCDNLYAIHPSVGFLEKLRILDAEGCSRLK 2184
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
+ +L SL L L C LE+ P +G
Sbjct: 2185 NF---PPIKLTSLEQLKLGFCHSLENFPEILG 2271
>TC225614 UP|Q84ZV7 (Q84ZV7) R 12 protein, partial (88%)
Length = 2367
Score = 56.2 bits (134), Expect = 6e-09
Identities = 34/113 (30%), Positives = 50/113 (44%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L K+L + P+ L L F GC +L+ + SIG L +L L+ GC L S
Sbjct: 1879 LKFDKCKFLTQIPDVSDLPNLRELSFVGCESLVAIDDSIGFLNKLEILNAAGCRKLTSF- 2055
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL L LS C+ LE P +G + +P ++P+ F +
Sbjct: 2056 --PPLNLTSLETLELSHCSSLEYFPEILGEMENITALHLERLPIKELPFSFQN 2208
>TC225611 homologue to UP|Q84ZU6 (Q84ZU6) R 1 protein, complete
Length = 2709
Score = 55.1 bits (131), Expect = 1e-08
Identities = 35/113 (30%), Positives = 53/113 (45%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L+ ++L + P+ L+ L F C +L+ V SIG L +L LS GC L S
Sbjct: 1897 LNFDQCEFLTQIPDVSDLPNLKELSFDWCESLIAVDDSIGFLNKLKKLSAYGCRKLRSF- 2073
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL L LSGC+ LE P +G + + +P ++P+ F +
Sbjct: 2074 --PPLNLTSLETLQLSGCSSLEYFPEILGEMENIKALDLDGLPIKELPFSFQN 2226
>TC233535 weakly similar to UP|Q6XZH4 (Q6XZH4) Nematode resistance-like
protein (Fragment), partial (4%)
Length = 741
Score = 54.3 bits (129), Expect = 2e-08
Identities = 31/91 (34%), Positives = 48/91 (52%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L ++D ++ ++L E P+ G L L C NL+++H S+G L L L+ GC+SL
Sbjct: 237 LTKMDFTDCEFLSEVPDISGIPDLRILYLDNCINLIKIHDSVGFLGNLEELTTIGCTSL- 413
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
+ + +L SLR L S C RL P +
Sbjct: 414 -KIIPSAFKLASLRELSFSECLRLVRFPEIL 503
Score = 40.4 bits (93), Expect = 3e-04
Identities = 25/73 (34%), Positives = 40/73 (54%)
Frame = +3
Query: 21 NFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRIL 80
NF + L ++DFT C L +V P I + +L L L+ C +L+ ++ +V L +L L
Sbjct: 216 NFKNMECLTKMDFTDCEFLSEV-PDISGIPDLRILYLDNCINLIKIH-DSVGFLGNLEEL 389
Query: 81 HLSGCTRLESTPN 93
GCT L+ P+
Sbjct: 390 TTIGCTSLKIIPS 428
>BU091472 similar to GP|9965103|gb|A resistance protein LM6 {Glycine max},
partial (10%)
Length = 426
Score = 53.9 bits (128), Expect = 3e-08
Identities = 33/88 (37%), Positives = 42/88 (47%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+ ++L E P+ LE L F C NL+ VH SIG L +L L C L
Sbjct: 166 LKVLNFEQCEFLTEIPDVSXLLNLEELSFNRCGNLITVHDSIGFLNKLKILGATRCRKLT 345
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTP 92
+ L SL L LS C+ LE+ P
Sbjct: 346 TF---PPLNLTSLERLELSCCSSLENFP 420
>TC225613 UP|Q9FVK5 (Q9FVK5) Resistance protein LM6 (Fragment), partial (98%)
Length = 2597
Score = 53.9 bits (128), Expect = 3e-08
Identities = 34/113 (30%), Positives = 52/113 (45%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L+ ++L + P+ L+ L F C +L+ V SIG L +L LS GC L S
Sbjct: 1855 LNFDRCEFLTKIPDVSDLPNLKELSFNWCESLVAVDDSIGFLNKLKTLSAYGCRKLTSF- 2031
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL L+L GC+ LE P +G + +P ++P+ F +
Sbjct: 2032 --PPLNLTSLETLNLGGCSSLEYFPEILGEMKNITVLALHDLPIKELPFSFQN 2184
>TC225608 UP|Q84ZU5 (Q84ZU5) R 8 protein, complete
Length = 3084
Score = 53.9 bits (128), Expect = 3e-08
Identities = 34/113 (30%), Positives = 52/113 (45%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L+ ++L + P+ L+ L F C +L+ V SIG L +L LS GC L S
Sbjct: 1894 LNFDRCEFLTKIPDVSDLPNLKELSFNWCESLVAVDDSIGFLNKLKTLSAYGCRKLTSF- 2070
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL L+L GC+ LE P +G + +P ++P+ F +
Sbjct: 2071 --PPLNLTSLETLNLGGCSSLEYFPEILGEMKNITVLALHDLPIKELPFSFQN 2223
Score = 31.6 bits (70), Expect = 0.15
Identities = 17/53 (32%), Positives = 22/53 (41%), Gaps = 2/53 (3%)
Frame = +1
Query: 76 SLRILHLSGCTRLESTPN--FIGNPSHFRCGFDIVVPGNKIPYWFDSQFKGGS 126
+L+ C L S+ + H G + V PG IP WFD Q G S
Sbjct: 2590 NLKHFDARNCASLTSSSKSMLLNQELHEAGGIEFVFPGTSIPEWFDQQSSGHS 2748
>TC225609 UP|Q84ZV3 (Q84ZV3) R 4 protein, complete
Length = 2688
Score = 53.5 bits (127), Expect = 4e-08
Identities = 33/95 (34%), Positives = 42/95 (43%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L K+L + P+ L L F C +L+ V SIG L +L LS GC L S
Sbjct: 1891 LKFDRCKFLTQIPDVSDLPNLRELSFEDCESLVAVDDSIGFLKKLKKLSAYGCRKLTSF- 2067
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L SL L LS C+ LE P +G + R
Sbjct: 2068 --PPLNLTSLETLQLSSCSSLEYFPEILGEMENIR 2166
>NP520295 disease resistance-like protein GS5-3 [Glycine max]
Length = 467
Score = 52.0 bits (123), Expect = 1e-07
Identities = 32/95 (33%), Positives = 44/95 (45%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L+ ++L + P+ LE L F C +L+ V SIG L +L LS GC L S
Sbjct: 94 LNFDQCEFLTKIPDVSDLPNLEELSFNWCESLVAVDDSIGFLNKLKKLSAYGCRKLTSF- 270
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L SL L LS C+ LE P +G + +
Sbjct: 271 --PPLNLTSLETLALSHCSSLEYFPEILGEMENIK 369
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,433,890
Number of Sequences: 63676
Number of extensions: 133478
Number of successful extensions: 848
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 786
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 827
length of query: 149
length of database: 12,639,632
effective HSP length: 89
effective length of query: 60
effective length of database: 6,972,468
effective search space: 418348080
effective search space used: 418348080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0388b.3


