

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
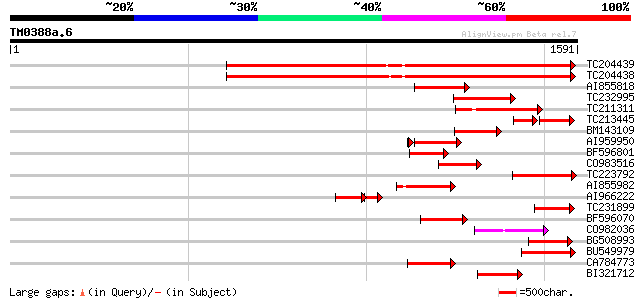
Query= TM0388a.6
(1591 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 958 0.0
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 954 0.0
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 229 6e-60
TC232995 224 2e-58
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 210 5e-54
TC213445 123 5e-47
BM143109 172 1e-42
AI959950 168 2e-41
BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Gl... 157 3e-38
CO983516 152 1e-36
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 149 1e-35
AI855982 148 2e-35
AI966222 112 5e-30
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 127 4e-29
BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vi... 127 5e-29
CO982036 124 3e-28
BG508993 117 5e-26
BU549979 116 7e-26
CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {V... 113 6e-25
BI321712 109 8e-24
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 958 bits (2476), Expect = 0.0
Identities = 475/979 (48%), Positives = 653/979 (66%)
Frame = +1
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI+G G + D P + VLL
Sbjct: 1678 EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHDGLPSLNKVLL 1857
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 1858 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 2037
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 2038 T-CLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 2214
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K E+
Sbjct: 2215 SHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSETFE 2394
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN +F S GI H+FS TPQQNG+VER
Sbjct: 2395 VFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAITPQQNGIVER 2574
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP++ +
Sbjct: 2575 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPSVKH 2754
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 2755 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 2934
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRITAAHPME 1087
+ + + E NV+D K+ E AE D +E+ + +RI HP E
Sbjct: 2935 SPARKKDVEEDVRTSGDNVADAAKSGENAE-NSDSATDESNINQPDKRSSTRIQKMHPKE 3111
Query: 1088 LIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSK 1147
LI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF +
Sbjct: 3112 LIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQFKR 3279
Query: 1148 NDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIA 1207
N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP+A
Sbjct: 3280 NEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPVA 3459
Query: 1208 RLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKS 1267
RLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D PDHV++LKK+
Sbjct: 3460 RLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADPTHPDHVYRLKKA 3639
Query: 1268 LYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQS 1327
LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 3640 LYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNE 3819
Query: 1328 LCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAK 1387
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QS+Y K ++KKF M ++ +
Sbjct: 3820 MLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKR 3999
Query: 1388 TPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLT 1447
TP L K++A V Q LYR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +HLT
Sbjct: 4000 TPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISHLT 4179
Query: 1448 AVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWE 1507
VKRIL+Y+ T++ G+MY S L GYCDAD+AG +RKSTSG C +LG+NL+SW
Sbjct: 4180 QVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSGGCFYLGNNLISWF 4359
Query: 1508 SKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKNP+
Sbjct: 4360 SKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKNPV 4539
Query: 1568 LHSRAKHIEVKYHFIRDYV 1586
HSR KHI++++H+IRD V
Sbjct: 4540 QHSRTKHIDIRHHYIRDLV 4596
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 954 bits (2466), Expect = 0.0
Identities = 476/981 (48%), Positives = 653/981 (66%), Gaps = 2/981 (0%)
Frame = +1
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ WYLDSGCSRHMTG + ++ V FG KGKI G G + D P + VLL
Sbjct: 1681 EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHDGLPSLNKVLL 1860
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N Y E +
Sbjct: 1861 VKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNCYLWTPQETSYSS 2040
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL S +E +WH+R GH +R + ++ VRG+PNLK +C CQ GK K+
Sbjct: 2041 T-CLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKM 2217
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDESHA 847
+ +TSR LELLH+DL GP++ ES+GGKRY ++VDD+SR+TWV F+ K ++
Sbjct: 2218 SHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSDTFE 2397
Query: 848 VFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
VF ++Q EK C I R+RSDHG EFEN KF S GI H+FS TPQQNG+VER
Sbjct: 2398 VFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVER 2577
Query: 908 KNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTLQE AR ML + + W EA+NTACYI NR+++R T YE+WK KP + +
Sbjct: 2578 KNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKH 2757
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFDDK 1027
FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T+ ESI+V DD
Sbjct: 2758 FHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDDL 2937
Query: 1028 LDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKS--RITAAHP 1085
+ + + E NV+D K+ E AE + +E P+ +Q K+ RI HP
Sbjct: 2938 TPARKKDVEEDVRTSGDNVADTAKSAENAENSDSATDE---PNINQPDKRPSIRIQKMHP 3108
Query: 1086 MELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQF 1145
ELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI AM+EEL QF
Sbjct: 3109 KELIIGDPNRGVTTRSR----EIEIVSNSCFVSKIEPKNVKEALTDEFWINAMQEELEQF 3276
Query: 1146 SKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAP 1205
+N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q EG+D+ ETFAP
Sbjct: 3277 KRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAP 3456
Query: 1206 IARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLK 1265
+ARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF D PDHV++LK
Sbjct: 3457 VARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHPDHVYRLK 3636
Query: 1266 KSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSAN 1325
K+LYGLKQAPRAWYERL+ FL + + +G +D TLF K +++++ QIYVDDI+FG +
Sbjct: 3637 KALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMS 3816
Query: 1326 QSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTV 1385
+ + F + MQ+EFEMS++GELTYFLG+QV Q + ++ QSKY K ++KKF M ++
Sbjct: 3817 NEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASH 3996
Query: 1386 AKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETH 1445
+TP L K++A V Q LYR MIGSLLYLTASRPDI ++V +CAR+Q++P+ +H
Sbjct: 3997 KRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISH 4176
Query: 1446 LTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVS 1505
L VKRIL+Y+ T++ G+MY S+ L GYCDAD+AG +RKSTSG C +LG+NL+S
Sbjct: 4177 LNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSADDRKSTSGGCFYLGTNLIS 4356
Query: 1506 WESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKN 1565
W SK+Q+ ++LSTAEAEYI+A +Q +WMK L++Y + + + +YCDN +AI++SKN
Sbjct: 4357 WFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINISKN 4536
Query: 1566 PILHSRAKHIEVKYHFIRDYV 1586
P+ HSR KHI++++H+IRD V
Sbjct: 4537 PVQHSRTKHIDIRHHYIRDLV 4599
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 229 bits (585), Expect = 6e-60
Identities = 108/153 (70%), Positives = 132/153 (85%)
Frame = -3
Query: 1136 LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQE 1195
+AM+EELNQF +N+VW LV+KPE+ VIGTKWVFRNKL+E G ++RNKARLVA+GY+Q+E
Sbjct: 461 IAMQEELNQFERNNVWKLVEKPENYPVIGTKWVFRNKLDEHGIIIRNKARLVAKGYNQEE 282
Query: 1196 GIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDE 1255
GIDY ET+AP+ARLE IR+L+++ N L+QMDVKSAFLNG I EEVYV QPPGFE
Sbjct: 281 GIDYEETYAPVARLEVIRMLLAYVSIMNFKLYQMDVKSAFLNGLIQEEVYVEQPPGFEIP 102
Query: 1256 KKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLE 1288
KP HV+KL+K+LYGLKQAPRAWYER+S+FLLE
Sbjct: 101 DKPTHVYKLQKALYGLKQAPRAWYERISNFLLE 3
>TC232995
Length = 1009
Score = 224 bits (571), Expect = 2e-58
Identities = 111/173 (64%), Positives = 133/173 (76%)
Frame = +2
Query: 1246 VHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTY 1305
V QPPGFE KP+HV+KL+K+LYGLKQAPRAWYERLS+FLLE EF RGKVDTTLF K
Sbjct: 2 VEQPPGFEISDKPNHVYKLQKALYGLKQAPRAWYERLSNFLLEKEFSRGKVDTTLFIKRK 181
Query: 1306 KDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYI 1365
+DIL+VQIYVDDIIFGS N SLCKEFS MQ+EFEMSMMGEL YFLG+Q+ QT G +I
Sbjct: 182 HNDILLVQIYVDDIIFGSTNDSLCKEFSLDMQSEFEMSMMGELKYFLGLQIKQTQ*GIFI 361
Query: 1366 HQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLL 1418
+QSKY KEL+K+F M + TPM C L+K+++ + K YR IG ++
Sbjct: 362 NQSKYCKELIKRFGMDSAKHMSTPMSTNCYLDKDESGQSIDIKQYRDAIGEVV 520
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 210 bits (534), Expect = 5e-54
Identities = 117/247 (47%), Positives = 156/247 (62%), Gaps = 2/247 (0%)
Frame = +3
Query: 1251 GFEDEKKPDHVFKLKKSL--YGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD 1308
GFED+++P HVF + L G+K ++ S ++ K+
Sbjct: 330 GFEDKERPCHVFMV*NKL*ELGMKG*VHF*FQMDSPEE*RTPHYSERLK--------KET 485
Query: 1309 ILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQS 1368
L++ IYVDDIIFG+ ++ +CKEF ++M+ FE SM GEL + LG+Q+ Q G +IHQ
Sbjct: 486 FLIIHIYVDDIIFGATSKRMCKEFFELMKDGFETSMKGELKFLLGLQIIQKVYGIFIHQE 665
Query: 1369 KYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDIL 1428
KYTK LK+F M E+ TPMH + I++K++ K Y GMI SL YLT+SRPDI+
Sbjct: 666 KYTKSHLKRFRMDEAKPMATPMHRSTIIDKDEKGNHTS*KEYSGMIDSLSYLTSSRPDIV 845
Query: 1429 FSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTE 1488
F V LCARFQS P+ +H+TAVKRILRYL TTN L +KK SE+ L GYCD +AGD+ E
Sbjct: 846 FVVCLCARFQSYPKISHVTAVKRILRYLVGTTNHCLWFKKRSEFDLLGYCDVYFAGDKVE 1025
Query: 1489 RKSTSGN 1495
RKSTS N
Sbjct: 1026 RKSTSRN 1046
>TC213445
Length = 705
Score = 123 bits (309), Expect(2) = 5e-47
Identities = 58/98 (59%), Positives = 76/98 (77%)
Frame = +1
Query: 1486 RTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI 1545
+T+R+STS C F+GS LVSW SK+Q+++ LSTAEAEYISA Q WM+ QL DY +
Sbjct: 400 KTDRESTSDTCHFIGSALVSWHSKKQNSVVLSTAEAEYISARSYYAQIFWMRQQLFDYGL 579
Query: 1546 LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIR 1583
+IPI CDNT+AI+LSKN IL+SR KHIE+++HF+R
Sbjct: 580 KLDHIPIRCDNTSAINLSKNHILYSRTKHIEIRHHFLR 693
Score = 84.7 bits (208), Expect(2) = 5e-47
Identities = 39/68 (57%), Positives = 50/68 (73%)
Frame = +2
Query: 1413 MIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEY 1472
MI S LYL+ SRP I+FSV +C R+Q++P+E+HL+ +KRI+RYL NLGL Y K S Y
Sbjct: 197 MIESFLYLSTSRPHIMFSVCMCVRYQANPKESHLSVIKRIMRYLLGIINLGLWYPKNSSY 376
Query: 1473 KLSGYCDA 1480
L GY DA
Sbjct: 377 NLVGYSDA 400
>BM143109
Length = 415
Score = 172 bits (436), Expect = 1e-42
Identities = 86/133 (64%), Positives = 103/133 (76%)
Frame = +1
Query: 1248 QPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKD 1307
QPP ++ +KP+HVFKLKK LYGLKQA RAWYE LS FLL+ F +GKVDT LF +
Sbjct: 4 QPPVRKNSEKPNHVFKLKKVLYGLKQALRAWYELLSKFLLDKGFSKGKVDTNLFI*KKLN 183
Query: 1308 DILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQ 1367
DIL+VQIYVDDIIFGS N SLCK+FS+ MQ EFEMSMM EL +FLG+Q+ QT G +I Q
Sbjct: 184 DILLVQIYVDDIIFGSTNDSLCKKFSQDMQNEFEMSMMRELNFFLGLQIKQTKNGIFISQ 363
Query: 1368 SKYTKELLKKFNM 1380
SKY K+L+ +F M
Sbjct: 364 SKYCKDLIHRFGM 402
>AI959950
Length = 466
Score = 168 bits (426), Expect(2) = 2e-41
Identities = 85/130 (65%), Positives = 102/130 (78%)
Frame = -1
Query: 1137 AMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEG 1196
AM+EEL+QF KN+V LVK P+ V+G KW+F NKL+E G VVR KARLVA+GYSQQEG
Sbjct: 391 AMQEELDQFQKNNV*KLVKLPKRKKVVGVKWIFCNKLDEDGKVVRYKARLVAKGYSQQEG 212
Query: 1197 IDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEK 1256
IDY +TFA +ARLE I +L+SF+ N+ L+QMDVKSAFLNG I +EVYV QPPGFE+E
Sbjct: 211 IDYPKTFALVARLEVICILLSFATYSNMKLYQMDVKSAFLNGLIQKEVYVEQPPGFENET 32
Query: 1257 KPDHVFKLKK 1266
HVFKL K
Sbjct: 31 LHQHVFKLNK 2
Score = 21.2 bits (43), Expect(2) = 2e-41
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = -2
Query: 1116 LVSLIEPKSIDEALQD 1131
L+ ++PK IDEA++D
Sbjct: 453 LIFEMKPKHIDEAIKD 406
>BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Glycine max},
partial (7%)
Length = 336
Score = 157 bits (398), Expect = 3e-38
Identities = 72/111 (64%), Positives = 95/111 (84%)
Frame = +3
Query: 1121 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPK+I EA+ D +WI+ M+EELNQF +N+VW LV+KPE+ VIGTKWVFRNKL+E G ++
Sbjct: 3 EPKNIKEAIVDDNWIIVMQEELNQFERNNVWKLVEKPENYPVIGTKWVFRNKLDEHGIII 182
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDV 1231
RNKARLVA+GY+Q+EGIDY ET+AP+ARLEAIR+L++++ N L+QMDV
Sbjct: 183 RNKARLVAKGYNQEEGIDYEETYAPVARLEAIRMLLAYASIMNFKLYQMDV 335
>CO983516
Length = 724
Score = 152 bits (383), Expect = 1e-36
Identities = 71/120 (59%), Positives = 92/120 (76%)
Frame = +2
Query: 1203 FAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVF 1262
F P+ARLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D PDHV+
Sbjct: 365 FHPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFIDPTHPDHVY 544
Query: 1263 KLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFG 1322
+LKK+LYGLKQAPRAWYERL+ L + + +G +D TLF K +++++ QIYVDDI+FG
Sbjct: 545 RLKKALYGLKQAPRAWYERLTELLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFG 724
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein, partial
(7%)
Length = 804
Score = 149 bits (376), Expect = 1e-35
Identities = 79/184 (42%), Positives = 119/184 (63%), Gaps = 4/184 (2%)
Frame = +1
Query: 1410 YRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYK-- 1467
+R +IGSL YL SRP+I F+V L +RF PR +H+ A KR+LR +K T G+++
Sbjct: 22 FRRLIGSLRYLCNSRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVLFPFK 201
Query: 1468 -KTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISA 1526
K+ + L GY D+D+ D + KST G V+ SK+Q IALST EAEY++A
Sbjct: 202 AKSGKPDLLGYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAEYVAA 381
Query: 1527 AICSTQKLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDY 1585
++ + Q +WM + LE+ ++ E + + DN +AI+L+K+P LH R+KHIE+++H+IRD
Sbjct: 382 SLGACQAVWMMNLLEELKLRERKPVNLLIDNKSAINLAKHPTLHGRSKHIELRFHYIRDQ 561
Query: 1586 V*KG 1589
V KG
Sbjct: 562 VSKG 573
>AI855982
Length = 484
Score = 148 bits (373), Expect = 2e-35
Identities = 78/165 (47%), Positives = 112/165 (67%)
Frame = +2
Query: 1085 PMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQ 1144
P++ I+G+ + V TR + + L + VS+IEPK+I EA+ D +WI+AM+EELNQ
Sbjct: 2 PLDNIIGDISKGVTTRHSLKD----LCNNMAFVSMIEPKNIKEAIVDDNWIIAMQEELNQ 169
Query: 1145 FSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFA 1204
F +N+VW LV+KP++ VI TKWVFRNKL+E ++ +KARLVA+GY+Q +G+DY T+A
Sbjct: 170 FERNNVWKLVEKPDNYPVI*TKWVFRNKLDEHRIIIIHKARLVAEGYNQVDGLDYEHTYA 349
Query: 1205 PIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQP 1249
IARL I + +S+ N L+ SA L+G + EVYV QP
Sbjct: 350 SIARL*VIIMPLSYVYIMNSTLYHYACVSALLHGLLLHEVYVDQP 484
>AI966222
Length = 430
Score = 112 bits (279), Expect(2) = 5e-30
Identities = 49/88 (55%), Positives = 63/88 (70%)
Frame = +1
Query: 914 EMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFHPFGC 973
EMART L + KHF E +N CY+QN+I +RPIL +TPYELWK KPNISYF+PF C
Sbjct: 1 EMARTTLNDNLTPKHF*AEVMNIVCYLQNKIYIRPILKRTPYELWKGRKPNISYFYPFRC 180
Query: 974 VCYVLNTKDRLHKFDAKSSKCLLLGYSE 1001
C+++NTKD L K D+KS + + YS+
Sbjct: 181 KCFIINTKDNLGKIDSKSDCGIFIAYSK 264
Score = 39.3 bits (90), Expect(2) = 5e-30
Identities = 22/51 (43%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +2
Query: 996 LLGYSERSKGFRFYNTDAKTIEESIHVRF-DDKLDSDQSKLVEKFADLSIN 1045
LL + SK FR YN+ IEE+IH+RF +K + + +L E FADL ++
Sbjct: 248 LLHTLKLSKAFRVYNSGTLVIEEAIHIRFGKNKPNKELLELDESFADLRLD 400
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment), partial
(30%)
Length = 687
Score = 127 bits (319), Expect = 4e-29
Identities = 57/113 (50%), Positives = 83/113 (73%), Gaps = 1/113 (0%)
Frame = +2
Query: 1473 KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQ 1532
+LSGYCDAD+AG +R+STSG C F+G NLVSW+SK+Q+ +A S+AEAEY S A+ + +
Sbjct: 14 QLSGYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCE 193
Query: 1533 KLWMKHQLEDYQILES-NIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRD 1584
+W+K L++ + E + +YCDN AA+ ++ NP+ H R KHIE+ HFIR+
Sbjct: 194 LMWIKQFLQELRFCEELQMKLYCDNQAALHIASNPVFHERTKHIEIDCHFIRE 352
>BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vinifera},
partial (34%)
Length = 407
Score = 127 bits (318), Expect = 5e-29
Identities = 60/130 (46%), Positives = 90/130 (69%)
Frame = -2
Query: 1154 VKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIR 1213
V P +G +WV+ K+ G+V R KARLVA+GY+Q GIDY +TF+P+A+L +R
Sbjct: 406 VPLPPGKTPVGCRWVYTVKVGPTGEVDRLKARLVAKGYTQVYGIDYCDTFSPVAKLTTVR 227
Query: 1214 LLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQ 1273
L ++ + + LHQ+D+K+AFL+G + E++Y+ QPPGF + + V KL +SLYGLKQ
Sbjct: 226 LFLAMAAICHWPLHQLDIKNAFLHGDLEEDIYMEQPPGFVAQGEYGLVCKLHRSLYGLKQ 47
Query: 1274 APRAWYERLS 1283
+PRAW+ + S
Sbjct: 46 SPRAWFGKFS 17
>CO982036
Length = 674
Score = 124 bits (311), Expect = 3e-28
Identities = 75/212 (35%), Positives = 119/212 (55%), Gaps = 5/212 (2%)
Frame = -2
Query: 1305 YKDDILVVQ--IYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEG 1362
YK IL V +YVD II GS+ +L + + + + F + ++G+L YF+ I+V P+
Sbjct: 673 YKTHILTVYLLVYVDIIITGSSC-TLIQNLTSKLNSSFPLKLLGKLDYFVEIEVKSMPDL 497
Query: 1363 TYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTA 1422
+ ++ + +K ++ +PM TC L K D+ YR ++G+L Y T
Sbjct: 496 LFSLRTSIFEIFCRKPR*QAQPIS-SPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTV 320
Query: 1423 SRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYK---KTSEYKLSGYCD 1479
RP+I F+V+ +F S+P ++H T VKRILRYLK + + GL K + + G+CD
Sbjct: 319 IRPEISFAVNKVCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAISSQPLPIRGFCD 140
Query: 1480 ADYAGDRTERKSTSGNCQFLGSNLVSWESKRQ 1511
AD+A +++STSG FLG NL+SW +Q
Sbjct: 139 ADWASAVDDKRSTSGAAVFLGPNLISWWXXKQ 44
>BG508993
Length = 374
Score = 117 bits (292), Expect = 5e-26
Identities = 52/123 (42%), Positives = 81/123 (65%), Gaps = 1/123 (0%)
Frame = +1
Query: 1457 KDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIAL 1516
K T + GL Y ++ YKL G+CD+D+AGD +RKST+G F+G + +W SK+Q + L
Sbjct: 4 KGTIDFGLFYSPSNNYKLVGFCDSDFAGDVDDRKSTTGFVFFMGDCVFTWSSKKQGIVTL 183
Query: 1517 STAEAEYISAAICSTQKLWMKHQLEDYQILE-SNIPIYCDNTAAISLSKNPILHSRAKHI 1575
T EAEY++A C+ +W++ LE+ Q+L+ + IY DN +A L+KN + H R+KHI
Sbjct: 184 FTCEAEYVAATSCTCHAIWLRRLLEELQLLQKESTKIYVDNRSAQELAKNSVFHERSKHI 363
Query: 1576 EVK 1578
+ +
Sbjct: 364 DTR 372
>BU549979
Length = 615
Score = 116 bits (291), Expect = 7e-26
Identities = 58/154 (37%), Positives = 96/154 (61%), Gaps = 3/154 (1%)
Frame = -1
Query: 1436 RFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGN 1495
R+QS+P H K+++RYL+ T + LMYK+T+ ++ GY D+D+AG R+STSG
Sbjct: 606 RYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYKQTNCLEVIGYSDSDFAGCVDSRRSTSGY 427
Query: 1496 CQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILES---NIPI 1552
L +VSW S +Q+ IA ST E E++ ++ +W+K + ++++S + +
Sbjct: 426 IFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSLRVVDSISRPLKL 247
Query: 1553 YCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
YCDN AA+ ++KN +R+KHI++KY IR+ V
Sbjct: 246 YCDNFAAVFMAKNNKSGNRSKHIDIKYLVIRERV 145
>CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {Vitis
vinifera}, partial (34%)
Length = 409
Score = 113 bits (283), Expect = 6e-25
Identities = 55/134 (41%), Positives = 81/134 (60%)
Frame = +3
Query: 1116 LVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNE 1175
L SL P +I EAL W AM +E+ N W LV P +G +WV+ K+
Sbjct: 3 LSSLTVPSTIREALDHPGWRQAMVDEMQALENNGTWELVPLPPGKTTVGCRWVYTVKVGP 182
Query: 1176 KGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAF 1235
G V R KARLVA+GY+Q GI+Y +TF+P+ L +RL ++ + + LHQ+D+K+AF
Sbjct: 183 NGKVDRLKARLVAKGYTQVYGIEYCDTFSPVFFLTTVRLFLAMAAIRHWPLHQLDIKNAF 362
Query: 1236 LNGYISEEVYVHQP 1249
L+G + E++Y+ QP
Sbjct: 363 LHGDLEEDIYMEQP 404
>BI321712
Length = 399
Score = 109 bits (273), Expect = 8e-24
Identities = 57/124 (45%), Positives = 78/124 (61%)
Frame = -3
Query: 1314 IYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKE 1373
+YVDD+IF N S+ +EF K M EFEM+ MG + Y+LGI+V Q +G +I Q Y KE
Sbjct: 379 LYVDDLIFTGNNPSMFEEFKKDMSNEFEMTDMGLMAYYLGIEVKQEDKGIFITQEGYAKE 200
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1433
+LKKF M ++ TPM L K + V LY+ +IGSL YLT +RPDIL+ V +
Sbjct: 199 VLKKFKMDDANPVGTPMECGSKLSKHEKGENVDPTLYKSLIGSLRYLTCTRPDILYVVGV 20
Query: 1434 CARF 1437
+R+
Sbjct: 19 VSRY 8
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.332 0.142 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,387,234
Number of Sequences: 63676
Number of extensions: 928536
Number of successful extensions: 7892
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 7651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7834
length of query: 1591
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1481
effective length of database: 5,635,272
effective search space: 8345837832
effective search space used: 8345837832
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0388a.6


