

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.4
(1489 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
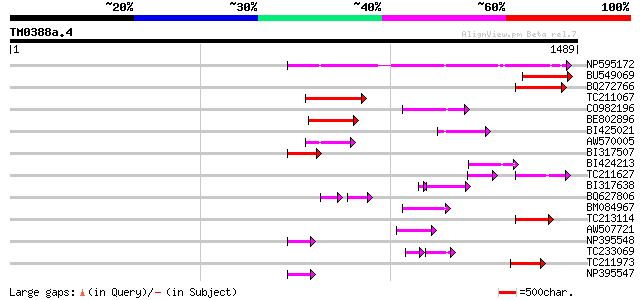
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 442 e-124
BU549069 158 2e-38
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 157 3e-38
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 147 3e-35
CO982196 113 5e-25
BE802896 107 4e-23
BI425021 105 1e-22
AW570005 99 2e-20
BI317507 86 2e-16
BI424213 83 1e-15
TC211627 77 7e-14
BI317638 weakly similar to GP|9294238|dbj| contains similarity t... 72 2e-12
BQ627806 52 4e-12
BM084967 69 1e-11
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 68 3e-11
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 67 5e-11
NP395548 reverse transcriptase [Glycine max] 65 2e-10
TC233069 58 3e-10
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 61 4e-09
NP395547 reverse transcriptase [Glycine max] 60 6e-09
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 442 bits (1138), Expect = e-124
Identities = 267/755 (35%), Positives = 407/755 (53%), Gaps = 9/755 (1%)
Frame = +1
Query: 730 KGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLA 789
K +L+A SKC F EV +LGH +S GV+++ +K+++++ W P +R F+GL
Sbjct: 2323 KQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLT 2502
Query: 790 GYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYD 849
GYYRRF+K YA + PLT L +K+ F W + E +F ++KK +T A +L+LP +P+
Sbjct: 2503 GYYRRFIKSYANIAGPLTDLLQKDS-FLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFI 2679
Query: 850 VYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSM 909
+ DAS G+G V+ Q IAY S++L + + EL AI AL +RHYL G+
Sbjct: 2680 LETDASGIGVGAVLGQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNK 2859
Query: 910 FTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSS 969
F I +D +SLK L DQ Q+ W+ Y+F ++ PGK N ADALSR M S
Sbjct: 2860 FIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIEYKPGKDNQAADALSRMFMLAWS 3039
Query: 970 LMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQ 1029
S L+E++++ +D L + + Q
Sbjct: 3040 -------------------------------EPHSIFLEELRARLISDPHLKQLMETYKQ 3126
Query: 1030 GKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWW 1089
G + +L K RV +P + + IL E H S + H G+ + LK F+W
Sbjct: 3127 GADASHYTVREGLLYWKDRVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLARLKAQFYW 3306
Query: 1090 PGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFD 1149
P M++ V Y+ CL C AK + PAG LQ L +P+ W+ ++MDF+T LP
Sbjct: 3307 PKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPLPIPQQVWEDVAMDFITGLP-NSFGLS 3483
Query: 1150 AIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGA 1209
I VV+DRLTK AHF+P+ +Y + +AE ++ IV+LHG+P SIVSDRD FTS FW
Sbjct: 3484 VIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQH 3663
Query: 1210 LQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSF 1269
L + GT L +SSAY+PQ+DGQ+E + LE LR +H W + LP EF YN ++
Sbjct: 3664 LFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAY 3843
Query: 1270 HASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SR--- 1326
H S+GM P+ ALYGR + P Q + P V++ + + K++I+ +R
Sbjct: 3844 HMSLGMTPFRALYGR--EPPTLTRQACS--IDDPAEVREQLTDRDALLAKLKINLTRAQQ 4011
Query: 1327 -QKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAI-KSKKLTPKFIGPYQITERVGPVAY 1384
K AD +R ++ FQ GD V +++ P + K++KL+ ++ GP+++ ++G VAY
Sbjct: 4012 VMKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAY 4191
Query: 1385 RIALPPFLSHIHDVLHVSQLRKY--MADDSHVLEPDDIQLKDDLTVVMPPIKIV-DRSTK 1441
++ LP + IH V HVSQL+ + A D ++ P + ++ VM P+KI+ R
Sbjct: 4192 KLELPS-AARIHPVFHVSQLKPFNGTAQDPYLPLPLTV---TEMGPVMQPVKILASRIII 4359
Query: 1442 RLRNK*VSLVKVVW-NQATGDATWELEDKMRESHP 1475
R N+ + + V W N +ATWE + ++ S+P
Sbjct: 4360 RGHNQ-IEQILVQWENGLQDEATWEDIEDIKASYP 4461
>BU549069
Length = 615
Score = 158 bits (399), Expect = 2e-38
Identities = 78/132 (59%), Positives = 102/132 (77%), Gaps = 1/132 (0%)
Frame = -1
Query: 1348 LRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKY 1407
L+VTP TGVG+A+KS+KLTP FIG +QI +R PVAY+IALPP LS++H+V HVSQLR Y
Sbjct: 615 LKVTPRTGVGQALKSRKLTPHFIGHFQILKRAXPVAYQIALPPSLSNLHNVFHVSQLRMY 436
Query: 1408 MADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWNQATG-DATWEL 1466
+ D SHV++ DD+Q+K++LT P++I DR TK LR K LVKV+W +G DATWEL
Sbjct: 435 IHDPSHVVKLDDVQVKENLTYETLPLRIEDRRTKHLRRKENPLVKVIWGGTSGEDATWEL 256
Query: 1467 EDKMRESHPDLF 1478
E +MR ++P LF
Sbjct: 255 ESQMRVAYPSLF 220
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo}, partial
(9%)
Length = 410
Score = 157 bits (397), Expect = 3e-38
Identities = 77/133 (57%), Positives = 101/133 (75%)
Frame = -3
Query: 1329 SYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIAL 1388
SY D RRK+LEF+ GDHVFLRVTP TGVGRA+KS KLTP FIGP+QI ++V VAY+IAL
Sbjct: 408 SYHDKRRKDLEFEVGDHVFLRVTP*TGVGRALKS*KLTPHFIGPFQILKKVDFVAYQIAL 229
Query: 1389 PPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*V 1448
PP L+ +H+V HVSQ+ KY+ D SH++ D +Q+K++L P++I D TK LR K +
Sbjct: 228 PPSLTSLHNVFHVSQIHKYINDPSHMVNLDVVQVKENLAHETLPLRIKDMRTKHLRGKEI 49
Query: 1449 SLVKVVWNQATGD 1461
LVKV+W A+G+
Sbjct: 48 LLVKVIWGGASGE 10
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 147 bits (372), Expect = 3e-35
Identities = 73/161 (45%), Positives = 104/161 (64%)
Frame = +1
Query: 777 KIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTA 836
K DIRSF GLA +YRRFV +++ + SPL +L KKN F W EK E++F +K++LT A
Sbjct: 106 KSVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKA 285
Query: 837 LILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVY 896
+LALP + +++ CDAS G+ V++Q IAY S +L NYPT+D EL A++
Sbjct: 286 PVLALPDFSKTFELECDASGVGVRAVLLQGGHPIAYFSEKLHSATLNYPTYDKELYALIR 465
Query: 897 ALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWME 937
A + W H+L F I+SDH+SLKY+ + LN R +W+E
Sbjct: 466 APQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWVE 588
>CO982196
Length = 812
Score = 113 bits (283), Expect = 5e-25
Identities = 70/176 (39%), Positives = 96/176 (53%)
Frame = +1
Query: 1031 KAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWP 1090
K P ++I L K R+ + ++T L+L E S L H G + ++ + +W
Sbjct: 289 KKPGYAIRGGK-LYFKDRLVLSKNSTKIPLLLKELQDSPLGGHSGFFRTFKRVANVVFWQ 465
Query: 1091 GMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDA 1150
GMKK +YV+ C C K PAG L L +P W ISMDF+ LP + + D
Sbjct: 466 GMKKTTRDYVAACEICRRNKTSTLSPAGLL*LLPIPTKVWTDISMDFIGGLPKAQGK-DN 642
Query: 1151 IWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRF 1206
I VVVDRLTK AHF +S Y +++AE+ I ++VRLHG P+SIVSD F S F
Sbjct: 643 ILVVVDRLTKYAHFFALSHPYTAKEVAELFIKELVRLHGFPASIVSDXXRLFMSLF 810
>BE802896
Length = 416
Score = 107 bits (267), Expect = 4e-23
Identities = 55/137 (40%), Positives = 85/137 (61%), Gaps = 4/137 (2%)
Frame = -2
Query: 784 SFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPK 843
SF+G AG+YRRF++D+ K+ PL+ L +K F + +KC+ +F +K+ L T I+ P
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPD 236
Query: 844 EGEPYDVYCDASHNGLGCVMMQE----KKVIAYASRQLKIHEKNYPTHDLELAAIVYALK 899
P+++ CDAS+ LG V+ Q+ +VI Y+SR L + NY T + EL AIV+AL+
Sbjct: 235 WTAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALE 56
Query: 900 IWRHYLYGSMFTIYSDH 916
+ YL G+ +Y DH
Sbjct: 55 KFHSYLLGTRIIVYIDH 5
>BI425021
Length = 426
Score = 105 bits (262), Expect = 1e-22
Identities = 57/140 (40%), Positives = 81/140 (57%)
Frame = -1
Query: 1123 LDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIA 1182
L VP+ W+ +SMDF+ LP I+VVV+R +K H + ++ +A + +
Sbjct: 417 LPVPQRPWEDLSMDFIVGLP-PYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASLFLN 241
Query: 1183 KIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDL 1242
+++LHG P SIVSDRD F S FW L + GT LR+SSAY+PQTDGQTE + +E
Sbjct: 240 IVIKLHGFPRSIVSDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQY 61
Query: 1243 LRACVLDHKGSWDELLPLIE 1262
LRA V + +P +E
Sbjct: 60 LRAFVHGRPRNLGRFIPWVE 1
>AW570005
Length = 413
Score = 98.6 bits (244), Expect = 2e-20
Identities = 52/132 (39%), Positives = 78/132 (58%)
Frame = -2
Query: 776 PKIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTT 835
P+ A +R F+ L G+YRRF+K YA + +PL+ L K+ F W+ + + +FQ +K +T
Sbjct: 406 PRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDS-FVWSPEADVAFQALKNVVTN 230
Query: 836 ALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIV 895
L+LALP +P+ V DAS + +G V+ QE IA+ S++ T+ ELAAI
Sbjct: 229 TLVLALPDFTKPFTVETDASGSDMGAVLSQEGHPIAFFSKEFCPKLVRSSTYVHELAAIT 50
Query: 896 YALKIWRHYLYG 907
+K WR YL G
Sbjct: 49 NVVKKWRQYLLG 14
>BI317507
Length = 359
Score = 85.5 bits (210), Expect = 2e-16
Identities = 43/90 (47%), Positives = 57/90 (62%), Gaps = 1/90 (1%)
Frame = -1
Query: 730 KGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLA 789
K + L AN KC F +++LGHVIS VA+D +K++S++ W PK + SF+ L
Sbjct: 284 KKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNKVKSVIEWPVPKNVKRVCSFLRLT 105
Query: 790 GYYRRFVKDYAKLT-SPLTQLTKKNQPFAW 818
GYYR+F+KDY KL PLT LT KN F W
Sbjct: 104 GYYRKFIKDYGKLAPRPLTDLT-KNDGFKW 18
>BI424213
Length = 426
Score = 82.8 bits (203), Expect = 1e-15
Identities = 50/133 (37%), Positives = 68/133 (50%)
Frame = +1
Query: 1204 SRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEF 1263
S FW L LGTKL S+ +PQTDGQT+ +SL LLRA + + SWDE LP +EF
Sbjct: 4 SHFWKTLWAKLGTKLLFSTTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEF 183
Query: 1264 TYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS 1323
YN H + +P+E +YG TPL D L + + + E +KM
Sbjct: 184 AYNRGVHRTTKQSPFEVVYGFNPLTPL----DLIPLPLDTSFIHKEGESRSEFVKKMH-- 345
Query: 1324 *SRQKSYADNRRK 1336
R K+ +N+ K
Sbjct: 346 -ERVKNQIENQTK 381
>TC211627
Length = 1034
Score = 76.6 bits (187), Expect = 7e-14
Identities = 40/78 (51%), Positives = 47/78 (59%)
Frame = +2
Query: 1202 FTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLI 1261
F S W L GTKLR S+AY+PQTDGQTE + LE LRA V DH W + L L
Sbjct: 5 FISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSLA 184
Query: 1262 EFTYNNSFHASIGMAPYE 1279
E YN S H+ IG +P+E
Sbjct: 185 E*CYNTSVHSGIGFSPFE 238
Score = 75.1 bits (183), Expect = 2e-13
Identities = 51/149 (34%), Positives = 74/149 (49%), Gaps = 4/149 (2%)
Frame = +3
Query: 1328 KSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIA 1387
K AD+ R++L F GD V++R+ P KL+ +F GPYQI RVG VAYR+
Sbjct: 384 KRIADSHRRDLTFNIGDWVYVRL*PYRQTSIQSTYTKLSKRFYGPYQIQARVGQVAYRLQ 563
Query: 1388 LPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMP---PIKIVDRSTKRLR 1444
LPP S IH + HVS L+ + + P+ + L T P P++ +D
Sbjct: 564 LPP-TSKIHPIFHVSLLKVHHGP----IPPELLALPPFSTTNHPLVQPLQFLDWKMDEST 728
Query: 1445 NK*VSLVKVVW-NQATGDATWELEDKMRE 1472
+ V V W N A D TWE ++++
Sbjct: 729 TPPIPQVLVQWTNLAPEDTTWESWTQLKD 815
>BI317638 weakly similar to GP|9294238|dbj| contains similarity to reverse
transcriptase~gene_id:K11J14.5 {Arabidopsis thaliana},
partial (5%)
Length = 420
Score = 72.0 bits (175), Expect(2) = 2e-12
Identities = 41/116 (35%), Positives = 61/116 (52%)
Frame = -2
Query: 1095 QVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVV 1154
+ A+ + CL C + K E ++ L L VP W+ +S+DF+T L L AI VV
Sbjct: 350 ECAQMLPNCLDCQHTKYETKRIVDLLCPLLVPHRPWEDLSLDFITGL-LPYHVHTAILVV 174
Query: 1155 VDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGAL 1210
VD +K H + ++ +A + I + +LHG+P S+VSD D F S FW L
Sbjct: 173 VDHFSKGIHLGMLPSSHTAHTVACLFIDSVAKLHGLPRSLVSDCDLLFVSHFWQEL 6
Score = 20.0 bits (40), Expect(2) = 2e-12
Identities = 8/26 (30%), Positives = 13/26 (49%)
Frame = -1
Query: 1073 HPGMNKMYQDLKLHFWWPGMKKQVAE 1098
H G+ K L + +W GM+ V +
Sbjct: 405 HTGIAKTLA*LSKNIYWFGMRTDVTQ 328
>BQ627806
Length = 435
Score = 51.6 bits (122), Expect(2) = 4e-12
Identities = 27/67 (40%), Positives = 38/67 (56%)
Frame = +1
Query: 886 THDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFD 945
T+ ELAAI A+K WR YL G F I +DH+SLK L Q Q+ ++ L +++
Sbjct: 232 TYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPEQQIYLARLMGFDYT 411
Query: 946 LQ*HPGK 952
+Q GK
Sbjct: 412 IQYRAGK 432
Score = 39.3 bits (90), Expect(2) = 4e-12
Identities = 18/57 (31%), Positives = 30/57 (52%)
Frame = +3
Query: 816 FAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAY 872
F W E+ + +F ++K L A +L LP + V DAS G+G ++ Q +A+
Sbjct: 21 FHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHHPLAF 191
>BM084967
Length = 426
Score = 69.3 bits (168), Expect = 1e-11
Identities = 45/125 (36%), Positives = 65/125 (52%)
Frame = -2
Query: 1033 PEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGM 1092
P+ S+ D IL+ G + +P+ + +L E H S H G+ K L +F W G+
Sbjct: 374 PDHSVIQDLILK-NGCIWLPSGFSFIPTLLLEYHSSPTDAHIGVTKTMARLSENFTWIGI 198
Query: 1093 KKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIW 1152
+K V ++V+ CL C K E QK AG L L VP W+ +S +F+ L R + AI
Sbjct: 197 RKDVEQFVAACLDCQYTKYEAQKMAGLLCPLPVPCRPWEDLSFNFIIGLS-EFRGYTAIL 21
Query: 1153 VVVDR 1157
VVV R
Sbjct: 20 VVVGR 6
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 68.2 bits (165), Expect = 3e-11
Identities = 39/101 (38%), Positives = 62/101 (60%), Gaps = 1/101 (0%)
Frame = +3
Query: 1328 KSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQITERVGPVAYRI 1386
K+YAD R+ + GD V+L++ P A K ++KL+P+F GPYQI +++G VA+ +
Sbjct: 6 KAYADRSRRAVTLSVGDWVYLKLQPYRLKSLAKKRNEKLSPRFYGPYQIKKQIGLVAFEL 185
Query: 1387 ALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLT 1427
LPP IH V H S L+K +A ++ +P + L +DL+
Sbjct: 186 DLPP-ARKIHPVFHASLLKKAVAATANP-QPLPLMLSEDLS 302
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 67.4 bits (163), Expect = 5e-11
Identities = 35/107 (32%), Positives = 62/107 (57%), Gaps = 3/107 (2%)
Frame = +1
Query: 1017 DEGLLEWRKLVTQ--GKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHP 1074
DE L++ +L + GK+ ++ + D I+ KGR+ +PND+ + ++I+ E H S++ H
Sbjct: 145 DESLMKIYQLCSNNAGKSGDYVLHHDVII-WKGRIMLPNDSQLLKMIMTESHASKVGGHA 321
Query: 1075 GMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQ-KPAGKL 1120
G + + F+WP M++ + ++V C+ C AKV H PAG L
Sbjct: 322 GTTRTIVRINAQFYWPKMREDIMKFVQECVICQQAKVTHSLLPAGLL 462
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 65.1 bits (157), Expect = 2e-10
Identities = 32/74 (43%), Positives = 43/74 (57%)
Frame = +1
Query: 729 CKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGL 788
C L N KC F + E LGH IS++G+ VD +KI+ I P IRSF+G
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 789 AGYYRRFVKDYAKL 802
A +YRRF+KD+ K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>TC233069
Length = 881
Score = 57.8 bits (138), Expect(2) = 3e-10
Identities = 34/82 (41%), Positives = 45/82 (54%), Gaps = 2/82 (2%)
Frame = +2
Query: 1092 MKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAI 1151
M + V +++ C C K QK G LQ L +P W ISMDFVT LP + + I
Sbjct: 224 MARDVCDHICACTNCQQNKYSTQK*FGLLQPLPIP*QVWKDISMDFVTHLPPS*GK-KMI 400
Query: 1152 WVVVDRLTKTAHF--VPISLNY 1171
WV+VD TK +HF +P L+Y
Sbjct: 401 WVIVDCWTKYSHFLSLPAHLSY 466
Score = 26.9 bits (58), Expect(2) = 3e-10
Identities = 14/50 (28%), Positives = 24/50 (48%)
Frame = +1
Query: 1040 DNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWW 1089
D +L + +P ++ +R ++L E H S L H G+ L + F W
Sbjct: 70 DGLLLYCTHLFIPLESELRAILLGECHDSLLGGHSGIKGTLNRLFVSFAW 219
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 60.8 bits (146), Expect = 4e-09
Identities = 33/92 (35%), Positives = 57/92 (61%), Gaps = 1/92 (1%)
Frame = +1
Query: 1316 IQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQ 1374
++E + S* + + RR+++E+ GD VFL++ P A + ++KL+P+F P+Q
Sbjct: 64 LRENLLKS*DIM*ANTNKRRRDIEYVVGD*VFLKMQPYRRRSLAKRINEKLSPRFYAPFQ 243
Query: 1375 ITERVGPVAYRIALPPFLSHIHDVLHVSQLRK 1406
+ +VG +AY++ LP + IH V HVS L+K
Sbjct: 244 VFNKVGTIAYKLDLPSHIK-IHPVFHVSLLKK 336
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 60.5 bits (145), Expect = 6e-09
Identities = 29/74 (39%), Positives = 41/74 (55%)
Frame = +1
Query: 729 CKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGL 788
C+ L N KC F ++E LGH IS +G+ V K++ I P I SF+G
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 789 AGYYRRFVKDYAKL 802
G+YRRF+KD+ K+
Sbjct: 721 VGFYRRFIKDFTKV 762
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.342 0.148 0.511
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,892,994
Number of Sequences: 63676
Number of extensions: 1100676
Number of successful extensions: 7814
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 7582
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7796
length of query: 1489
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1380
effective length of database: 5,698,948
effective search space: 7864548240
effective search space used: 7864548240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0388a.4


