

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.2
(422 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
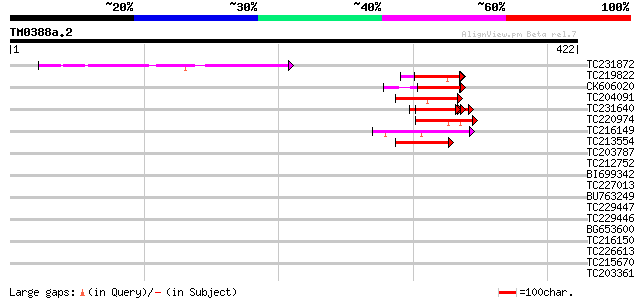
Score E
Sequences producing significant alignments: (bits) Value
TC231872 52 4e-07
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 46 4e-05
CK606020 46 4e-05
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 44 1e-04
TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated anti... 44 1e-04
TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus pro... 42 4e-04
TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 42 4e-04
TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protei... 42 5e-04
TC203787 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 41 0.001
TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific... 40 0.002
BI699342 40 0.002
TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1... 40 0.002
BU763249 40 0.002
TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, parti... 40 0.002
TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 39 0.003
BG653600 39 0.003
TC216150 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 39 0.004
TC226613 similar to UP|Q8H0V8 (Q8H0V8) Spliceosome associated pr... 39 0.006
TC215670 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, part... 38 0.008
TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 38 0.010
>TC231872
Length = 564
Score = 52.4 bits (124), Expect = 4e-07
Identities = 51/194 (26%), Positives = 84/194 (43%), Gaps = 4/194 (2%)
Frame = -2
Query: 22 FDRSRFLSAIKEDFYRAHLAHKECVKERGVLYREGRRDLLGMQEAMEIRRWKSWVNPIPF 81
+D RF SA+ + + A + ER V R+ + QE + RRW S V P+
Sbjct: 563 YDSHRFRSAVHQQRFEA-IKGWSFFGERRVQLRDD--EYTDFQEEIGRRRWTSLVTPMAK 393
Query: 82 ACEVMVREFYANTYCDNQEDKSRPPVNSSWFRGDVIDYNPAVIRRHLG---LLTEDQEKE 138
+V EFYAN + + + SW RG I ++ I + LG +L EDQE E
Sbjct: 392 FDPEIVLEFYANAWPTEEGVRDM----RSWVRGQWIPFDADAIGQLLGYPLVLKEDQECE 225
Query: 139 LFPEKATYHELLKEKSPEILEDMKAVIVRPGRDW-EAVDDEIIFVRRKDMTPLARVWAEF 197
Y + E + ++ PG D+ + + R +MT L ++W
Sbjct: 224 -------YGQRRNRSDGFDEEAIAQLLCIPG*DFARTAAGRRVRIMRTNMTTLTQIWMTL 66
Query: 198 LQDSLIPSSNKYEV 211
L +++PS + ++
Sbjct: 65 LLSNIMPSDHNSDL 24
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 45.8 bits (107), Expect = 4e-05
Identities = 23/39 (58%), Positives = 30/39 (75%), Gaps = 2/39 (5%)
Frame = +1
Query: 302 PQEDEEEEDPEEPAGEDKEEEEE--EEEEEENVEVVNEV 338
P+E+EEEE+ EE E++EEEEE EEEEEE +VNE+
Sbjct: 421 PEEEEEEEEEEEEENEEEEEEEEGEEEEEEEARSMVNEL 537
Score = 41.2 bits (95), Expect = 9e-04
Identities = 21/48 (43%), Positives = 29/48 (59%)
Frame = +1
Query: 292 QLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
QL + + + +EEEE+ EE E++EEEEEEE EEE E +V
Sbjct: 385 QL*LNYAPALAEPEEEEEEEEEEEEENEEEEEEEEGEEEEEEEARSMV 528
Score = 33.5 bits (75), Expect = 0.18
Identities = 24/85 (28%), Positives = 40/85 (46%)
Frame = +1
Query: 255 HLICSLIHAVRDRARRPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEP 314
H S I + A ++ + L AP L++ + +E +E+E EE+ EE
Sbjct: 322 HTSLSDITDLASNAMEDMNAFQL*LNYAPALAEPEEEEEEEE-----EEEEENEEEEEEE 486
Query: 315 AGEDKEEEEEEEEEEENVEVVNEVV 339
GE++EEEE E ++ N +V
Sbjct: 487 EGEEEEEEEARSMVNELAQLDNMLV 561
>CK606020
Length = 340
Score = 45.8 bits (107), Expect = 4e-05
Identities = 26/60 (43%), Positives = 36/60 (59%)
Frame = +1
Query: 279 LPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
LP +P+ + + I + +E+EEEE+ EE E++EEE EEEEEEE EV EV
Sbjct: 148 LPDSPVANDDEI-------DPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEV 306
Score = 43.5 bits (101), Expect = 2e-04
Identities = 22/36 (61%), Positives = 26/36 (72%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
E+EEEE+ EE E++EEEE EEEEEE E V E V
Sbjct: 199 EEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEV 306
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 44.3 bits (103), Expect = 1e-04
Identities = 25/52 (48%), Positives = 33/52 (63%), Gaps = 2/52 (3%)
Frame = +2
Query: 288 ERIAQLYKEWTARMPQEDEEEE--DPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E Q Y+E + +E+ EEE + EE ED+EEEEEEEEEEE +VV +
Sbjct: 107 ENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEENDVVKQ 262
Score = 35.8 bits (81), Expect = 0.037
Identities = 18/49 (36%), Positives = 27/49 (54%)
Frame = +2
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEE 328
P P E + + +E E+EEEE+ E+ E++EEEEEEE +
Sbjct: 104 PENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEEND 250
Score = 35.0 bits (79), Expect = 0.064
Identities = 22/62 (35%), Positives = 32/62 (51%)
Frame = +2
Query: 276 EKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+K L + P+ K ++A K + P +++ PE P E +E EEE EEEE EV
Sbjct: 5 KKTLVMPPMGRKRKVAP--KSSLSTEPPLKQQQPKPENPPQEYEEVEEEVEEEEVEEEVE 178
Query: 336 NE 337
E
Sbjct: 179 EE 184
Score = 30.4 bits (67), Expect = 1.6
Identities = 19/81 (23%), Positives = 43/81 (52%), Gaps = 2/81 (2%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKE--EEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFL 360
+EDEEEE+ EE E+ + +++++++++E+ +++++ P D + A +
Sbjct: 197 EEDEEEEEEEEEEEEENDVVKQQQQQQQDEDDIPISKLLEPLGKDQLLNLLCDAAAK--- 367
Query: 361 RQDRLHRDLDLHWRGGSTSDP 381
HRD++ R + DP
Sbjct: 368 -----HRDVEERIRKAADGDP 415
>TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated antigen, partial
(7%)
Length = 466
Score = 43.9 bits (102), Expect = 1e-04
Identities = 21/35 (60%), Positives = 26/35 (74%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
QE+EEE++ EE E++EEEEEEEEEEE E E
Sbjct: 22 QEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEE 126
Score = 43.9 bits (102), Expect = 1e-04
Identities = 20/33 (60%), Positives = 27/33 (81%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+E+EEEE+ EE E++EE+EEEEEEE NV V+
Sbjct: 52 EEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLVI 150
Score = 42.7 bits (99), Expect = 3e-04
Identities = 20/35 (57%), Positives = 26/35 (74%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+EE+E+ EE E++EEEEEEEEEEE E E
Sbjct: 25 EEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEE 129
Score = 42.4 bits (98), Expect = 4e-04
Identities = 20/37 (54%), Positives = 28/37 (75%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
+++EEEE+ EE E++EEEEEEE+EEE E N +V
Sbjct: 37 EKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLV 147
Score = 42.0 bits (97), Expect = 5e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Frame = +1
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
T +E EEEE+ EE E++EEEEEEEE+EE E V+
Sbjct: 19 TQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVL 144
Score = 41.6 bits (96), Expect = 7e-04
Identities = 20/43 (46%), Positives = 29/43 (66%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
+E+EEEE+ EE E++EEEEE+EEEEE V ++ + D
Sbjct: 43 EEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLVIIDEDKHD 171
Score = 38.1 bits (87), Expect = 0.008
Identities = 18/33 (54%), Positives = 23/33 (69%)
Frame = +1
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D +EE+ E+ E++EEEEEEEEEEE E E
Sbjct: 16 DTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKE 114
Score = 32.7 bits (73), Expect = 0.32
Identities = 17/33 (51%), Positives = 19/33 (57%)
Frame = +1
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
D D +E E +EEEEEEEEEEE E E
Sbjct: 1 DNVLSDTQEEEEEKEEEEEEEEEEEEEEEEEEE 99
>TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus protein p62,
partial (7%)
Length = 436
Score = 42.4 bits (98), Expect = 4e-04
Identities = 24/56 (42%), Positives = 37/56 (65%), Gaps = 10/56 (17%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEE------EEEEENVEV----VNEVVTPPRADPSP 348
Q+++EEED E +GE++EE+EEE EEEEE+ ++ V++ PP A+P P
Sbjct: 166 QKNKEEEDEEVSSGEEEEEDEEEEEAASSEEEEEDEDLPPPPVSKNPPPPPANPQP 333
Score = 32.3 bits (72), Expect = 0.41
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 4/58 (6%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEE----EEEENVEVVNEVVT 340
K R + L + TA +EEE ++P+ + +EEE+EE EEEE E E +
Sbjct: 76 KLRPSPLDEPPTASSSDSEEEEPQQQQPSSQKNKEEEDEEVSSGEEEEEDEEEEEAAS 249
>TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (4%)
Length = 2040
Score = 42.4 bits (98), Expect = 4e-04
Identities = 28/88 (31%), Positives = 44/88 (49%), Gaps = 12/88 (13%)
Frame = +3
Query: 271 PVHVIEKR--------LPVAPLLSKERIAQLYKEWTARMPQED----EEEEDPEEPAGED 318
PV+V+ K L V P E+ + KE P+++ +EEE EEP E+
Sbjct: 1164 PVNVVSKLRKYWQTDILSVGPAKEPEKKEEPKKEEKKEEPKKEGEAKKEEEKKEEPKKEE 1343
Query: 319 KEEEEEEEEEEENVEVVNEVVTPPRADP 346
+++EE ++EEE+ E + PP DP
Sbjct: 1344 EKKEEPKKEEEKKEEEKKKEAPPPPPDP 1427
>TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1,
partial (18%)
Length = 451
Score = 42.0 bits (97), Expect = 5e-04
Identities = 20/43 (46%), Positives = 29/43 (66%)
Frame = +2
Query: 288 ERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
E AQ Y+E +E+E EE+ EE E+++EE+EEEEEE+
Sbjct: 170 ENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEED 298
Score = 38.5 bits (88), Expect = 0.006
Identities = 21/42 (50%), Positives = 27/42 (64%)
Frame = +2
Query: 296 EWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E A+ +E+ EEE EE E+ EEEEEEEE+EE+ E E
Sbjct: 170 ENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEEE 295
Score = 36.6 bits (83), Expect = 0.022
Identities = 20/43 (46%), Positives = 25/43 (57%)
Frame = +2
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
K A E+E EE+ EE E++ EEEEEEEE+E E E
Sbjct: 164 KPENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDEEEEE 292
Score = 30.8 bits (68), Expect = 1.2
Identities = 17/36 (47%), Positives = 20/36 (55%)
Frame = +2
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
P + + PE A E +EE EEE EEEE E V E
Sbjct: 140 PPLKQHQPKPENAAQEYEEEVEEEVEEEEVEEEVEE 247
Score = 29.3 bits (64), Expect = 3.5
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +2
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+ + ++ EE E+ EEEE EEE EE E
Sbjct: 161 PKPENAAQEYEEEVEEEVEEEEVEEEVEEEEE 256
Score = 28.5 bits (62), Expect = 5.9
Identities = 15/37 (40%), Positives = 20/37 (53%)
Frame = +2
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ Q + E+ + E+ EEE EEEE EE VE E
Sbjct: 146 LKQHQPKPENAAQEYEEEVEEEVEEEEVEEEVEEEEE 256
Score = 28.5 bits (62), Expect = 5.9
Identities = 16/54 (29%), Positives = 28/54 (51%)
Frame = +2
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P+ +L+ R ++ + + ++ + E A ++ EEE EEE EEE VE
Sbjct: 71 PINQILAMGRKRKVAPKSSVSAEPPLKQHQPKPENAAQEYEEEVEEEVEEEEVE 232
>TC203787 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 1056
Score = 40.8 bits (94), Expect = 0.001
Identities = 16/40 (40%), Positives = 28/40 (70%)
Frame = -3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
ED++++D E+ AGE+++E+ E+EE E E E + PP+
Sbjct: 406 EDDDDDDEEDEAGEEEDEDGAEDEENEEDEEDEEALQPPK 287
>TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein T2
precursor, partial (35%)
Length = 533
Score = 40.4 bits (93), Expect = 0.002
Identities = 20/38 (52%), Positives = 24/38 (62%)
Frame = +3
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
T+ Q+ EEEE+P +D E EEEEEEEEE E V
Sbjct: 9 TSSDKQKQEEEEEPHRMEEDDDESEEEEEEEEEEEEEV 122
Score = 33.5 bits (75), Expect = 0.18
Identities = 14/35 (40%), Positives = 24/35 (68%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
+ +++E+ EEP +++++E EEEEEE E EV
Sbjct: 18 DKQKQEEEEEPHRMEEDDDESEEEEEEEEEEEEEV 122
Score = 33.1 bits (74), Expect = 0.24
Identities = 19/43 (44%), Positives = 27/43 (62%), Gaps = 5/43 (11%)
Frame = +3
Query: 300 RMPQEDEEE-----EDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ QE+EEE ED +E E++EEEEEEEE ++ E +E
Sbjct: 21 KQKQEEEEEPHRMEEDDDESEEEEEEEEEEEEEVWDDWEGEDE 149
>BI699342
Length = 325
Score = 40.4 bits (93), Expect = 0.002
Identities = 20/42 (47%), Positives = 31/42 (73%)
Frame = -1
Query: 293 LYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEV 334
++K T+ +E+EEEE+ EE E++EEEEEEEEEE+ + +
Sbjct: 292 IFK**TSNAEEEEEEEEEEEE---EEEEEEEEEEEEEKAINL 176
Score = 31.2 bits (69), Expect = 0.92
Identities = 15/23 (65%), Positives = 17/23 (73%)
Frame = -1
Query: 315 AGEDKEEEEEEEEEEENVEVVNE 337
A E++EEEEEEEEEEE E E
Sbjct: 268 AEEEEEEEEEEEEEEEEEEEEEE 200
Score = 31.2 bits (69), Expect = 0.92
Identities = 18/32 (56%), Positives = 20/32 (62%)
Frame = -1
Query: 306 EEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
EEEE E++EEEEEEEEEEE E E
Sbjct: 265 EEEE-------EEEEEEEEEEEEEEEEEEEEE 191
>TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1 regulatory
subunit GAC1, partial (3%)
Length = 1149
Score = 40.4 bits (93), Expect = 0.002
Identities = 24/52 (46%), Positives = 32/52 (61%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAF 355
E++ E + EE ED+ EEEEEEEEEE E + V+T +D SP + AF
Sbjct: 188 EEKVENENEEKHNEDEIEEEEEEEEEEEEEEFSFVLT--NSDGSPISADDAF 337
Score = 36.6 bits (83), Expect = 0.022
Identities = 18/43 (41%), Positives = 27/43 (61%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
+ + EE+ E+ E++EEEEEEEEEE + + N +P AD
Sbjct: 200 ENENEEKHNEDEIEEEEEEEEEEEEEEFSFVLTNSDGSPISAD 328
>BU763249
Length = 412
Score = 40.4 bits (93), Expect = 0.002
Identities = 33/118 (27%), Positives = 52/118 (43%)
Frame = +3
Query: 12 AAASSSTSRSFDRSRFLSAIKEDFYRAHLAHKECVKERGVLYREGRRDLLGMQEAMEIRR 71
++A+ FD RF S + + A + +KER V REG + Q+ + R+
Sbjct: 69 SSAAPQADIEFDGHRFRSEKHQRCFEA-IKS*AFLKERRVQLREG--EYTEFQKEIARRK 239
Query: 72 WKSWVNPIPFACEVMVREFYANTYCDNQEDKSRPPVNSSWFRGDVIDYNPAVIRRHLG 129
W P+ +V EFYAN + + K + SW RG I ++ I + LG
Sbjct: 240 WTQLAGPMAKYDPKIVMEFYANAWPTEEGVKDK----RSWVRGQWIPFDEDAINQFLG 401
>TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, partial (4%)
Length = 582
Score = 40.0 bits (92), Expect = 0.002
Identities = 24/85 (28%), Positives = 40/85 (46%)
Frame = +3
Query: 270 RPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEE 329
+PV ++ L ++ I + E + M ++ + E EEP + E+E EEEEEE
Sbjct: 141 KPVEPQQQNHETVELDQQQPIPEPTLEDSNTMAIDEPKHEQEEEPQNAESEQEVEEEEEE 320
Query: 330 ENVEVVNEVVTPPRADPSPAWMEAA 354
E+ + + P P +EAA
Sbjct: 321 EDEQQEEQEPETREGQPGPETLEAA 395
>TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(20%)
Length = 850
Score = 39.3 bits (90), Expect = 0.003
Identities = 16/39 (41%), Positives = 26/39 (66%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
+E+EE+E+P+ E++EEE EE++EE E + E P
Sbjct: 252 EEEEEDEEPQNAESEEEEEEAEEQQEERPPETLEEQQDP 368
Score = 38.1 bits (87), Expect = 0.008
Identities = 16/29 (55%), Positives = 22/29 (75%)
Frame = +3
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
P+++EEEED E E +EEEEE EE++E
Sbjct: 243 PKQEEEEEDEEPQNAESEEEEEEAEEQQE 329
Score = 33.9 bits (76), Expect = 0.14
Identities = 23/85 (27%), Positives = 43/85 (50%), Gaps = 1/85 (1%)
Frame = +3
Query: 254 PHLICSLIHAVRDRARRPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEE 313
P+L+ ++ + R+ P E P+ P + +L ++ + P ++ EDP
Sbjct: 66 PYLLAAMAKKRKLRSSDP----EPTKPIEPQQQDQEPVELDQQ--QQQPNPEQTLEDPNT 227
Query: 314 PAGEDKEEEEEEEEEE-ENVEVVNE 337
A ++ ++EEEEE+EE +N E E
Sbjct: 228 MAVDEPKQEEEEEDEEPQNAESEEE 302
Score = 32.7 bits (73), Expect = 0.32
Identities = 32/111 (28%), Positives = 48/111 (42%), Gaps = 21/111 (18%)
Frame = +3
Query: 286 SKERIAQLYKEWTARMPQEDEEEEDPEEPAG----------EDKEEEEEEEE---EEENV 332
S+E + ++ R P+ EE++DPE A E EEEEEE+ EEE V
Sbjct: 291 SEEEEEEAEEQQEERPPETLEEQQDPETLAAANHAEVNGGNEGNNEEEEEEDLTLEEEPV 470
Query: 333 EVVNEVVTPPR--------ADPSPAWMEAAFGRMFLRQDRLHRDLDLHWRG 375
E + E T + D P ++++ R D HR + +H G
Sbjct: 471 EKLLEPFTKEQLHSLVTQAVDMFPEFVDSV--RRLADVDPAHRKIFVHGLG 617
>BG653600
Length = 411
Score = 39.3 bits (90), Expect = 0.003
Identities = 30/97 (30%), Positives = 48/97 (48%), Gaps = 3/97 (3%)
Frame = -1
Query: 2 PVKIARTNTR---AAASSSTSRSFDRSRFLSAIKEDFYRAHLAHKECVKERGVLYREGRR 58
P K+A +R AA +S++ +D RF SA+ + + A + ++ER V R+
Sbjct: 381 PRKLASKRSRKDKAAEGTSSAPEYDSHRFRSAVHQQRFEA-IKGWSFLRERRVQLRDD-- 211
Query: 59 DLLGMQEAMEIRRWKSWVNPIPFACEVMVREFYANTY 95
+ QE + RRW V P+ +V EFYAN +
Sbjct: 210 EYTDFQEEIGRRRWAPLVTPMAKFDPEIVLEFYANAW 100
>TC216150 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (5%)
Length = 928
Score = 38.9 bits (89), Expect = 0.004
Identities = 19/52 (36%), Positives = 30/52 (57%)
Frame = +3
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADP 346
KE + + +EEE EEP E++++EE ++EEE+ E + PP DP
Sbjct: 438 KEEPKKEGEAKKEEEKKEEPKKEEEKKEETKKEEEKKEEEKKKEAPPPPPDP 593
>TC226613 similar to UP|Q8H0V8 (Q8H0V8) Spliceosome associated protein-like
(At4g21660), partial (59%)
Length = 1844
Score = 38.5 bits (88), Expect = 0.006
Identities = 18/44 (40%), Positives = 28/44 (62%)
Frame = +2
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+EEEE+ EE E++EEE EEEE E ++ V+ + + P +P
Sbjct: 863 EEEEEEEEEEEEEEEEEEMEEEELEAGIQSVDSLSSTPTGVETP 994
Score = 34.3 bits (77), Expect = 0.11
Identities = 17/32 (53%), Positives = 21/32 (65%)
Frame = +2
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
P +EE D + G+ +EEEEEEEEEEE E
Sbjct: 812 PNYEEEPVDKTKHWGDLEEEEEEEEEEEEEEE 907
Score = 32.0 bits (71), Expect = 0.54
Identities = 19/48 (39%), Positives = 27/48 (55%)
Frame = +2
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWME 352
D EEE+ EE ++EEEEEEEEE E E+ + + +P +E
Sbjct: 857 DLEEEEEEE----EEEEEEEEEEEMEEEELEAGIQSVDSLSSTPTGVE 988
Score = 30.0 bits (66), Expect = 2.0
Identities = 17/39 (43%), Positives = 21/39 (53%), Gaps = 4/39 (10%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKE----EEEEEEEEEENVEVVNE 337
Q+ ++ EEP + K EEEEEEEEEE E E
Sbjct: 797 QQQDQPNYEEEPVDKTKHWGDLEEEEEEEEEEEEEEEEE 913
>TC215670 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, partial (34%)
Length = 809
Score = 38.1 bits (87), Expect = 0.008
Identities = 21/56 (37%), Positives = 26/56 (45%), Gaps = 5/56 (8%)
Frame = +2
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEE-----EEEEEENVEVVNEVVTPPRADPSP 348
T R+ EEE PEEP ++K EEE + E+E EV PP P P
Sbjct: 110 TRRIAPMGEEENKPEEPMAKEKNPEEEATQEKKPEQESKEEVAAAAAAPPPPPPPP 277
>TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 1090
Score = 37.7 bits (86), Expect = 0.010
Identities = 18/41 (43%), Positives = 26/41 (62%)
Frame = -1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPR 343
+EDE E+ +E ED+E EE+EE+EEE E + PP+
Sbjct: 349 EEDEGGEEEDEDGAEDEENEEDEEDEEE------EALQPPK 245
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,851,639
Number of Sequences: 63676
Number of extensions: 279774
Number of successful extensions: 5013
Number of sequences better than 10.0: 233
Number of HSP's better than 10.0 without gapping: 2298
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3563
length of query: 422
length of database: 12,639,632
effective HSP length: 100
effective length of query: 322
effective length of database: 6,272,032
effective search space: 2019594304
effective search space used: 2019594304
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0388a.2


