

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.1
(728 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
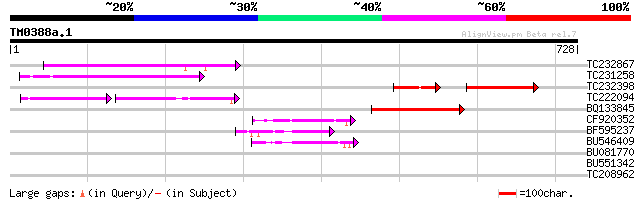
Score E
Sequences producing significant alignments: (bits) Value
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 144 2e-34
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 125 7e-29
TC232398 87 4e-27
TC222094 74 1e-24
BQ133845 107 2e-23
CF920352 59 1e-08
BF595237 57 4e-08
BU546409 55 1e-07
BU081770 40 0.003
BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [impor... 30 5.0
TC208962 UP|Q9ARG2 (Q9ARG2) Amino acid transporter, complete 29 8.5
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 144 bits (362), Expect = 2e-34
Identities = 94/267 (35%), Positives = 142/267 (52%), Gaps = 14/267 (5%)
Frame = -3
Query: 44 PALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEW 103
P+LI L+ N + G EDP+AH+ +I CNT R+ V D IRLSL FSL A+ W
Sbjct: 815 PSLIQLIQSNFFHGLPNEDPYAHLATYIEICNTIRLAGVPADAIRLSLLSFSLSGEAKRW 636
Query: 104 LNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRR 163
L+S S+ SW+++ EKF ++ P + + K I +F Q DE+L EA ERF+ LLR+
Sbjct: 635 LHSFKGNSLKSWDEVVEKFLKKYFPESKTAEGKAAISSFHQFPDESLSEALERFRGLLRK 456
Query: 164 CPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSND 223
P H ++ Q+ F D L+ S+ +DA+ G+ P +LIE MA + D
Sbjct: 455 TPTHGFSEPIQLNIFIDELRPESKQLVDASVGGKIKMKTPDEAMDLIESMAASDIAILRD 276
Query: 224 R---QARRGVLEVEAYDQLMASNKQLSKQ-------MNEIQNQMKTTKIG-SRVAKVEFV 272
R ++ +LE+ + D L+A NK LSKQ ++++ Q+ + + S + +V
Sbjct: 275 RAHIPTKKSLLELTSQDTLLAQNKLLSKQLETLTKTLSKLPTQLHSAQTSHSSILQVTGC 96
Query: 273 TC-GGPHDSEEC--TETRPEEEVKAMG 296
T G H+S C E + EV MG
Sbjct: 95 TIFGEAHESGCCIPNEEQIAHEVNYMG 15
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 125 bits (314), Expect = 7e-29
Identities = 81/242 (33%), Positives = 121/242 (49%), Gaps = 4/242 (1%)
Frame = +2
Query: 13 RSLRDLTSAAMSYDYPGSIVS--PDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERF 70
R+LR++ + +Y+ S+ + PD + L+ LI+L+ + + G EDPH H++ F
Sbjct: 437 RTLREMVAPDFTYE---SLCNQYPDEDVPYVLKTGLIHLLPK--FHGLVGEDPHKHLKEF 601
Query: 71 IRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRA 130
C+T + +V D I L FP SL A++WL SI +W+DL F +F P +
Sbjct: 602 HIVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRSIFNWDDLKRVFLEKFFPAS 781
Query: 131 LLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGL 190
++ DI Q + E+LYE WERFKK CP H +++ + FY+ L R +
Sbjct: 782 RTTTIRKDISGIRQLSRESLYEYWERFKKSCASCPHHQISKQLLL*YFYEELSNMKRSMI 961
Query: 191 DAASSGEFDALLPQVGYELIEKMAIRAMNSS--NDRQARRGVLEVEAYDQLMASNKQLSK 248
DAAS G + P LI+KMA + S ND RGV EV NK L +
Sbjct: 962 DAASGGALGDMTPTEARNLIKKMASNSQQFSARNDAIVLRGVHEVATDSSSSTENKSLRE 1141
Query: 249 QM 250
+
Sbjct: 1142NL 1147
>TC232398
Length = 1054
Score = 87.0 bits (214), Expect(2) = 4e-27
Identities = 39/92 (42%), Positives = 65/92 (70%)
Frame = +1
Query: 587 RLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILG 646
R+ ++ PT ++LQ+AD S+ +G+VED++V V + +F VDFV++D+EED +I LILG
Sbjct: 175 RIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLILG 354
Query: 647 RPFLATGRAKIDVDKGHLILPVGKEKVRFSVF 678
PF+ T + +D+ KG+L + V +K F++F
Sbjct: 355 WPFMVTAKCVVDMGKGNLEMSVEDQKATFNLF 450
Score = 53.5 bits (127), Expect(2) = 4e-27
Identities = 27/60 (45%), Positives = 42/60 (70%)
Frame = +2
Query: 494 LHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRRDPG 553
L I+IPF + ++QMP+Y KF+KDIL K+ + ET+++ E C A++Q K+P K +D G
Sbjct: 2 LEITIPFGERIQQMPLYKKFLKDILIKKGKYIN-SETIVVGEYCRALIQ-KLPPKFKDLG 175
>TC222094
Length = 984
Score = 73.9 bits (180), Expect(2) = 1e-24
Identities = 42/118 (35%), Positives = 61/118 (51%), Gaps = 1/118 (0%)
Frame = -2
Query: 14 SLRDLTSAAMSYDYPGSIVSPD-GTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIR 72
+L D +S + Y SI P+ T NF +LI L+ N + G EDP+AH+ +I
Sbjct: 845 TLEDYSSPIIP-QYFTSIARPEVQTANFSYPYSLIQLIQGNLFYGLPTEDPYAHLATYID 669
Query: 73 NCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRA 130
CNT ++ V D I L LF FSL A WL S ++ +W + EKF ++ P +
Sbjct: 668 ICNTVKIVGVPEDAIHLDLFCFSLAGEARTWLRSFKGNNLRTWNEXXEKFLKKYFPES 495
Score = 58.2 bits (139), Expect(2) = 1e-24
Identities = 47/163 (28%), Positives = 77/163 (46%), Gaps = 3/163 (1%)
Frame = -3
Query: 136 KNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASS 195
K +I +F Q E+L EA +RF LL + P H ++ Q+ F DG+Q S+ LDA++
Sbjct: 478 KVEISSFHQHPHESLSEALDRFHGLLWKTPTHGFSEPVQLNIFIDGMQPHSKQLLDASAG 299
Query: 196 GEFDALLPQVGYELIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQN 255
G+ P+ ELIE MA +ND R E ++ + L + M E ++
Sbjct: 298 GKIKLKTPEEAIELIENMA------ANDYVILRD-QEPSPQEESTRLEELLVQFMQETRS 140
Query: 256 QMKTTKIGSRVAKVEFVTCGGPHDSEEC---TETRPEEEVKAM 295
K+T R +V+ +E T+ +P++E K +
Sbjct: 139 HQKSTDAAIRNLEVQLAKLVPERPTETFAVNTKMKPKKECKVI 11
Score = 38.9 bits (89), Expect = 0.008
Identities = 25/64 (39%), Positives = 37/64 (57%)
Frame = -3
Query: 362 LEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASV 421
LEEL+ F+ T ++ K+ +AA +NLE QLAK + ERP F + PK+E V
Sbjct: 181 LEELLVQFMQETRSHQKSTDAAIRNLE---VQLAKLVPERPTETFAVNTKMKPKKE-CKV 14
Query: 422 VATR 425
+ T+
Sbjct: 13 ILTK 2
>BQ133845
Length = 389
Score = 107 bits (267), Expect = 2e-23
Identities = 54/120 (45%), Positives = 82/120 (68%)
Frame = +2
Query: 465 ETSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRL 524
E ++P+P + K ++ +KF+D+FKKL I++PF +AL+QMP+YA F+KD+L K+
Sbjct: 23 ECKEVPYPLVPS*KDKEQHLAKFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNWY 202
Query: 525 KGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTM 584
D+ +++ CSA++QR +P DPG T+P I +A +AL DLGASINLMPL+M
Sbjct: 203 IHSDK-IVVEGNCSAVIQRILPP*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSM 379
>CF920352
Length = 571
Score = 58.5 bits (140), Expect = 1e-08
Identities = 48/136 (35%), Positives = 68/136 (49%), Gaps = 4/136 (2%)
Frame = -2
Query: 312 RNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFIN 371
RNHP GN F G QG S +PQ++G + K LEE + F+
Sbjct: 477 RNHP-----------GNQFN------GDQGGSSIRPQQQGSSLYDRTTK-LEETLAQFM* 352
Query: 372 RTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVM- 430
+ +N K+ E+A KNLE Q GQLAK++A+R F ++ NPK+E V TRS
Sbjct: 351 VSMSNQKSTESAIKNLEVQVGQLAKKLADRSSSSFSANTEKNPKEE-CKAVMTRSKMTTH 175
Query: 431 ---SELKKKTEGEKRE 443
+ +KK E K++
Sbjct: 174 VDEGKAEKKMEEHKQQ 127
>BF595237
Length = 415
Score = 56.6 bits (135), Expect = 4e-08
Identities = 46/135 (34%), Positives = 64/135 (47%), Gaps = 8/135 (5%)
Frame = +3
Query: 291 EVKAMGQARNDPFSNTYN----PGWRNHPNF----SWRQGSSGQGNGFQRQFPSQGFQGQ 342
EV MG + F YN PG+ NF SW+ Q N QR P Q Q
Sbjct: 33 EVNYMGAQNHHGFQG-YNQGGPPGFNQGRNFTQG*SWKNHPGNQFNKEQRSQPIQN-SNQ 206
Query: 343 SLRQPQERGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAERP 402
+ ++ + LEE + F+ +NY + +++ KNLE Q GQLAKQ+AERP
Sbjct: 207 VVNLYEKTSK---------LEETLNQFM*MFMSNYMSTKSSIKNLEIQMGQLAKQMAERP 359
Query: 403 QGIFPSDCIPNPKQE 417
F ++ NPK+E
Sbjct: 360 TNNFGANTEKNPKKE 404
>BU546409
Length = 528
Score = 55.1 bits (131), Expect = 1e-07
Identities = 48/149 (32%), Positives = 72/149 (48%), Gaps = 12/149 (8%)
Frame = -1
Query: 311 WRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFI 370
WR+HP+ + + G N P Q QG ++ Q + LEE + F+
Sbjct: 420 WRSHPSNQFNKDQGGPSNK-----PIQ--QGPNIFQRTXK-----------LEETLTQFM 295
Query: 371 NRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGR-- 428
+N+K+ E+A +NLE Q GQLAKQI ++ F + NPK+E V+ TRS R
Sbjct: 294 KVIMSNHKSTESALENLEVQVGQLAKQITDKSSNSFMVNTEKNPKEECKDVM-TRSKRFV 118
Query: 429 -------VMSELK---KKTEGEKREEIVG 447
V+S+ K KK EK+ ++ G
Sbjct: 117 EAEDEDNVVSKKKFAEKKGTDEKKNDVRG 31
>BU081770
Length = 426
Score = 40.4 bits (93), Expect = 0.003
Identities = 30/90 (33%), Positives = 46/90 (50%), Gaps = 3/90 (3%)
Frame = +1
Query: 342 QSLRQPQERGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAER 401
Q +QPQ+ E + SLEELV + + +A+ ++L NQ GQLA Q+ ++
Sbjct: 97 QQQQQPQK*QTVEAP-PQPSLEELVRQMTMQNIQFQQETKASIQSLTNQMGQLATQLNQQ 273
Query: 402 P---QGIFPSDCIPNPKQENASVVATRSGR 428
PS + NPK N S ++ RSG+
Sbjct: 274 QSQNSDKLPSQAVQNPK--NVSAISLRSGK 357
>BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [imported] -
Arabidopsis thaliana, partial (5%)
Length = 467
Score = 29.6 bits (65), Expect = 5.0
Identities = 30/133 (22%), Positives = 58/133 (43%), Gaps = 8/133 (6%)
Frame = -3
Query: 112 ITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQ 171
+ SWE + RFVP R+L N + TQ + E +K++ + N+ +
Sbjct: 453 VDSWEXMKXLMRRRFVPSLYQRELHNKLQRLTQGS----RSVDEYYKEMEVSMIRANVME 286
Query: 172 --AEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVG--YELIEKMAIRAMNSSN----D 223
+A+F GL R ++ + E + L+ Q + +++ ++ +SSN D
Sbjct: 285 D*EATMARFLHGLNSDVRDVVEWQNHVELENLVHQASKLEQQLKRKSVMRRSSSNFHSPD 106
Query: 224 RQARRGVLEVEAY 236
+ R +E + Y
Sbjct: 105 WKVRNDYMESD*Y 67
>TC208962 UP|Q9ARG2 (Q9ARG2) Amino acid transporter, complete
Length = 1789
Score = 28.9 bits (63), Expect = 8.5
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 3/52 (5%)
Frame = +3
Query: 59 ALEDPHAHMERFIRNC--NTYRVQNVSTDTIRLSLFPF-SLRDTAEEWLNSQ 107
A++DPH+H F + C NT+R ++ + SLFP+ + E+W Q
Sbjct: 1272 AVDDPHSHAHAFFQRCGWNTWRFWVLALN----SLFPYRHVYFAKEDWTMDQ 1415
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,455,896
Number of Sequences: 63676
Number of extensions: 351731
Number of successful extensions: 1730
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1720
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1728
length of query: 728
length of database: 12,639,632
effective HSP length: 104
effective length of query: 624
effective length of database: 6,017,328
effective search space: 3754812672
effective search space used: 3754812672
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0388a.1


