

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0383.4
(221 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
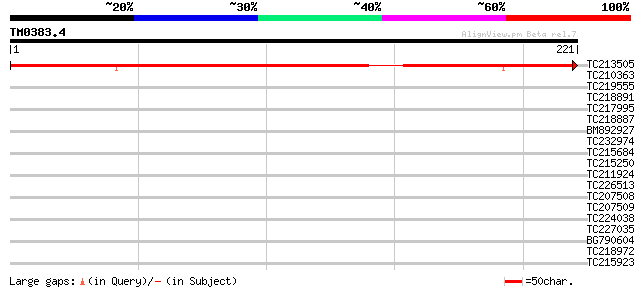
Score E
Sequences producing significant alignments: (bits) Value
TC213505 330 2e-91
TC210363 34 0.045
TC219555 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone n... 30 0.66
TC218891 homologue to UP|GAC1_YEAST (P28006) Protein phosphatase... 30 1.1
TC217995 similar to GB|AAB04607.1|1448917|ATHTHSY threonine synt... 28 2.5
TC218887 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone n... 28 2.5
BM892927 28 3.3
TC232974 weakly similar to GB|AAK54457.1|20384750|AY032998 DNA r... 28 4.2
TC215684 similar to UP|P93396 (P93396) Transformer-SR ribonucleo... 28 4.2
TC215250 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), parti... 27 5.5
TC211924 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor... 27 5.5
TC226513 similar to UP|O65673 (O65673) Nonclathrin coat protein ... 27 5.5
TC207508 weakly similar to UP|RL1_COXBU (Q83ET3) 50S ribosomal p... 27 7.2
TC207509 27 7.2
TC224038 weakly similar to UP|Q9LCV5 (Q9LCV5) Atp operon (Fragme... 27 7.2
TC227035 similar to UP|Q9M1H3 (Q9M1H3) ABC transporter-like prot... 27 7.2
BG790604 similar to GP|13562004|gb major ampullate spidroin 2-li... 27 7.2
TC218972 similar to UP|Q9ZPL6 (Q9ZPL6) DNA-binding protein 2, pa... 27 9.5
TC215923 similar to GB|AAS68109.1|45592910|BT011785 At2g38360 {A... 27 9.5
>TC213505
Length = 941
Score = 330 bits (847), Expect = 2e-91
Identities = 179/235 (76%), Positives = 188/235 (79%), Gaps = 14/235 (5%)
Frame = +1
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAK-------------MQEVAGE 47
MAGREVREYTNLSDPKDKKWGKGKDKIDDED+TFQRMVAK MQEVAGE
Sbjct: 97 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDITFQRMVAKGYWLPVLGG*CKQMQEVAGE 276
Query: 48 RGGYLHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPL 107
RGGYLHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASL DSSPASVPL
Sbjct: 277 RGGYLHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLADSSPASVPL 456
Query: 108 PLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQ 167
RVEPKPKSGIRQQDLLKKVVEIKPKRPRS++ NRKPDH KE Q
Sbjct: 457 AFRVEPKPKSGIRQQDLLKKVVEIKPKRPRSDA-------------NRKPDHDPSKEKEQ 597
Query: 168 SLSGVKKVEEQSLSGSTKVEDLSS-GSHDADAKPKINNSGGGLLGLAYASSDDDE 221
SL KKVEEQ LSGS K E+ + GS +A+AKPK+ +S GGLLGLAY SSDDDE
Sbjct: 598 SLPRTKKVEEQPLSGSKKAEEHAPLGSQEAEAKPKVESSSGGLLGLAYDSSDDDE 762
>TC210363
Length = 1015
Score = 34.3 bits (77), Expect = 0.045
Identities = 29/97 (29%), Positives = 44/97 (44%)
Frame = +3
Query: 18 KKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDSDDLLYLKEQMEAEEDAE 77
K+ K K+K + T + +K E + G+ L S L ++Q+ + AE
Sbjct: 48 KEMRKAKEKKAQVEATLPKNDSKTIEKQVTASTSVKGQSQLKSGVQLSKEDQIRMAQAAE 227
Query: 78 RLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPK 114
R EKRA AA KR A+L + ++ P EPK
Sbjct: 228 R-----EKRAAAAEKRIAALKIQANSATTAPTMSEPK 323
>TC219555 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone na-481-5),
partial (16%)
Length = 647
Score = 30.4 bits (67), Expect = 0.66
Identities = 32/124 (25%), Positives = 55/124 (43%), Gaps = 2/124 (1%)
Frame = +3
Query: 66 LKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLL 125
+++Q+ A++ + + +K A K+ +S DSS S +P PKS + + +
Sbjct: 240 IEKQLSAKKQKINEVAQKQKEAKVQ-KKESSSDDSSSESED----EKPAPKSAVLSKKIP 404
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGV--KKVEEQSLSGS 183
K PK+ + S S SSD K + E ++ + V KKVE S S
Sbjct: 405 AKNGAAPPKKGKLASSSSEDDSSDEDGVRAKKPQKMVAEKQKNGATVQKKKVESSSDDSS 584
Query: 184 TKVE 187
++ E
Sbjct: 585 SESE 596
>TC218891 homologue to UP|GAC1_YEAST (P28006) Protein phosphatase 1
regulatory subunit GAC1, partial (3%)
Length = 1463
Score = 29.6 bits (65), Expect = 1.1
Identities = 23/80 (28%), Positives = 34/80 (41%), Gaps = 2/80 (2%)
Frame = -3
Query: 98 VDSSPASVPLPLRVEPKPKSGIR--QQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNR 155
+ SPASVP P P P +R L + +E+ +ST A++ ++ NR
Sbjct: 999 ISESPASVPSPRFPNPSPSVFLRFLSWCLNSSTAASSSRNSPAENCRSTPATTRLAPPNR 820
Query: 156 KPDHGRLKETGQSLSGVKKV 175
P R T S S + V
Sbjct: 819 TPPEDRGAGTESSGSSTEPV 760
>TC217995 similar to GB|AAB04607.1|1448917|ATHTHSY threonine synthase
{Arabidopsis thaliana;} , partial (51%)
Length = 992
Score = 28.5 bits (62), Expect = 2.5
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -2
Query: 44 VAGERGGYLHGRGALDSDD 62
V GERGG + G GA D+DD
Sbjct: 724 VGGERGGSIAGGGAADADD 668
>TC218887 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone na-481-5),
partial (13%)
Length = 559
Score = 28.5 bits (62), Expect = 2.5
Identities = 27/98 (27%), Positives = 40/98 (40%), Gaps = 5/98 (5%)
Frame = +1
Query: 67 KEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLK 126
K+ ++ E + ++ +K A K+ +S DSS + P PKP +L
Sbjct: 259 KQVSAKKQKIEEVAQKQKKEAKVQKKKESSSDDSSESEEEKPA---PKP--------VLS 405
Query: 127 KVVEIK-----PKRPRSESKQSTSASSDVSVTNRKPDH 159
K V K PK+ + S S SSD N K H
Sbjct: 406 KKVPAKNGAALPKKDKPVSSSSEDDSSDEDEVNAKKPH 519
>BM892927
Length = 428
Score = 28.1 bits (61), Expect = 3.3
Identities = 15/52 (28%), Positives = 25/52 (47%)
Frame = -1
Query: 134 KRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTK 185
+R RS + AS + KPDHGR + + L ++ E +S S ++
Sbjct: 272 RRRRSPCACAGGASCRIECRGTKPDHGRRRRRNRFLRAWRRREVRSRSAESR 117
>TC232974 weakly similar to GB|AAK54457.1|20384750|AY032998 DNA repair
protein XRCC3 {Arabidopsis thaliana;} , partial (40%)
Length = 909
Score = 27.7 bits (60), Expect = 4.2
Identities = 20/68 (29%), Positives = 32/68 (46%)
Frame = +1
Query: 92 KRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVS 151
+R+ L+ P VPLPL P P + R L + ++P RPR +++ A
Sbjct: 187 RRSLRLLHLHPHRVPLPLPPPPPPLT--RLPRLPPRPPLLRPLRPRLPTRRPLRA----R 348
Query: 152 VTNRKPDH 159
+T P+H
Sbjct: 349 ITQPNPNH 372
>TC215684 similar to UP|P93396 (P93396) Transformer-SR ribonucleoprotein
(Fragment), partial (84%)
Length = 1263
Score = 27.7 bits (60), Expect = 4.2
Identities = 17/61 (27%), Positives = 30/61 (48%), Gaps = 1/61 (1%)
Frame = +2
Query: 106 PLPLRVEPKPKSGIRQQDLLKKVVEIKPK-RPRSESKQSTSASSDVSVTNRKPDHGRLKE 164
P P + + + +S R + + +P+ RPRS S+ + S + +R P HGR +
Sbjct: 83 PSPWKAQSRSRSRSRSRSRSRS----RPRQRPRSRSRSRGRSRSPIRGRSRSPIHGRSEP 250
Query: 165 T 165
T
Sbjct: 251 T 253
>TC215250 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (67%)
Length = 1148
Score = 27.3 bits (59), Expect = 5.5
Identities = 26/96 (27%), Positives = 42/96 (43%), Gaps = 4/96 (4%)
Frame = +1
Query: 102 PASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRK-PDHG 160
P +VP P V+P+P L KK V + P P + + T A V N K P+
Sbjct: 163 PVTVPAPAPVDPEPPLA-EAPPLEKKAVAVPP--PVPAAAEETKALVVVEKENEKIPEPV 333
Query: 161 RLKETGQSLS---GVKKVEEQSLSGSTKVEDLSSGS 193
+ +G SL + ++E++ + K + S S
Sbjct: 334 KKNASGGSLDRDIALAEIEKEKRLSNVKAWEESEKS 441
>TC211924 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor SC35
(CG5442-PB) (LD32469p), partial (9%)
Length = 510
Score = 27.3 bits (59), Expect = 5.5
Identities = 21/87 (24%), Positives = 36/87 (41%), Gaps = 8/87 (9%)
Frame = +3
Query: 105 VPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTN--------RK 156
V LPL + P + + + + E P+ S T ++S ++++ +K
Sbjct: 249 VLLPLHLLPNKPDPPQASNFITPIQEPDPEPEPEHSPSITESTSTTALSSSKRWKDIFKK 428
Query: 157 PDHGRLKETGQSLSGVKKVEEQSLSGS 183
D + + VKK E QS SGS
Sbjct: 429 SDKKNAENNNEEKEKVKKKERQSGSGS 509
>TC226513 similar to UP|O65673 (O65673) Nonclathrin coat protein gamma-like
protein, partial (41%)
Length = 1493
Score = 27.3 bits (59), Expect = 5.5
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = -3
Query: 190 SSGSHDADAKPKINNSGGGLLGLAYASSDD 219
S+GS D + PK NSG L+YAS+ D
Sbjct: 435 STGSEDLKSFPKFANSGMDRSSLSYASAVD 346
>TC207508 weakly similar to UP|RL1_COXBU (Q83ET3) 50S ribosomal protein L1,
partial (35%)
Length = 983
Score = 26.9 bits (58), Expect = 7.2
Identities = 28/140 (20%), Positives = 52/140 (37%), Gaps = 17/140 (12%)
Frame = +1
Query: 72 AEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEI 131
A EDA RR + R A + VP PL ++PK DL + + ++
Sbjct: 409 AAEDAVESERRMIEAENEMRMRMAKAAEDERMKVPFPLLIKPKKNEKPPPLDLNEAIRQV 588
Query: 132 KPKRPRSESKQSTSASSDVSVTNRKPD----------HGRLKET-------GQSLSGVKK 174
K +++ ++ A + V +++ + HG K G K
Sbjct: 589 K-TNAKAKFDETVEAHIRLGVDSKRTEMAVRGTVILPHGAPKAVSVAVFAEGAEAEEAKA 765
Query: 175 VEEQSLSGSTKVEDLSSGSH 194
+ G +E+++SG +
Sbjct: 766 AGADIVGGKELIEEIASGKN 825
>TC207509
Length = 641
Score = 26.9 bits (58), Expect = 7.2
Identities = 26/104 (25%), Positives = 40/104 (38%), Gaps = 1/104 (0%)
Frame = +3
Query: 30 EDVTFQRMVAKMQEVA-GERGGYLHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAF 88
ED+ + + V + VA RGG + L + EDA RR +
Sbjct: 327 EDIRYVKDVPSIAPVAYASRGG------------VAPLPDDRAPVEDAAESERRMIEAEN 470
Query: 89 AAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIK 132
R A V+ VP PL ++PK DL + + ++K
Sbjct: 471 EMRLRMAKAVEEERMKVPFPLLIKPKKNEKPPPLDLNEAIRQVK 602
>TC224038 weakly similar to UP|Q9LCV5 (Q9LCV5) Atp operon (Fragment), partial
(4%)
Length = 559
Score = 26.9 bits (58), Expect = 7.2
Identities = 23/69 (33%), Positives = 29/69 (41%), Gaps = 8/69 (11%)
Frame = -2
Query: 77 ERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQ-QDLLKKVVE----- 130
+RLLRR +R P PLP R EP+P+ + LL VVE
Sbjct: 402 QRLLRRRTQRPL-------------PPCAPLPQRREPRPRRRVPPINSLLLAVVEPYLLR 262
Query: 131 --IKPKRPR 137
+P RPR
Sbjct: 261 RCAEPHRPR 235
>TC227035 similar to UP|Q9M1H3 (Q9M1H3) ABC transporter-like protein
(At3g54540), partial (56%)
Length = 1561
Score = 26.9 bits (58), Expect = 7.2
Identities = 9/28 (32%), Positives = 19/28 (67%)
Frame = +2
Query: 4 REVREYTNLSDPKDKKWGKGKDKIDDED 31
++V++ + K+K GKGK K+D+++
Sbjct: 377 KKVKDQAKFAAAKEKSKGKGKGKVDEDE 460
>BG790604 similar to GP|13562004|gb major ampullate spidroin 2-like protein
{Nephila madagascariensis}, partial (1%)
Length = 411
Score = 26.9 bits (58), Expect = 7.2
Identities = 20/56 (35%), Positives = 24/56 (42%), Gaps = 2/56 (3%)
Frame = -1
Query: 104 SVPLPLRVEPKPKSGIRQQDLLKKVVEIK--PKRPRSESKQSTSASSDVSVTNRKP 157
S+PLPL VEP+ +G VV P P SESK S + S P
Sbjct: 288 SIPLPLAVEPETPAGGGGSPSSSVVVPPTSWPPPPNSESKPQLSCPPNPSEAAAAP 121
>TC218972 similar to UP|Q9ZPL6 (Q9ZPL6) DNA-binding protein 2, partial (33%)
Length = 1177
Score = 26.6 bits (57), Expect = 9.5
Identities = 15/36 (41%), Positives = 22/36 (60%)
Frame = +2
Query: 118 GIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVT 153
G + ++ +K VE PKR ++E QS ASS +VT
Sbjct: 392 GDHETEVDEKNVEPDPKRRKAEVSQSDPASSHRTVT 499
>TC215923 similar to GB|AAS68109.1|45592910|BT011785 At2g38360 {Arabidopsis
thaliana;} , partial (76%)
Length = 1055
Score = 26.6 bits (57), Expect = 9.5
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 6/57 (10%)
Frame = +2
Query: 89 AAFKRAASLVDSSPASVPLPLRVEPKP------KSGIRQQDLLKKVVEIKPKRPRSE 139
+ +R SL DSSP+S LP + P P R + L ++V PRS+
Sbjct: 707 STIRRILSLPDSSPSSAALPPPLPPPPPLFPPLPHAFR*KKNLSEIVRSGDVSPRSD 877
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.307 0.127 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,890,377
Number of Sequences: 63676
Number of extensions: 74847
Number of successful extensions: 337
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 333
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 334
length of query: 221
length of database: 12,639,632
effective HSP length: 94
effective length of query: 127
effective length of database: 6,654,088
effective search space: 845069176
effective search space used: 845069176
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0383.4


