

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
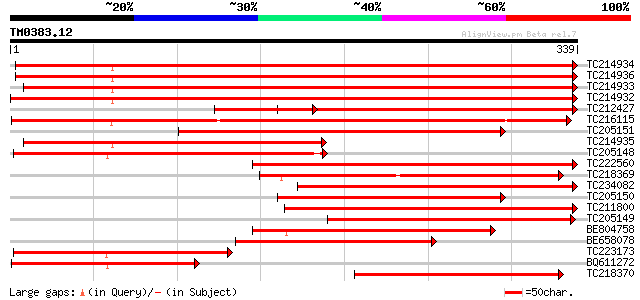
Query= TM0383.12
(339 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC214934 homologue to UP|Q43016 (Q43016) Alcohol dehydrogenase-1... 381 e-106
TC214936 UP|O82478 (O82478) Alcohol dehydrogenase Adh-1 (Fragme... 363 e-101
TC214933 UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment), ... 358 3e-99
TC214932 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1... 353 8e-98
TC212427 similar to UP|Q8RV10 (Q8RV10) AT5g24760/T4C12_30, parti... 297 4e-81
TC216115 weakly similar to UP|ADHX_UROHA (P80467) Alcohol dehydr... 290 5e-79
TC205151 similar to UP|ADHX_ORYSA (P93436) Alcohol dehydrogenase... 215 2e-56
TC214935 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1... 206 9e-54
TC205148 similar to GB|AAB06322.1|1498024|ATU63931 glutathione-d... 194 4e-50
TC222560 weakly similar to UP|Q41241 (Q41241) Alcohol dehydrogen... 183 8e-47
TC218369 weakly similar to UP|ADHX_HUMAN (P11766) Alcohol dehydr... 178 3e-45
TC234082 similar to UP|Q41241 (Q41241) Alcohol dehydrogenase ADH... 171 4e-43
TC205150 homologue to UP|ADHX_ORYSA (P93436) Alcohol dehydrogena... 171 4e-43
TC211800 weakly similar to UP|ADHX_HUMAN (P11766) Alcohol dehydr... 166 1e-41
TC205149 similar to UP|ADHX_ORYSA (P93436) Alcohol dehydrogenase... 140 1e-33
BE804758 137 7e-33
BE658078 similar to GP|7705215|gb| alcohol dehydrogenase ADH {L... 132 2e-31
TC223173 similar to UP|Q9FH04 (Q9FH04) Alcohol dehydrogenase, pa... 127 7e-30
BQ611272 homologue to GP|1498024|gb| glutathione-dependent forma... 127 9e-30
TC218370 weakly similar to UP|ADHX_DROME (P46415) Alcohol dehydr... 118 4e-27
>TC214934 homologue to UP|Q43016 (Q43016) Alcohol dehydrogenase-1F ,
complete
Length = 1450
Score = 381 bits (978), Expect = e-106
Identities = 184/340 (54%), Positives = 243/340 (71%), Gaps = 4/340 (1%)
Frame = +3
Query: 4 STSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH--- 60
S++ QVI CKAAV+W AG+PLVIEEV+V+PPQ E+R+K++ TSLC +D+ WE+
Sbjct: 96 SSTAGQVIKCKAAVSWEAGKPLVIEEVEVAPPQAGEVRLKILYTSLCHTDVYFWEAKGQT 275
Query: 61 PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLER 120
P+FPRIFGHEA GIVESVG GVT K GDH L VF GEC EC C S +SN+C +L +
Sbjct: 276 PLFPRIFGHEAGGIVESVGEGVTHLKPGDHALPVFTGECGECPHCKSEESNMCDLLRINT 455
Query: 121 N-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVA 179
+ G+M D +TRFSIKG+P+YH+ G S+FSEYTVVH+GC KV+P APL+KIC+LSCG+
Sbjct: 456 DRGVMIHDSQTRFSIKGQPIYHFVGTSTFSEYTVVHAGCVAKVNPAAPLDKICVLSCGIC 635
Query: 180 AGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGI 239
GLGA NVA GS+V IFGLG VGL+ A+GA++ GASRIIGVD + E AK FG+
Sbjct: 636 TGLGATINVAKPKPGSSVAIFGLGAVGLAAAEGARISGASRIIGVDLVSSRFEEAKKFGV 815
Query: 240 TEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKV 299
E V+P +++P+ +VI +T+GG D + EC G + +A + DGWG+ V +GVP
Sbjct: 816 NEFVNPKDHDKPVQEVIAAMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNK 995
Query: 300 KPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
H L RTLKG+ +G +KP++DLPS+V+KY+N
Sbjct: 996 DDAFKTHPVNFLNERTLKGTFYGNYKPRTDLPSVVEKYMN 1115
>TC214936 UP|O82478 (O82478) Alcohol dehydrogenase Adh-1 (Fragment) ,
complete
Length = 1353
Score = 363 bits (931), Expect = e-101
Identities = 174/340 (51%), Positives = 239/340 (70%), Gaps = 4/340 (1%)
Frame = +3
Query: 4 STSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH--- 60
S++ Q I CKAA+AW AG+PLVIEEV+V+PPQ E+R+K++ TSLC +D+ W++
Sbjct: 33 SSTVGQTIKCKAAIAWEAGKPLVIEEVEVAPPQAGEVRLKILYTSLCHTDVYFWDAKGQT 212
Query: 61 PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLER 120
P+FPRIFGHEASGIVESVG GVT K GDH L VF GEC +C C S +SN+C++L +
Sbjct: 213 PLFPRIFGHEASGIVESVGEGVTHLKPGDHALPVFTGECGDCAHCKSEESNMCELLRINT 392
Query: 121 N-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVA 179
+ G+M D ++RFS G+P++H+ G S+FSEYTVVH+GC K++P APL+K+C+LSCG+
Sbjct: 393 DRGVMIHDGQSRFSKNGQPIHHFLGTSTFSEYTVVHAGCVAKINPAAPLDKVCVLSCGIC 572
Query: 180 AGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGI 239
G GA NVA GS+V IFGLG V L+ A+GA++ GASRIIGVD + E AK FG+
Sbjct: 573 TGFGATVNVAKPKPGSSVAIFGLGAVALAAAEGARVSGASRIIGVDLVSARFEEAKKFGV 752
Query: 240 TEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKV 299
E V+P +++P+ QVI +T+GG D + EC G + +A + DGWGL V +GVP
Sbjct: 753 NEFVNPKDHDKPVQQVIAEMTNGGVDRAVECTGSIQAMVSAFECVHDGWGLAVLVGVPSK 932
Query: 300 KPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
L RTLKG+ +G +KP++DLPS+V+KY++
Sbjct: 933 DDAFKTAPINFLNERTLKGTFYGNYKPRTDLPSVVEKYMS 1052
>TC214933 UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment), complete
Length = 1477
Score = 358 bits (918), Expect = 3e-99
Identities = 168/335 (50%), Positives = 233/335 (69%), Gaps = 4/335 (1%)
Frame = +1
Query: 9 QVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH---PIFPR 65
QVI C+AAVAW AG+PL IE ++V+PPQ E+R++++ SLCRSD+ W++ P+FPR
Sbjct: 67 QVIKCRAAVAWEAGKPLSIETIEVAPPQKGEVRLRILFNSLCRSDVYWWDAKDQTPLFPR 246
Query: 66 IFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERN-GLM 124
I GHEASGIVESVG GVT K GDH L +F GEC EC C S +SN+C++L + + G+M
Sbjct: 247 ILGHEASGIVESVGEGVTHLKPGDHALPIFTGECGECTYCKSEESNLCELLRINTDRGVM 426
Query: 125 HSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGA 184
SD KTRFS G+P+YH+ G S+FSEYTV+H GC K++PNAPL+K+ ++SCG G GA
Sbjct: 427 LSDGKTRFSKNGQPIYHFVGTSTFSEYTVLHEGCVAKINPNAPLDKVAIVSCGFCTGFGA 606
Query: 185 AWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVD 244
NVA +TV +FGLG VGL+ +GA++ GASRIIGVD P + E AK FG+T+ V+
Sbjct: 607 TVNVAKPKPNNTVAVFGLGAVGLAACEGARVSGASRIIGVDLLPNRFEQAKKFGVTDFVN 786
Query: 245 PNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMS 304
P + +P+ +VI +T+GG D + EC G +A + DGWG V +GVPK
Sbjct: 787 PKDHNKPVQEVIAEMTNGGVDRAIECTGSIQASISAFECTHDGWGTAVLVGVPKKDVEFK 966
Query: 305 AHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
+ + GRTLKG+ +G ++P++D+P +V+KY+N
Sbjct: 967 TNPMKFMEGRTLKGTFYGHYRPRTDIPGVVEKYLN 1071
>TC214932 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment),
complete
Length = 1456
Score = 353 bits (905), Expect = 8e-98
Identities = 166/343 (48%), Positives = 234/343 (67%), Gaps = 4/343 (1%)
Frame = +3
Query: 1 MSSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH 60
+ + ++ QVI C+AAVAW AG+PL IE ++V+PPQ E+R+K++ SLCR+D+ W++
Sbjct: 84 VEAMSTAGQVIKCRAAVAWEAGKPLSIETIEVAPPQKGEVRLKILFNSLCRTDVYWWDAK 263
Query: 61 ---PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLG 117
P+FPRI GHEASGIVESVG GVT K GDH L +F GEC EC C S +SN+C++L
Sbjct: 264 GQTPLFPRILGHEASGIVESVGKGVTHLKPGDHALPIFTGECGECTYCKSEESNLCELLR 443
Query: 118 LERN-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC 176
+ + G+M SD KTRFS G+P+YH+ G S+FSEYTV+H GC K++P APL+K+ ++SC
Sbjct: 444 INTDRGVMLSDGKTRFSKNGQPIYHFVGTSTFSEYTVLHEGCVAKINPAAPLDKVAVVSC 623
Query: 177 GVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKA 236
G G GA NVA +TV +FGLG VGL+ +GA++ GASRIIGVD + E AK
Sbjct: 624 GFCTGFGATVNVAKPKPNNTVAVFGLGAVGLAACEGARVSGASRIIGVDLLTNRFEQAKQ 803
Query: 237 FGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGV 296
FG+T+ V+P + +P+ +VI +T+GG D + EC G +A + DGWG V + V
Sbjct: 804 FGVTDFVNPKDHNKPVQEVIAEMTNGGVDRAIECTGSIQASISAFECTHDGWGTAVLVSV 983
Query: 297 PKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
PK H + GRTLKG+ +G ++P++D+P +V+KY+N
Sbjct: 984 PKKDAEFKTHPMKFMEGRTLKGTFYGHYRPRTDIPGVVEKYLN 1112
>TC212427 similar to UP|Q8RV10 (Q8RV10) AT5g24760/T4C12_30, partial (55%)
Length = 670
Score = 297 bits (761), Expect = 4e-81
Identities = 148/179 (82%), Positives = 159/179 (88%)
Frame = +1
Query: 161 KVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASR 220
K +P L K + G+ L +NVADVSKGSTVVIFGLGTVGLSVAQ +KL+GASR
Sbjct: 118 KSAPLHLLRKYVF*AVGLLQVLVHLFNVADVSKGSTVVIFGLGTVGLSVAQASKLRGASR 297
Query: 221 IIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTA 280
IIGVDNNPQKCENAKAFG+TEVVDPNSY+EPIAQVI RITDGGADFSFECVGDTD ITTA
Sbjct: 298 IIGVDNNPQKCENAKAFGVTEVVDPNSYKEPIAQVIKRITDGGADFSFECVGDTDTITTA 477
Query: 281 LQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
LQSCCDGWGLTVTLGVPKVKP MSAHYGLLLMGRTLKGS+FGGWKPKSDLPSLV+KY+N
Sbjct: 478 LQSCCDGWGLTVTLGVPKVKPEMSAHYGLLLMGRTLKGSLFGGWKPKSDLPSLVEKYLN 654
Score = 125 bits (313), Expect = 4e-29
Identities = 59/62 (95%), Positives = 60/62 (96%)
Frame = +3
Query: 123 LMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGL 182
LMHSDQKTRFS+KGKPVYHYC VSSFSEYTVVHSGCAVKVSP APLEKICLLSCGVAAGL
Sbjct: 3 LMHSDQKTRFSLKGKPVYHYCAVSSFSEYTVVHSGCAVKVSPLAPLEKICLLSCGVAAGL 182
Query: 183 GA 184
GA
Sbjct: 183 GA 188
>TC216115 weakly similar to UP|ADHX_UROHA (P80467) Alcohol dehydrogenase
class III (Glutathione-dependent formaldehyde
dehydrogenase) (FDH) , partial (42%)
Length = 1433
Score = 290 bits (743), Expect = 5e-79
Identities = 145/338 (42%), Positives = 213/338 (62%), Gaps = 3/338 (0%)
Frame = +1
Query: 2 SSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWES-- 59
S+ + T ++ITCKAA+ WG G+P+ +EE+QV PP+ E+RVK++C S+C +DIS+ +
Sbjct: 55 STMSKTSEIITCKAAICWGIGKPVTVEEIQVDPPKATEVRVKMLCASICSTDISSTKGFP 234
Query: 60 HPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLE 119
H FP GHE GI+ESVG VT KEGD V+ +IGEC EC+ C S K+N+C +
Sbjct: 235 HTNFPIALGHEGVGIIESVGDQVTNLKEGDVVIPTYIGECQECENCVSEKTNLCMTYPVR 414
Query: 120 RNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVA 179
GLM D +R SI+G+ +YH +++SEY V + +KV P +SCG +
Sbjct: 415 WTGLM-PDNTSRMSIRGERIYHIFSCATWSEYMVSDANYVLKVDPTIDRAHASFISCGFS 591
Query: 180 AGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGI 239
G GAAW A V GSTV +FGLG VGL G+K++GASRIIG+D N K +AFGI
Sbjct: 592 TGFGAAWKEAKVESGSTVAVFGLGAVGLGAVIGSKMQGASRIIGIDTNENKRAKGEAFGI 771
Query: 240 TEVVDPNSYEEPIAQVINRITDG-GADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPK 298
T+ ++P + ++++ ++ G GAD+SFEC G + +++ +L++ G G + +GV
Sbjct: 772 TDFINPGDSNKSASELVKELSGGMGADYSFECTGVSTLLSESLEATKIGTGKAIVIGV-G 948
Query: 299 VKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDK 336
++ + +L+GRTLKGS+FGG + SDL L DK
Sbjct: 949 IEITLPLGLFAILLGRTLKGSVFGGLRAISDLSILADK 1062
>TC205151 similar to UP|ADHX_ORYSA (P93436) Alcohol dehydrogenase class III
(Glutathione-dependent formaldehyde dehydrogenase)
(FDH) (FALDH) (GSH-FDH) , partial (54%)
Length = 667
Score = 215 bits (548), Expect = 2e-56
Identities = 105/196 (53%), Positives = 138/196 (69%), Gaps = 1/196 (0%)
Frame = +2
Query: 102 CKPCTSGKSNVC-QVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAV 160
CK SGK+N+C +V G+M +D K+RFSI GKP+YH+ G S+FS+YT H
Sbjct: 2 CKTRKSGKTNLCGKVRSATGVGVMLNDGKSRFSINGKPIYHFMGTSTFSQYTGAHDLSVA 181
Query: 161 KVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASR 220
K+ P APLEK+CLL CGV+ GLGA WN A V GS V IFGLGTVGL+VAQGAK GASR
Sbjct: 182 KIDPVAPLEKVCLLGCGVSTGLGAVWNTAKVESGSIVAIFGLGTVGLAVAQGAKTAGASR 361
Query: 221 IIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTA 280
+IG+D + +K + AK FG+TE ++P +++PI QVI TDGG D+SFEC+G+ ++ A
Sbjct: 362 VIGIDIDSKKFDIAKNFGVTEFINPYEHDKPIQQVIIDRTDGGVDYSFECIGNVSVMRAA 541
Query: 281 LQSCCDGWGLTVTLGV 296
L+ C G G +V +GV
Sbjct: 542 LECCHHGSGTSVIVGV 589
>TC214935 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment),
partial (47%)
Length = 790
Score = 206 bits (525), Expect = 9e-54
Identities = 101/186 (54%), Positives = 131/186 (70%), Gaps = 5/186 (2%)
Frame = +2
Query: 9 QVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH---PIFPR 65
QVI C+AAVAW AG+PL IE ++V+PPQ E+R++++ SLCRSD+ W++ P+FPR
Sbjct: 47 QVIKCRAAVAWEAGKPLSIETIEVAPPQKGEVRLRILFNSLCRSDVYWWDAKDQTPLFPR 226
Query: 66 IFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERN-GLM 124
I GHEASGIVESVG GVT K GDH L +F GEC EC C S +SN+C++L + + G+M
Sbjct: 227 ILGHEASGIVESVGEGVTHLKPGDHALPIFTGECGECTYCKSEESNLCELLRINTDRGVM 406
Query: 125 HSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLS-CGVAAGLG 183
SD KTRFS G+P+YH+ G +FSEYT +H GC K++PNAPL+K C G G
Sbjct: 407 LSDGKTRFSKNGQPIYHFVGTFTFSEYTGLHEGCVAKINPNAPLDKSCYWQLVDSCTGFG 586
Query: 184 AAWNVA 189
A NVA
Sbjct: 587 ATGNVA 604
>TC205148 similar to GB|AAB06322.1|1498024|ATU63931 glutathione-dependent
formaldehyde dehydrogenase {Arabidopsis thaliana;} ,
partial (48%)
Length = 678
Score = 194 bits (494), Expect = 4e-50
Identities = 97/192 (50%), Positives = 129/192 (66%), Gaps = 4/192 (2%)
Frame = +1
Query: 3 SSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAW---ES 59
S + QVITCKAAVAW +PL +++VQV+PPQ E+RV+++ T+LC +D W +
Sbjct: 64 SMATQGQVITCKAAVAWEPNKPLTVQDVQVAPPQAGEVRVQILFTALCHTDAYTWGGKDP 243
Query: 60 HPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVC-QVLGL 118
+FP I GHEA+GIVESVG GVT + GDHV+ + EC ECK C SGK+N+C +V
Sbjct: 244 EGLFPCILGHEAAGIVESVGEGVTNVQPGDHVIPCYQAECGECKTCKSGKTNLCGKVRSA 423
Query: 119 ERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGV 178
G+M +D K+RFSI GKP+YH+ G S+FS+YTVVH K+ P PLEK LL CG
Sbjct: 424 TGVGVMLNDGKSRFSINGKPIYHFMGTSTFSQYTVVHDVSVAKIDPKPPLEKGWLLGCGG 603
Query: 179 AAGLGAAWNVAD 190
+ GL WN+ +
Sbjct: 604 STGL---WNLLE 630
>TC222560 weakly similar to UP|Q41241 (Q41241) Alcohol dehydrogenase ADH,
partial (60%)
Length = 704
Score = 183 bits (465), Expect = 8e-47
Identities = 93/196 (47%), Positives = 132/196 (66%), Gaps = 2/196 (1%)
Frame = +3
Query: 146 SSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTV 205
SSFSEYTVV +K+ P P + CL+SCG++AG+GAAW A V GSTV IFGLG++
Sbjct: 3 SSFSEYTVVDIAHLIKIDPAIPPNRACLISCGISAGIGAAWRAAGVEPGSTVAIFGLGSI 182
Query: 206 GLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYE-EPIAQVINRITDGGA 264
GL+VA+GA+L GA++IIGVD NP++ E K FG+T+ V E + ++QVI +T GGA
Sbjct: 183 GLAVAEGARLCGATKIIGVDVNPERYEIGKRFGLTDFVHSGECENKSVSQVIIEMTGGGA 362
Query: 265 DFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYG-LLLMGRTLKGSMFGG 323
D+ FECVG ++ A SC GWG T+ LGV K ++ +L+ G++L G +FGG
Sbjct: 363 DYCFECVGMASLMHEAYASCRKGWGKTIVLGVDKPGSKLNLSCSEVLVSGKSLMGCLFGG 542
Query: 324 WKPKSDLPSLVDKYVN 339
P+S +P L+ +Y++
Sbjct: 543 LIPESHVPILLKRYMD 590
>TC218369 weakly similar to UP|ADHX_HUMAN (P11766) Alcohol dehydrogenase
class III chi chain (Glutathione-dependent formaldehyde
dehydrogenase) (FDH) , partial (34%)
Length = 850
Score = 178 bits (452), Expect = 3e-45
Identities = 90/185 (48%), Positives = 122/185 (65%), Gaps = 3/185 (1%)
Frame = +1
Query: 150 EYTVVHSGCAVK---VSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVG 206
EYTVV S C VK + ++ + LLSCGV+ G+GAAWN A+V GSTV +FGLG VG
Sbjct: 1 EYTVVDSACVVKFVSTDHSLSIKNLTLLSCGVSTGVGAAWNTANVHSGSTVAVFGLGAVG 180
Query: 207 LSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADF 266
L+VA+GA+ +GAS+IIGVD NP K KA G+T ++P E+P+ + I +TDGG +
Sbjct: 181 LAVAEGARARGASKIIGVDINPDKF--IKAMGVTNFINPKDEEKPVYERIREMTDGGVHY 354
Query: 267 SFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKP 326
SFEC G+ D++ A S +GWGLTV LG+ ++ H L GR + GS+FGG+K
Sbjct: 355 SFECTGNVDVLRDAFLSAHEGWGLTVVLGIHASPTLLPIHPMELFDGRNIVGSVFGGFKG 534
Query: 327 KSDLP 331
+S LP
Sbjct: 535 RSHLP 549
>TC234082 similar to UP|Q41241 (Q41241) Alcohol dehydrogenase ADH, partial
(53%)
Length = 705
Score = 171 bits (433), Expect = 4e-43
Identities = 88/169 (52%), Positives = 117/169 (69%), Gaps = 2/169 (1%)
Frame = +2
Query: 173 LLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCE 232
LL CGV+ G+GAAW A V GSTV IFGLG++GL+VA+GA+L GA+RIIGVD NP+K E
Sbjct: 2 LLGCGVSTGVGAAWRTAGVEPGSTVAIFGLGSIGLAVAEGARLCGATRIIGVDINPEKFE 181
Query: 233 NAKAFGITEVVDPNSY-EEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLT 291
K FG+T+ V+ +P+ QVI ITDGGAD+ FECVG ++ A SC GWG T
Sbjct: 182 IGKKFGVTDFVNAGECGGKPVGQVIIEITDGGADYCFECVGMASLVHEAYASCRKGWGKT 361
Query: 292 VTLGVPKV-KPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
+ LGV K + + Y +L G++L GS+FGG KPKS +P L+ +Y++
Sbjct: 362 IVLGVDKPGARINLSSYEVLHDGKSLMGSLFGGLKPKSHVPILLKRYMD 508
>TC205150 homologue to UP|ADHX_ORYSA (P93436) Alcohol dehydrogenase class III
(Glutathione-dependent formaldehyde dehydrogenase)
(FDH) (FALDH) (GSH-FDH) , partial (38%)
Length = 441
Score = 171 bits (433), Expect = 4e-43
Identities = 78/136 (57%), Positives = 102/136 (74%)
Frame = +3
Query: 161 KVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASR 220
K+ P APLEK+CLL CGV+ GLGA WN A V GS V IFGLGTVGL+VA+GAK GASR
Sbjct: 12 KIDPKAPLEKVCLLGCGVSTGLGAVWNTAKVESGSIVAIFGLGTVGLAVAEGAKTAGASR 191
Query: 221 IIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTA 280
+IG+D + +K + AK FG+TE ++PN +++PI QVI TDGG D+SFEC+G+ ++ A
Sbjct: 192 VIGIDIDSKKFDVAKNFGVTEFINPNEHDKPIQQVIIDRTDGGVDYSFECIGNVSVMRAA 371
Query: 281 LQSCCDGWGLTVTLGV 296
L+ C GWG +V +GV
Sbjct: 372 LECCHKGWGTSVIVGV 419
>TC211800 weakly similar to UP|ADHX_HUMAN (P11766) Alcohol dehydrogenase
class III chi chain (Glutathione-dependent formaldehyde
dehydrogenase) (FDH) , partial (20%)
Length = 700
Score = 166 bits (420), Expect = 1e-41
Identities = 81/175 (46%), Positives = 115/175 (65%)
Frame = +3
Query: 165 NAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGV 224
N ++++ LLSCGV++G+GAAWN ADV GSTV +FGLG VGL+VA+GA+ +GASRIIGV
Sbjct: 18 NRNIKRLTLLSCGVSSGVGAAWNTADVHFGSTVAVFGLGVVGLAVAEGARARGASRIIGV 197
Query: 225 DNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSC 284
D N K A+ GIT+ ++P E P+ ++I +T GG +SFEC G+ +++ A S
Sbjct: 198 DINSDKFIKAREMGITDFINPKDDERPVYEIIGEMTGGGVHYSFECAGNLNVLRDAFLSA 377
Query: 285 CDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
+GWGLTV +G+ ++ H L GR + GS FGG K K+ LP + +N
Sbjct: 378 HEGWGLTVLVGIHLSPKMLPIHPMELFHGRRIVGSNFGGIKGKTQLPHFAKECMN 542
>TC205149 similar to UP|ADHX_ORYSA (P93436) Alcohol dehydrogenase class III
(Glutathione-dependent formaldehyde dehydrogenase)
(FDH) (FALDH) (GSH-FDH) , partial (49%)
Length = 790
Score = 140 bits (352), Expect = 1e-33
Identities = 69/148 (46%), Positives = 97/148 (64%)
Frame = +3
Query: 191 VSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEE 250
V G V I VGL+VA+GAK GASR+IG+D + +K + AK FG+TE ++PN +++
Sbjct: 6 VESGXXVAIXXXXXVGLAVAEGAKTAGASRVIGIDIDSKKFDIAKNFGVTEFINPNEHDK 185
Query: 251 PIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLL 310
PI QVI TDGG D+SFEC+G+ ++ AL+ C GWG +V +GV +S L
Sbjct: 186 PIQQVIIDRTDGGVDYSFECIGNVSVMRAALECCHKGWGTSVIVGVAASGQEISTRPFQL 365
Query: 311 LMGRTLKGSMFGGWKPKSDLPSLVDKYV 338
+ GR KG+ FGG+K +S +P LVDKY+
Sbjct: 366 VSGRVWKGTAFGGFKSRSQVPWLVDKYL 449
>BE804758
Length = 457
Score = 137 bits (345), Expect = 7e-33
Identities = 73/151 (48%), Positives = 97/151 (63%), Gaps = 6/151 (3%)
Frame = +2
Query: 146 SSFSEYTVVHSGCAVKVSP------NAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVI 199
S+F+EYTVV S C VK+ N ++++ LLSCGV+ G+GAAWN ADV GS V +
Sbjct: 5 STFTEYTVVDSACVVKIHVDGDGDLNPYIKRLTLLSCGVSTGVGAAWNTADVHFGSPVAV 184
Query: 200 FGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRI 259
FGL VGLSVA+GA+ +GAS+IIGVD N K A+A GIT+ +DP E+P+ + I +
Sbjct: 185 FGLRAVGLSVAEGARSRGASKIIGVDINSDKYIKARAMGITDFIDPRDDEKPVYERIRDM 364
Query: 260 TDGGADFSFECVGDTDMITTALQSCCDGWGL 290
T GG S EC D +++ A S G GL
Sbjct: 365 TRGGVHSSSECADDVHVLSEAFLSVHAGCGL 457
>BE658078 similar to GP|7705215|gb| alcohol dehydrogenase ADH {Lycopersicon
esculentum}, partial (32%)
Length = 583
Score = 132 bits (332), Expect = 2e-31
Identities = 64/121 (52%), Positives = 86/121 (70%), Gaps = 1/121 (0%)
Frame = -2
Query: 136 GKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGS 195
G+ +YH+ +SSFSEYTVV +K+ P P ++ CLL CGV+ G+GAAW A V GS
Sbjct: 564 GEIIYHFLFISSFSEYTVVDIANLIKIDPEIPPDRACLLGCGVSTGVGAAWRTAGVEPGS 385
Query: 196 TVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSY-EEPIAQ 254
TV IFGLG++GL+VA+GA+L GA+RIIGVD NP+K E K FG+T+ V+ +P+ Q
Sbjct: 384 TVAIFGLGSIGLAVAEGARLCGATRIIGVDINPEKFEIGKKFGVTDFVNAGECGGKPVGQ 205
Query: 255 V 255
V
Sbjct: 204 V 202
>TC223173 similar to UP|Q9FH04 (Q9FH04) Alcohol dehydrogenase, partial (31%)
Length = 470
Score = 127 bits (319), Expect = 7e-30
Identities = 67/134 (50%), Positives = 89/134 (66%), Gaps = 3/134 (2%)
Frame = +1
Query: 3 SSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISA---WES 59
+ST+ Q I CKAAV+ AGEPLVIE++ V+PP+P E R++++C+SLC SDI+ +
Sbjct: 43 ASTTEGQPIRCKAAVSRRAGEPLVIEDIIVAPPKPREARIRIICSSLCHSDITLRNLQDP 222
Query: 60 HPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLE 119
IFPRI GHEA+G+VESVG VTE +GD V+ V + EC EC C S KSN C +
Sbjct: 223 PAIFPRILGHEATGVVESVGKDVTEVTKGDVVIPVILPECGECIDCKSTKSNRCTNFPFK 402
Query: 120 RNGLMHSDQKTRFS 133
+ M D TRF+
Sbjct: 403 VSPWMPRDGTTRFT 444
>BQ611272 homologue to GP|1498024|gb| glutathione-dependent formaldehyde
dehydrogenase {Arabidopsis thaliana}, partial (29%)
Length = 456
Score = 127 bits (318), Expect = 9e-30
Identities = 61/115 (53%), Positives = 81/115 (70%), Gaps = 3/115 (2%)
Frame = +2
Query: 2 SSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAW---E 58
S +T+ QV TCKAAVAW +PL IE+VQV+PPQ E+R++++ T+LC +D W +
Sbjct: 110 SMATTQGQVTTCKAAVAWEPNKPLSIEDVQVAPPQNGEVRIQILYTALCHTDAYTWSGKD 289
Query: 59 SHPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVC 113
+FP I GHEA+GIVESVG GVT + GDHV+ + EC ECK C SGK+N+C
Sbjct: 290 PEGLFPCILGHEAAGIVESVGEGVTAVQPGDHVIPCYQAECGECKFCKSGKTNLC 454
>TC218370 weakly similar to UP|ADHX_DROME (P46415) Alcohol dehydrogenase
class III (Glutathione-dependent formaldehyde
dehydrogenase) (FDH) (FALDH) (Octanol dehydrogenase) ,
partial (8%)
Length = 640
Score = 118 bits (295), Expect = 4e-27
Identities = 57/125 (45%), Positives = 82/125 (65%)
Frame = +2
Query: 207 LSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADF 266
+ VA+GA+ +GAS+IIGVD NP K A+ G+T+ ++P+ E+P+ + I ITDGG +
Sbjct: 137 IHVAEGARARGASKIIGVDINPDKFIKAQTMGVTDFINPDDEEKPVYERIREITDGGVHY 316
Query: 267 SFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKP 326
SFEC G+ D++ A S +GWGLTV LGV ++ H LL GR + G +FGG+K
Sbjct: 317 SFECTGNVDVLRDAFLSSHEGWGLTVILGVHASPKLLPIHPMELLDGRNIVGCVFGGFKG 496
Query: 327 KSDLP 331
+S LP
Sbjct: 497 RSQLP 511
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,117,494
Number of Sequences: 63676
Number of extensions: 253788
Number of successful extensions: 1174
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 1132
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1139
length of query: 339
length of database: 12,639,632
effective HSP length: 98
effective length of query: 241
effective length of database: 6,399,384
effective search space: 1542251544
effective search space used: 1542251544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0383.12


