

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0381.8
(460 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
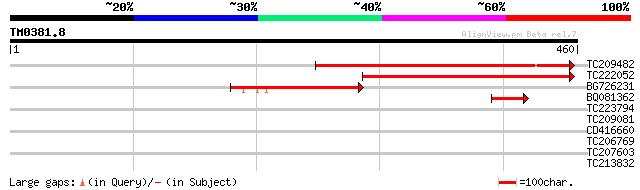
TC209482 355 2e-98
TC222052 223 2e-58
BG726231 127 1e-29
BQ081362 48 8e-06
TC223794 32 0.59
TC209081 similar to UP|Q8LD35 (Q8LD35) Nuclear matrix protein 1,... 28 6.6
CD416660 28 6.6
TC206769 similar to UP|Q93ZG9 (Q93ZG9) AT4g25340/T30C3_20, parti... 28 6.6
TC207603 UP|C773_SOYBN (O48928) Cytochrome P450 77A3 , complete 28 8.6
TC213832 similar to UP|AGA1_YEAST (P32323) A-agglutinin attachme... 28 8.6
>TC209482
Length = 921
Score = 355 bits (912), Expect = 2e-98
Identities = 173/210 (82%), Positives = 196/210 (92%)
Frame = +1
Query: 249 YISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVCGILGTLGGGLILDRMTSTISN 308
Y++YNFVIGAYSYWGPKAGYSIYHM+N DLLFGGITIVCGI+GTL GGL LDR++STISN
Sbjct: 1 YLAYNFVIGAYSYWGPKAGYSIYHMNNADLLFGGITIVCGIVGTLAGGLFLDRISSTISN 180
Query: 309 AFKLLSGATLLGAIFCLVAFLFKNLSGFIVFFSIGELLIFATQAPVNYVSLRCVKPSLRP 368
AFKLLSGAT LGAIFCL+AFLFK+LSGFIVFFS+GELLIF TQAPVNYVSLRCVKPSLRP
Sbjct: 181 AFKLLSGATFLGAIFCLIAFLFKSLSGFIVFFSMGELLIFVTQAPVNYVSLRCVKPSLRP 360
Query: 369 LSMAISTVSIHIFGDVPSAPLVGVLQDRINDWRKSALCLTSVFFLAAGIWFIGIFLRTVD 428
LSMAISTVSIH+FGDVPS+PLVGVLQD INDWRK+ALCLTS+FFLAA IWFIGIFL++ D
Sbjct: 361 LSMAISTVSIHVFGDVPSSPLVGVLQDHINDWRKTALCLTSIFFLAAVIWFIGIFLKS-D 537
Query: 429 LSTEDDEDQSATSLRGKMKPLLEGNSDASS 458
+ +DDE+QSAT+ GK+ PLL+G+SD +S
Sbjct: 538 VYDKDDEEQSATTRGGKLTPLLDGSSDTTS 627
>TC222052
Length = 832
Score = 223 bits (567), Expect = 2e-58
Identities = 110/172 (63%), Positives = 135/172 (77%)
Frame = +2
Query: 287 CGILGTLGGGLILDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIVFFSIGELL 346
CGILGT+ GG +LD MT+TISNAFKLLS AT +G C AFLFK+ GF+ F++GELL
Sbjct: 2 CGILGTVAGGFVLDFMTNTISNAFKLLSIATFIGGACCFGAFLFKSQYGFLALFAVGELL 181
Query: 347 IFATQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDRINDWRKSALC 406
+FATQ PVNYV L CVKPSLRPLSMA+STV+IHIFGDVPS+PLVG++QD+IN+WR +AL
Sbjct: 182 VFATQGPVNYVCLHCVKPSLRPLSMAMSTVAIHIFGDVPSSPLVGLMQDKINNWRTTALI 361
Query: 407 LTSVFFLAAGIWFIGIFLRTVDLSTEDDEDQSATSLRGKMKPLLEGNSDASS 458
LT++FF AA IWFIGIFL +VD ED E + ++ R PLLE N+ +S
Sbjct: 362 LTTIFFPAAAIWFIGIFLPSVDRFNEDSEHEVSSVERTSTAPLLEENTAETS 517
>BG726231
Length = 415
Score = 127 bits (319), Expect = 1e-29
Identities = 63/119 (52%), Positives = 90/119 (74%), Gaps = 11/119 (9%)
Frame = +1
Query: 180 GFAPLESKK--TISSIETNVSE----HGDDGVLA-----QDQAFIRGTRSTSKLRNQFTR 228
GFAP +S+K T+ ++ + VS+ +G D L+ +D++ +RS + +QF+R
Sbjct: 58 GFAPTDSEKALTLGTVASEVSDVGASNGKDEALSLKAEFRDKSSHEPSRSKCTILDQFSR 237
Query: 229 FSKDMQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVC 287
F KDM+ELL D+V+++NVLGYI+YNFVIGAYSYWGPKAGYSIY+M+N D++FGGIT+VC
Sbjct: 238 FLKDMKELLLDKVFVVNVLGYIAYNFVIGAYSYWGPKAGYSIYNMTNADMMFGGITVVC 414
>BQ081362
Length = 421
Score = 48.1 bits (113), Expect = 8e-06
Identities = 19/30 (63%), Positives = 26/30 (86%)
Frame = +1
Query: 392 VLQDRINDWRKSALCLTSVFFLAAGIWFIG 421
++QD+IN+WR +AL LT++FF AA IWFIG
Sbjct: 76 IMQDKINNWRTTALILTTIFFPAAAIWFIG 165
>TC223794
Length = 457
Score = 32.0 bits (71), Expect = 0.59
Identities = 13/20 (65%), Positives = 17/20 (85%)
Frame = +3
Query: 332 NLSGFIVFFSIGELLIFATQ 351
++ GF+ FSIGELL+FATQ
Sbjct: 3 SMYGFLALFSIGELLVFATQ 62
>TC209081 similar to UP|Q8LD35 (Q8LD35) Nuclear matrix protein 1, partial
(65%)
Length = 876
Score = 28.5 bits (62), Expect = 6.6
Identities = 14/31 (45%), Positives = 19/31 (61%)
Frame = -1
Query: 223 RNQFTRFSKDMQELLYDQVYIINVLGYISYN 253
R QFT F K + L + VY ++VLGY+ N
Sbjct: 138 RKQFTFFRKYLSLLFCNGVYQLDVLGYLLVN 46
>CD416660
Length = 542
Score = 28.5 bits (62), Expect = 6.6
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -1
Query: 421 GIFLRTVDLSTEDDEDQSATSLRGKMKPLLEGNSDASS 458
GIFL +VD ED E Q + R PLL+ + +S
Sbjct: 272 GIFLHSVDRFDEDSEHQVSNVERSNTMPLLQEKTGETS 159
>TC206769 similar to UP|Q93ZG9 (Q93ZG9) AT4g25340/T30C3_20, partial (10%)
Length = 1057
Score = 28.5 bits (62), Expect = 6.6
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Frame = -1
Query: 134 VGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYVYGFAPLE--SKKTIS 191
VG+G+ S SL+ + A + L+ + ++ G P S TIS
Sbjct: 436 VGIGKPSSSSLSTRSVSPLATIGSPELLLSFSFN*ATCFFFFNFLVGSYPFSGLSSSTIS 257
Query: 192 SIETNVSEHGDDGVLAQDQAFIRGTRSTSKL 222
SI T + G+DG ++ + I+ + S L
Sbjct: 256 SITTPLFGTGEDGYMSVSLSTIKSSAYPSSL 164
>TC207603 UP|C773_SOYBN (O48928) Cytochrome P450 77A3 , complete
Length = 1929
Score = 28.1 bits (61), Expect = 8.6
Identities = 37/127 (29%), Positives = 56/127 (43%), Gaps = 2/127 (1%)
Frame = -1
Query: 19 LLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGDFNLNNFQ--DGVL 76
+L I I T+ G I NG + T + V +A S + + + +NF + L
Sbjct: 1539 ILAIMR*ICTVAIAKPGQILLPTPNGII--FTPVTPVMSASSPPEMNLSGSNFSGFNQFL 1366
Query: 77 SSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGV 136
S+ M G+ ++ FAS+ S+ P + V V T V GC + F T CR G+
Sbjct: 1365 GSSAMAGVYTST--FASIGMSYPPKVVGSVTACVSTKCVGGCFLRSSF--TTACR--YGI 1204
Query: 137 GEASFIS 143
SF S
Sbjct: 1203 FSTSFSS 1183
>TC213832 similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment subunit
precursor, partial (4%)
Length = 751
Score = 28.1 bits (61), Expect = 8.6
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 8/136 (5%)
Frame = -1
Query: 9 PNP-SWFTPKRLLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGD-- 65
P+P S + RLL+ F NT+N+ D+ ++ + PTC S T+ S
Sbjct: 688 PSPLSSVSSLRLLV*FSCCNTMNHFDKTVFDASLSLSSTPTCFSSSMSSTSTSSSSSSSS 509
Query: 66 ----FNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHS-PFRLIGVGLSVWTFAVAGCGI 120
++ L S M+ + + I + L S S R LSVW F+++
Sbjct: 508 SSVAMSVPENDPKHLRSNLMLRKHVKNCIISFLKPSVSAQLRRFF*FLSVWIFSIS---- 341
Query: 121 SFDFWSITICRMLVGV 136
SIT C++L V
Sbjct: 340 -----SITDCQLLHAV 308
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,050,896
Number of Sequences: 63676
Number of extensions: 304851
Number of successful extensions: 1456
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1451
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1455
length of query: 460
length of database: 12,639,632
effective HSP length: 101
effective length of query: 359
effective length of database: 6,208,356
effective search space: 2228799804
effective search space used: 2228799804
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0381.8


