

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0380.7
(155 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
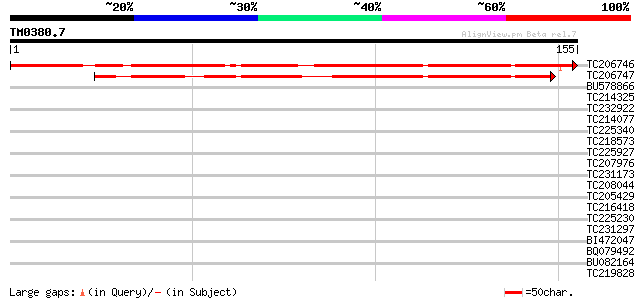
Sequences producing significant alignments: (bits) Value
TC206746 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_1... 165 9e-42
TC206747 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_1... 119 6e-28
BU578866 32 0.094
TC214325 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {A... 31 0.21
TC232922 weakly similar to UP|Q84VZ3 (Q84VZ3) At5g52990, partial... 31 0.27
TC214077 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {A... 30 0.61
TC225340 similar to UP|Q9SPI1 (Q9SPI1) Splicing factor SR1, part... 30 0.61
TC218573 weakly similar to UP|B1AR_MELGA (P07700) Beta-1 adrener... 30 0.61
TC225927 similar to UP|Q8VWX5 (Q8VWX5) Ribosome-like protein (Fr... 29 0.79
TC207976 similar to UP|Q6YU70 (Q6YU70) Cyclin G-associated kinas... 29 0.79
TC231173 29 1.0
TC208044 29 1.0
TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich prote... 29 1.0
TC216418 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor... 29 1.0
TC225230 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein ... 29 1.0
TC231297 28 1.8
BI472047 28 1.8
BQ079492 28 1.8
BU082164 weakly similar to GP|11994124|d receptor protein kinase... 28 1.8
TC219828 similar to GB|AAN15551.1|23198048|BT000232 expressed pr... 28 1.8
>TC206746 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_120, partial
(41%)
Length = 685
Score = 165 bits (417), Expect = 9e-42
Identities = 101/160 (63%), Positives = 112/160 (69%), Gaps = 5/160 (3%)
Frame = +3
Query: 1 MSSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV 60
MSS+G Q RS F S+ LPVS SRS SE GLR+RLSS + S ++S+
Sbjct: 78 MSSMGESQQRSLFGSTKASS---LPVSTSRS--DSEGGLRRRLSSLSLKIQLESPNSSS- 239
Query: 61 WSSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRG 120
WS FPRSKSLSSMGDYA T WWDW+WAWILSRK IFA DLE+NDQETKLL SHNRG
Sbjct: 240 WS-FPRSKSLSSMGDYART----WWDWSWAWILSRKLIFATDLEMNDQETKLL-ASHNRG 401
Query: 121 TWRHVFYKLRSEIRRFIPPTNSHHHLPRT-----TPQADS 155
TWRHVFYKLRSEIRRFI T+ H LP+T TPQ S
Sbjct: 402 TWRHVFYKLRSEIRRFI-NTSDHTTLPKTYHHKSTPQVFS 518
>TC206747 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_120, partial
(41%)
Length = 885
Score = 119 bits (298), Expect = 6e-28
Identities = 76/126 (60%), Positives = 82/126 (64%)
Frame = +3
Query: 24 LPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKN 83
LPVS S E GLR+RLSS + S WS F RSKSLSSMGDY T
Sbjct: 183 LPVSSR----SEEGGLRRRLSSLSL-----KIQDSPTWS-FSRSKSLSSMGDYVRTC--- 323
Query: 84 WWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTNSH 143
WAWILSRK IFA DLE+NDQ+TKLL GS NRGTW HVFYKLRSEIRRFI T+ H
Sbjct: 324 -----WAWILSRKLIFATDLEMNDQKTKLL-GSPNRGTWSHVFYKLRSEIRRFI-STSDH 482
Query: 144 HHLPRT 149
LP+T
Sbjct: 483 ATLPKT 500
>BU578866
Length = 409
Score = 32.3 bits (72), Expect = 0.094
Identities = 20/64 (31%), Positives = 34/64 (52%), Gaps = 2/64 (3%)
Frame = +2
Query: 19 SNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKS--LSSMGDY 76
S K+ + +++ S+H L K+ + ++ PA + ++ V SSFP S LS+M D
Sbjct: 53 SPRKRKSEAAAKTPSPSKHSLEKKRAGWDLPPAGTNNPSAVVSSSFPVSNCAVLSNMHDV 232
Query: 77 AGTS 80
TS
Sbjct: 233 VSTS 244
>TC214325 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {Arabidopsis
thaliana;} , partial (48%)
Length = 1186
Score = 31.2 bits (69), Expect = 0.21
Identities = 23/63 (36%), Positives = 28/63 (43%)
Frame = -3
Query: 41 KRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKPIFA 100
K L+SF+ T AAS AS +S R SS A + WW AW RK +
Sbjct: 242 KVLASFSKTLQAASRQASRATNSKGRILMASSGLGLASARVWWWWRRILAWSS*RKGVTE 63
Query: 101 RDL 103
R L
Sbjct: 62 RSL 54
>TC232922 weakly similar to UP|Q84VZ3 (Q84VZ3) At5g52990, partial (15%)
Length = 584
Score = 30.8 bits (68), Expect = 0.27
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = +2
Query: 127 YKLRSEIRRFIPPTNSHHHLP 147
+++RS R+ +PPTN+HH P
Sbjct: 29 HRIRSSFRQTLPPTNNHHFAP 91
>TC214077 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {Arabidopsis
thaliana;} , partial (50%)
Length = 1125
Score = 29.6 bits (65), Expect = 0.61
Identities = 23/65 (35%), Positives = 28/65 (42%)
Frame = -3
Query: 41 KRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKPIFA 100
K L+SF+ T AAS AS +S R SS A + WW AW RK +
Sbjct: 493 KVLASFSKTLQAASRQASRATNSKGRILMASSGLGLASARVWWWWRRILAWSS*RKGVTE 314
Query: 101 RDLEL 105
EL
Sbjct: 313 GSREL 299
>TC225340 similar to UP|Q9SPI1 (Q9SPI1) Splicing factor SR1, partial (81%)
Length = 1349
Score = 29.6 bits (65), Expect = 0.61
Identities = 25/67 (37%), Positives = 30/67 (44%)
Frame = +1
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G RS P+ S ++ S SRS+ S R L N +P SAS S S
Sbjct: 871 GKSSQRSPAKSPSRSPSRSRSRSRSRSLSGSRSRSRSPLPPRNKSPKKRSASRSPS-RSR 1047
Query: 65 PRSKSLS 71
RSKSLS
Sbjct: 1048SRSKSLS 1068
>TC218573 weakly similar to UP|B1AR_MELGA (P07700) Beta-1 adrenergic receptor
(Beta-1 adrenoceptor) (Beta-1 adrenoreceptor) (Beta-T),
partial (4%)
Length = 578
Score = 29.6 bits (65), Expect = 0.61
Identities = 20/56 (35%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Frame = +3
Query: 97 PIFARDLELNDQ-ETKLLGGSHN-RGTWRHVFYKLRSEIRR--FIPPTNSHHHLPR 148
P+ +L+ ND+ E++LL + N R WR + S I F+PP HLPR
Sbjct: 330 PLAVTNLQRNDETESRLLHRAANGRRRWRRRRHAYSSVINSVTFLPPIRQRSHLPR 497
>TC225927 similar to UP|Q8VWX5 (Q8VWX5) Ribosome-like protein (Fragment),
partial (59%)
Length = 718
Score = 29.3 bits (64), Expect = 0.79
Identities = 23/80 (28%), Positives = 36/80 (44%), Gaps = 2/80 (2%)
Frame = -2
Query: 12 FFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLS 71
FF+ S N+ L HS + LRKRLSS ++ + +S + +++ S
Sbjct: 354 FFSSCLFSKNRLL--LHS*MTPALMTSLRKRLSSLSSG-SFSSTTTCTLYADRKNSVFAC 184
Query: 72 SMGDYA--GTSIKNWWDWTW 89
+ +A TS WW W W
Sbjct: 183 AWDGFATADTSANEWWWWPW 124
>TC207976 similar to UP|Q6YU70 (Q6YU70) Cyclin G-associated kinase-like
protein, partial (3%)
Length = 702
Score = 29.3 bits (64), Expect = 0.79
Identities = 19/59 (32%), Positives = 31/59 (52%), Gaps = 4/59 (6%)
Frame = +2
Query: 29 SRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFP----RSKSLSSMGDYAGTSIKN 83
+R+ S GL+ +S T+ A+ASAS+ W +FP + +SLS+ +KN
Sbjct: 56 ARTPSPSREGLK--ISPAVTSSASASASSDKSWQAFPEEPQQQRSLSAENTSKSVRVKN 226
>TC231173
Length = 548
Score = 28.9 bits (63), Expect = 1.0
Identities = 18/45 (40%), Positives = 22/45 (48%)
Frame = -1
Query: 26 VSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSL 70
VS S + S HGL KRL AAA +S+S W S S+
Sbjct: 200 VSPSPTTYISSHGLSKRLLRPLVVAAAARSSSSGTWLSLMDPNSI 66
>TC208044
Length = 802
Score = 28.9 bits (63), Expect = 1.0
Identities = 12/32 (37%), Positives = 18/32 (55%), Gaps = 1/32 (3%)
Frame = -2
Query: 66 RSKSLSSMGDYAGTSIKNW-WDWTWAWILSRK 96
R K + S G +G + + W W W WAW+ R+
Sbjct: 114 RMKGIWSWG-VSGEAFRAWDWAWAWAWLCRRR 22
>TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich protein, partial
(49%)
Length = 611
Score = 28.9 bits (63), Expect = 1.0
Identities = 12/37 (32%), Positives = 16/37 (42%)
Frame = -1
Query: 61 WSSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKP 97
W++F RS+ + WW W W W SR P
Sbjct: 482 WATFRRSRCRN-----------RWWFWRWTWWRSRWP 405
>TC216418 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor NtEIG-D48,
partial (51%)
Length = 1461
Score = 28.9 bits (63), Expect = 1.0
Identities = 7/12 (58%), Positives = 8/12 (66%)
Frame = -2
Query: 84 WWDWTWAWILSR 95
WW W WAW+ R
Sbjct: 269 WWAWAWAWVWER 234
>TC225230 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein
(Fibrillarin 2) (AtFib2), partial (66%)
Length = 773
Score = 28.9 bits (63), Expect = 1.0
Identities = 7/15 (46%), Positives = 11/15 (72%)
Frame = +3
Query: 78 GTSIKNWWDWTWAWI 92
G+S ++WW W W W+
Sbjct: 27 GSSKRSWWFWWWRWV 71
>TC231297
Length = 979
Score = 28.1 bits (61), Expect = 1.8
Identities = 7/13 (53%), Positives = 8/13 (60%)
Frame = -1
Query: 79 TSIKNWWDWTWAW 91
+S WWDW W W
Sbjct: 355 SSTFGWWDWDWGW 317
>BI472047
Length = 399
Score = 28.1 bits (61), Expect = 1.8
Identities = 18/70 (25%), Positives = 32/70 (45%), Gaps = 3/70 (4%)
Frame = +1
Query: 28 HSRSVDSSEHGLRK---RLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNW 84
HS + ++ L+K S+F ++ +A+A+A+ ++ P SL G +
Sbjct: 28 HSFLITMPQNQLQKIPTNNSTFQSSRFSAAAAATTTTTTQPLRSSLHIGDSRVG*VLLAQ 207
Query: 85 WDWTWAWILS 94
DW W W S
Sbjct: 208MDWLWLWFTS 237
>BQ079492
Length = 426
Score = 28.1 bits (61), Expect = 1.8
Identities = 15/37 (40%), Positives = 24/37 (64%)
Frame = +3
Query: 44 SSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTS 80
+S +TTP+++S+S+S+ S+ P S SS GTS
Sbjct: 267 TSSSTTPSSSSSSSSSPSSATPSPSSSSSSSPPPGTS 377
>BU082164 weakly similar to GP|11994124|d receptor protein kinase
{Arabidopsis thaliana}, partial (12%)
Length = 459
Score = 28.1 bits (61), Expect = 1.8
Identities = 17/71 (23%), Positives = 31/71 (42%)
Frame = -2
Query: 19 SNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAG 78
S PV+ ++ EH L K LS+ + + S + + P++ S +
Sbjct: 455 STRISFPVAGLPDLEEPEHMLSKALST--NVSLLLTEALSRLTAITPKTMSKDADIATIA 282
Query: 79 TSIKNWWDWTW 89
+++N W WTW
Sbjct: 281 DTLRNLWRWTW 249
>TC219828 similar to GB|AAN15551.1|23198048|BT000232 expressed protein
{Arabidopsis thaliana;} , partial (95%)
Length = 996
Score = 28.1 bits (61), Expect = 1.8
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +2
Query: 85 WDWTWAWILSRKPIFARDLELNDQETKL 112
W W AW LS P+ R+ E ++ TKL
Sbjct: 233 WSWFCAWSLSGSPLPPREEEASEPCTKL 316
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.127 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,960,508
Number of Sequences: 63676
Number of extensions: 138840
Number of successful extensions: 2068
Number of sequences better than 10.0: 112
Number of HSP's better than 10.0 without gapping: 1831
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2024
length of query: 155
length of database: 12,639,632
effective HSP length: 89
effective length of query: 66
effective length of database: 6,972,468
effective search space: 460182888
effective search space used: 460182888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0380.7


