

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0377b.5
(372 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
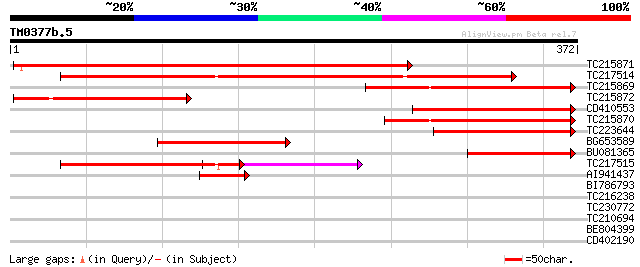
Score E
Sequences producing significant alignments: (bits) Value
TC215871 weakly similar to UP|O48971 (O48971) Aldose-1-epimerase... 411 e-115
TC217514 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like p... 287 6e-78
TC215869 weakly similar to UP|Q947H4 (Q947H4) Non-cell-autonomou... 213 1e-55
TC215872 similar to UP|Q947H4 (Q947H4) Non-cell-autonomous prote... 202 2e-52
CD410553 similar to GP|15824567|gb| non-cell-autonomous protein ... 196 1e-50
TC215870 weakly similar to UP|Q947H4 (Q947H4) Non-cell-autonomou... 182 3e-46
TC223644 similar to UP|Q947H4 (Q947H4) Non-cell-autonomous prote... 174 8e-44
BG653589 weakly similar to GP|15824565|gb| non-cell-autonomous p... 148 4e-36
BU081365 weakly similar to GP|15824567|gb| non-cell-autonomous p... 115 3e-26
TC217515 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like p... 114 9e-26
AI941437 44 9e-05
BI786793 30 1.8
TC216238 30 2.3
TC230772 similar to UP|O24648 (O24648) Gibberellin 3 beta-hydrox... 28 6.7
TC210694 28 8.7
BE804399 similar to GP|23170542|gb| CG12586-PA {Drosophila melan... 28 8.7
CD402190 28 8.7
>TC215871 weakly similar to UP|O48971 (O48971) Aldose-1-epimerase-like
protein , partial (58%)
Length = 927
Score = 411 bits (1057), Expect = e-115
Identities = 199/264 (75%), Positives = 228/264 (85%), Gaps = 2/264 (0%)
Frame = +1
Query: 3 TKIF--VLLCLLFLAASGFVNGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDK 60
TKIF +LLCL+ ++SGFVN S ++++ I +FELKKGDLSL+V+NWGA+++SLV+PDK
Sbjct: 136 TKIFFVLLLCLIIASSSGFVNCSDVEKEEKIGIFELKKGDLSLRVTNWGASILSLVIPDK 315
Query: 61 HGKLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHG 120
+GKLGDVVLGYDS+K Y+ND+TYFG T GRVANRIGGAQFTLNGIHYKL+ANEGNNTLHG
Sbjct: 316 NGKLGDVVLGYDSVKDYSNDTTYFGATVGRVANRIGGAQFTLNGIHYKLVANEGNNTLHG 495
Query: 121 GPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMK 180
G RGFSDVLWKV+RY KEG P ITFSYHS DGE+GFPGDLLVT +YIL KN L I+MK
Sbjct: 496 GARGFSDVLWKVKRYQKEGPSPSITFSYHSIDGEQGFPGDLLVTGNYILTGKNQLVILMK 675
Query: 181 AKALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTP 240
K LNKPTPVNL NHAYWNLG HNSGNIL EVVQIFGS+IT +D+ LIPTG+ ASVKGT
Sbjct: 676 GKTLNKPTPVNLANHAYWNLGGHNSGNILNEVVQIFGSKITPVDNKLIPTGKFASVKGTA 855
Query: 241 YDFLKPQVVGARINQLPKTNGYDI 264
YDFLKPQ VG+RINQL T GYDI
Sbjct: 856 YDFLKPQTVGSRINQLAGTKGYDI 927
>TC217514 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like protein,
partial (85%)
Length = 1207
Score = 287 bits (734), Expect = 6e-78
Identities = 149/300 (49%), Positives = 195/300 (64%), Gaps = 1/300 (0%)
Frame = +2
Query: 34 FELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND-STYFGTTAGRVA 92
FEL G + + +SN G T++SL +P K G L D+VLG +S++ Y + YFG GRVA
Sbjct: 128 FELNNGSMQVLLSNLGCTIISLSVPGKDGLLSDIVLGLESVESYQKGLAPYFGCIVGRVA 307
Query: 93 NRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFD 152
NRI +FTL+GI Y L N+ N+LHGG GF +W+V Y K+G+ P ITF HS D
Sbjct: 308 NRIKDGKFTLDGIQYSLPINKPPNSLHGGNVGFDKKVWEVVEY-KKGNTPSITFRCHSHD 484
Query: 153 GEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGEV 212
GEEG+PGD+ VT +Y L S +L + M+ +KPT VNL H YWNL HNSGNIL
Sbjct: 485 GEEGYPGDVTVTATYTLTSSTTLRLDMEGVPKDKPTIVNLAQHTYWNLAGHNSGNILDHS 664
Query: 213 VQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGK 272
+Q++ + IT +D + +PTG+I VKGTP+DF +G I+Q+ GYD NYVLD G+
Sbjct: 665 IQLWANHITPVDENTVPTGKIMPVKGTPFDFTSENRIGNTISQVGL--GYDHNYVLDCGE 838
Query: 273 SKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQV 332
K ++ AA V D S RV+ L+TNAPG+QFYT N V +GKGG +Y AGLCLE QV
Sbjct: 839 EKEGLRHAAKVRDPSSSRVLNLWTNAPGVQFYTTNYVNGVQGKGGAIYGKHAGLCLEIQV 1018
>TC215869 weakly similar to UP|Q947H4 (Q947H4) Non-cell-autonomous protein
pathway2, partial (25%)
Length = 768
Score = 213 bits (541), Expect = 1e-55
Identities = 106/138 (76%), Positives = 116/138 (83%)
Frame = +2
Query: 234 ASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMK 293
ASVKGT YDF KPQ VG+RINQL +T GYDINYVLDG K + IKLAAIV DKKSGRV++
Sbjct: 5 ASVKGTAYDFPKPQTVGSRINQLAETKGYDINYVLDGEKGQK-IKLAAIVQDKKSGRVLE 181
Query: 294 LFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPY 353
LFTNAPGLQFYT N VK+ KGKGG+VYQ +GLCLESQ FPDSVN PNFPS+IVTPEKPY
Sbjct: 182 LFTNAPGLQFYTGNYVKDLKGKGGYVYQSHSGLCLESQAFPDSVNQPNFPSTIVTPEKPY 361
Query: 354 KHVMLLKFSTKAPYAFSQ 371
KH ML KFSTK PY S+
Sbjct: 362 KHYMLFKFSTKNPYISSK 415
>TC215872 similar to UP|Q947H4 (Q947H4) Non-cell-autonomous protein pathway2,
partial (24%)
Length = 795
Score = 202 bits (515), Expect = 2e-52
Identities = 97/117 (82%), Positives = 110/117 (93%)
Frame = +1
Query: 3 TKIFVLLCLLFLAASGFVNGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHG 62
TKIF+LLCLLFLA+SGFV+GS+KK DHIRLFELKKGD S+KV+NWGATLVS++LPDK+G
Sbjct: 115 TKIFLLLCLLFLASSGFVHGSSKK--DHIRLFELKKGDFSIKVTNWGATLVSVILPDKNG 288
Query: 63 KLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLH 119
KLGD+VLGYDS K YTND++YFG T GRVANRIGGAQFTLNGIHYKL+ANEGNNTLH
Sbjct: 289 KLGDIVLGYDSPKAYTNDTSYFGATVGRVANRIGGAQFTLNGIHYKLLANEGNNTLH 459
>CD410553 similar to GP|15824567|gb| non-cell-autonomous protein pathway2;
plasmodesmal receptor {Nicotiana tabacum}, partial (23%)
Length = 460
Score = 196 bits (498), Expect = 1e-50
Identities = 94/107 (87%), Positives = 99/107 (91%)
Frame = -1
Query: 265 NYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRA 324
NYVLDG K G IKLAAIV+DKKSGRVMKLFTNAPGLQFYTAN VKN+KGKGGFVYQPR+
Sbjct: 451 NYVLDGEKGNGEIKLAAIVVDKKSGRVMKLFTNAPGLQFYTANFVKNDKGKGGFVYQPRS 272
Query: 325 GLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAPYAFSQ 371
LCLESQ FPDSVNHPNFPS+IVTP+KPYKHVMLLKFSTK PYAFSQ
Sbjct: 271 ALCLESQAFPDSVNHPNFPSTIVTPDKPYKHVMLLKFSTKVPYAFSQ 131
>TC215870 weakly similar to UP|Q947H4 (Q947H4) Non-cell-autonomous protein
pathway2, partial (27%)
Length = 631
Score = 182 bits (461), Expect = 3e-46
Identities = 90/125 (72%), Positives = 99/125 (79%)
Frame = +3
Query: 247 QVVGARINQLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTA 306
Q RINQL T GYDINYV+DG K + IKLAAIV DKK GRV++LFTNAPGLQFYT
Sbjct: 3 QTXXXRINQLXXTKGYDINYVIDGEKGQK-IKLAAIVQDKKXGRVLELFTNAPGLQFYTG 179
Query: 307 NKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAP 366
N +K+ KGKGG+VY AGLCLESQ FPDSVNHPNFPS+IVT EKPYKH ML KFSTK P
Sbjct: 180 NYIKDLKGKGGYVYNAHAGLCLESQAFPDSVNHPNFPSTIVTQEKPYKHYMLFKFSTKNP 359
Query: 367 YAFSQ 371
Y S+
Sbjct: 360 YISSK 374
>TC223644 similar to UP|Q947H4 (Q947H4) Non-cell-autonomous protein pathway2,
partial (24%)
Length = 404
Score = 174 bits (440), Expect = 8e-44
Identities = 83/93 (89%), Positives = 88/93 (94%)
Frame = -2
Query: 279 LAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVN 338
LAAIV+DKKSGRVMKLFTNAPGLQFYTAN VKNEKGKGGFVYQPR+ LCLESQ FPDSVN
Sbjct: 403 LAAIVVDKKSGRVMKLFTNAPGLQFYTANFVKNEKGKGGFVYQPRSALCLESQAFPDSVN 224
Query: 339 HPNFPSSIVTPEKPYKHVMLLKFSTKAPYAFSQ 371
HPNFPS+IVT EKPYKHV+LLKFSTK P+AFSQ
Sbjct: 223 HPNFPSTIVTTEKPYKHVLLLKFSTKVPHAFSQ 125
>BG653589 weakly similar to GP|15824565|gb| non-cell-autonomous protein
pathway1; plasmodesmal receptor {Nicotiana tabacum},
partial (20%)
Length = 358
Score = 148 bits (373), Expect = 4e-36
Identities = 71/87 (81%), Positives = 75/87 (85%)
Frame = +1
Query: 98 AQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGF 157
AQFTLNGI+YKL ANEGNNTLHGG RGFSDVLWKV+RY KEG P ITFSYHS DGE+GF
Sbjct: 16 AQFTLNGIYYKLFANEGNNTLHGGARGFSDVLWKVKRYQKEGPSPSITFSYHSVDGEQGF 195
Query: 158 PGDLLVTVSYILGSKNSLSIIMKAKAL 184
PGDLLVTVSYIL KN L I+MK KAL
Sbjct: 196 PGDLLVTVSYILTGKNQLVILMKGKAL 276
Score = 37.0 bits (84), Expect = 0.014
Identities = 16/22 (72%), Positives = 17/22 (76%)
Frame = +2
Query: 350 EKPYKHVMLLKFSTKAPYAFSQ 371
EKPYKH ML KFSTK PY S+
Sbjct: 266 EKPYKHYMLFKFSTKNPYISSK 331
>BU081365 weakly similar to GP|15824567|gb| non-cell-autonomous protein
pathway2; plasmodesmal receptor {Nicotiana tabacum},
partial (20%)
Length = 387
Score = 115 bits (288), Expect = 3e-26
Identities = 53/71 (74%), Positives = 59/71 (82%)
Frame = +2
Query: 301 LQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLK 360
+QFYT N VK+ KGKGG+VYQ +GLCLESQ FPDSVN PNFPS+IVTPEKPYKH ML K
Sbjct: 107 MQFYTGNYVKDLKGKGGYVYQSHSGLCLESQAFPDSVNQPNFPSTIVTPEKPYKHYMLFK 286
Query: 361 FSTKAPYAFSQ 371
FSTK PY S+
Sbjct: 287 FSTKNPYISSK 319
>TC217515 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like protein,
partial (54%)
Length = 726
Score = 114 bits (284), Expect = 9e-26
Identities = 58/122 (47%), Positives = 78/122 (63%), Gaps = 1/122 (0%)
Frame = +1
Query: 34 FELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND-STYFGTTAGRVA 92
FEL G + + ++N G T++SL +P K G L DVVLG +S++ Y + YFG GRVA
Sbjct: 130 FELNNGSMQVLLTNLGCTIISLSVPGKDGVLSDVVLGLESVESYQKGLAPYFGCIVGRVA 309
Query: 93 NRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFD 152
NRI +FTL+GIHY L N N+LHGG GF +W++ + K+G+ P ITF HS D
Sbjct: 310 NRIKDGKFTLDGIHYSLPINNPPNSLHGGNVGFDKKVWEIVEH-KKGETPSITFKCHSHD 486
Query: 153 GE 154
GE
Sbjct: 487 GE 492
Score = 81.3 bits (199), Expect = 7e-16
Identities = 40/110 (36%), Positives = 62/110 (56%), Gaps = 5/110 (4%)
Frame = +2
Query: 127 DVLWKVERY-----VKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKA 181
++L + RY +++ P+ S EG+PGD+ VT +Y L S +L + M+
Sbjct: 395 EMLGLIRRYGR*LNIRKAKPPQSLLSATVMMESEGYPGDITVTATYTLTSSTTLRLDMEE 574
Query: 182 KALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRITVLDSHLIPTG 231
+KPT VNL H YWNL HNSGNIL + ++ + IT ++ + +PTG
Sbjct: 575 VPKDKPTIVNLAQHTYWNLAGHNSGNILDHSIHLWANHITPVNENTVPTG 724
>AI941437
Length = 100
Score = 44.3 bits (103), Expect = 9e-05
Identities = 19/33 (57%), Positives = 23/33 (69%)
Frame = +2
Query: 125 FSDVLWKVERYVKEGDKPRITFSYHSFDGEEGF 157
F DVL + RY +G +P ITF YHS DGE+GF
Sbjct: 2 FMDVLCRETRYELDGPRPNITFRYHSVDGEQGF 100
>BI786793
Length = 426
Score = 30.0 bits (66), Expect = 1.8
Identities = 14/24 (58%), Positives = 17/24 (70%)
Frame = +2
Query: 90 RVANRIGGAQFTLNGIHYKLIANE 113
RVANRI +FTL+GI Y L N+
Sbjct: 350 RVANRIKDGKFTLDGIQYSLPINK 421
>TC216238
Length = 1821
Score = 29.6 bits (65), Expect = 2.3
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Frame = -1
Query: 279 LAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKG-GFVYQPRAGLCLESQVFPDSV 337
++ ++++K SG F A + + +E G G G QP L L S +FP S
Sbjct: 360 ISPLLLNKSSGGTKGPFAAAATVTGAGIPQDIDEAGNGTGLTKQPAGALQLNSPLFPSSA 181
Query: 338 NHPNFPSS 345
P+F S+
Sbjct: 180 LSPSFKSN 157
>TC230772 similar to UP|O24648 (O24648) Gibberellin 3 beta-hydroxylase
(2-oxoglutarate-dependent dioxygenase), partial (85%)
Length = 1264
Score = 28.1 bits (61), Expect = 6.7
Identities = 14/67 (20%), Positives = 32/67 (46%)
Frame = -3
Query: 131 KVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPV 190
++ + +K G F H F GEEG + ++ + G+ + ++ + + P P+
Sbjct: 395 ELAKMLKRGSNNCEAFRPHEF-GEEGRDASEAIAINTVRGTSCFVFLVRRERE*GLPNPL 219
Query: 191 NLINHAY 197
N++ A+
Sbjct: 218 NVLEEAH 198
>TC210694
Length = 778
Score = 27.7 bits (60), Expect = 8.7
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = -3
Query: 303 FYTANKVKNEKGKGGFVY 320
F+ +N KNEKG GG VY
Sbjct: 278 FHCSNTTKNEKGGGGVVY 225
>BE804399 similar to GP|23170542|gb| CG12586-PA {Drosophila melanogaster},
partial (3%)
Length = 394
Score = 27.7 bits (60), Expect = 8.7
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 2/62 (3%)
Frame = +2
Query: 299 PGLQFYTANKVKNEKGKGGFVYQPRAGLC-LESQVFPDSVNHPNFPSSIVTPEKPY-KHV 356
P L+F+TA +++ + F+ QPR L P +PNFP P P+ +H+
Sbjct: 209 PPLRFHTAPHLQSPQNPSLFLTQPRNPLVPRPPPQQPPHPPNPNFPLQNSHPPCPHRRHL 388
Query: 357 ML 358
ML
Sbjct: 389 ML 394
>CD402190
Length = 504
Score = 27.7 bits (60), Expect = 8.7
Identities = 14/58 (24%), Positives = 25/58 (42%)
Frame = -2
Query: 297 NAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYK 354
N P L+ YT+ G Q L + P ++ FP+S+++ E+P +
Sbjct: 482 NLPSLRSYTSRSKMELMKVGNLCCQLSRSTRLFQERHPLNIRQAGFPTSVISTERPIR 309
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,624,751
Number of Sequences: 63676
Number of extensions: 206806
Number of successful extensions: 941
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 933
length of query: 372
length of database: 12,639,632
effective HSP length: 99
effective length of query: 273
effective length of database: 6,335,708
effective search space: 1729648284
effective search space used: 1729648284
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0377b.5


