

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372b.7
(224 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
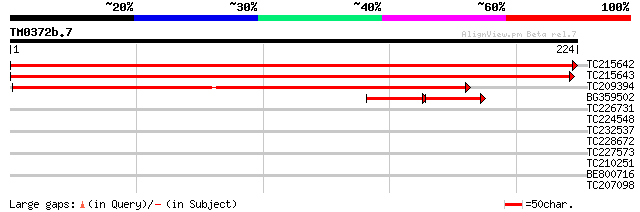
Score E
Sequences producing significant alignments: (bits) Value
TC215642 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%) 414 e-116
TC215643 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%) 404 e-113
TC209394 weakly similar to GB|AAB64690.1|603403|SCE8229 Yer163cp... 207 4e-54
BG359502 similar to GP|2827524|emb| predicted protein {Arabidops... 50 1e-14
TC226731 31 0.51
TC224548 similar to UP|Q8H8S1 (Q8H8S1) Ribosomal protein L15, co... 29 1.5
TC232537 homologue to GB|AAH40943.1|26454860|BC040943 WAS protei... 28 3.3
TC228672 27 5.7
TC227573 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis r... 27 7.4
TC210251 similar to UP|Q9FME7 (Q9FME7) Kinesin-like protein, par... 27 9.7
BE800716 27 9.7
TC207098 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, parti... 27 9.7
>TC215642 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%)
Length = 1146
Score = 414 bits (1064), Expect = e-116
Identities = 190/224 (84%), Positives = 207/224 (91%)
Frame = +1
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MVFWVFGYGSLVWNPGF+YDEK+IG+IKDYRRVFDLACIDHRGTPE+PARTCTLEEKEG
Sbjct: 145 MVFWVFGYGSLVWNPGFDYDEKIIGFIKDYRRVFDLACIDHRGTPENPARTCTLEEKEGA 324
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGA YCVRGGPEKEKL MQYLERRECEYD KTLV+F+KEGDS HP LT VIVFTSTPD
Sbjct: 325 ICWGAAYCVRGGPEKEKLAMQYLERRECEYDRKTLVNFFKEGDSLHPTLTDVIVFTSTPD 504
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KVNNKYYLGPAPL+ MARQIATA+GPCGNNRDY+FLLEKAMH+IGHEDD+VIELA+EV+K
Sbjct: 505 KVNNKYYLGPAPLEDMARQIATAYGPCGNNRDYIFLLEKAMHDIGHEDDLVIELANEVKK 684
Query: 181 VLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEPIALDS 224
VLGVGNV+P +KK G Q+ H PH PIP LQLLP+PEPIALDS
Sbjct: 685 VLGVGNVVPKEKKQVGAAQLPHPPHVPIPTLQLLPLPEPIALDS 816
>TC215643 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%)
Length = 1100
Score = 404 bits (1038), Expect = e-113
Identities = 185/223 (82%), Positives = 205/223 (90%)
Frame = +1
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MVFWVFGYGSLVWNPGF+Y+EK+IG+IKDYRRVFDLACIDHRGTPE+PARTCTLEEKEG
Sbjct: 145 MVFWVFGYGSLVWNPGFDYNEKIIGFIKDYRRVFDLACIDHRGTPENPARTCTLEEKEGA 324
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGA YCVRGGPEKEKL MQYLERRECEYD KTLV+F+KEGDS HP LT VIVFTSTPD
Sbjct: 325 ICWGAAYCVRGGPEKEKLAMQYLERRECEYDRKTLVNFFKEGDSLHPTLTDVIVFTSTPD 504
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KVNNKYYLGPAPL+ MARQIATA+GPCGNNRDY+FLLEKAM++IGHEDD+VIELA+EVRK
Sbjct: 505 KVNNKYYLGPAPLEDMARQIATAYGPCGNNRDYIFLLEKAMYDIGHEDDLVIELANEVRK 684
Query: 181 VLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEPIALD 223
VLG+GNV+P +KKL G Q+ H PH IP LQL P+PEPIA+D
Sbjct: 685 VLGMGNVVPKEKKLVGAAQLPHPPHVTIPTLQLHPLPEPIAMD 813
>TC209394 weakly similar to GB|AAB64690.1|603403|SCE8229 Yer163cp
{Saccharomyces cerevisiae;} , partial (24%)
Length = 792
Score = 207 bits (526), Expect = 4e-54
Identities = 101/181 (55%), Positives = 130/181 (71%)
Frame = +3
Query: 2 VFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEI 61
V WVFGYGSL+ GF YDE+++G+IK YRRVF DHRGTPE P RT TLE EGEI
Sbjct: 6 VMWVFGYGSLISKAGFHYDERIVGFIKGYRRVFYQGSTDHRGTPEFPGRTVTLEPAEGEI 185
Query: 62 CWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDK 121
CWGA Y + + E ++ YLE RE +YD K +DFY + ++ PA++GV+V+ +TPDK
Sbjct: 186 CWGAAYKIAKKEDAETALV-YLEVREKQYDKKEYLDFYTDLTATSPAVSGVMVYIATPDK 362
Query: 122 VNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKV 181
+N YLGPA ++ +ARQI A GP G NRDYLF LEKA+H IG +D VI+LA+EVR++
Sbjct: 363 KSNVNYLGPASIEDIARQIIQAKGPAGPNRDYLFNLEKALHEIGCKDKHVIDLANEVRRI 542
Query: 182 L 182
L
Sbjct: 543 L 545
>BG359502 similar to GP|2827524|emb| predicted protein {Arabidopsis
thaliana}, partial (16%)
Length = 313
Score = 49.7 bits (117), Expect(2) = 1e-14
Identities = 20/24 (83%), Positives = 24/24 (99%)
Frame = +2
Query: 142 TAHGPCGNNRDYLFLLEKAMHNIG 165
TA+GPCGNNRDY+FLLEKAM++IG
Sbjct: 53 TAYGPCGNNRDYIFLLEKAMYDIG 124
Score = 46.6 bits (109), Expect(2) = 1e-14
Identities = 21/24 (87%), Positives = 24/24 (99%)
Frame = +3
Query: 165 GHEDDMVIELAHEVRKVLGVGNVL 188
GHEDD+VIELA+EVRKVLGVGNV+
Sbjct: 240 GHEDDLVIELANEVRKVLGVGNVV 311
>TC226731
Length = 782
Score = 30.8 bits (68), Expect = 0.51
Identities = 10/27 (37%), Positives = 18/27 (66%)
Frame = -2
Query: 196 GPVQIQHSPHAPIPALQLLPMPEPIAL 222
GP+Q++H+P P+P + L P+ +L
Sbjct: 118 GPIQMEHNPRGPVPVTRTLRAPKSYSL 38
>TC224548 similar to UP|Q8H8S1 (Q8H8S1) Ribosomal protein L15, complete
Length = 820
Score = 29.3 bits (64), Expect = 1.5
Identities = 16/60 (26%), Positives = 26/60 (42%)
Frame = -2
Query: 116 TSTPDKVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELA 175
++TP N Y++ L+ A + A PC N L + A H + D+ + LA
Sbjct: 285 SATPPNTNTVYHIALLSLEAQATSLVRAGRPCQANNGRLLPILPAPHPLQEAHDIRLLLA 106
>TC232537 homologue to GB|AAH40943.1|26454860|BC040943 WAS protein family,
member 2 {Homo sapiens;} , partial (4%)
Length = 496
Score = 28.1 bits (61), Expect = 3.3
Identities = 16/60 (26%), Positives = 27/60 (44%), Gaps = 2/60 (3%)
Frame = +3
Query: 162 HNIGHEDDMVIELAHEVRKVLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQ--LLPMPEP 219
H + ++ H + + +++P KK P QHSP +P+P Q L P +P
Sbjct: 27 HILSSSSSTIVHFIHSEKAKIQ-RHMIPQWKKQHQPSSCQHSPLSPLPNSQI*LAPFSQP 203
>TC228672
Length = 552
Score = 27.3 bits (59), Expect = 5.7
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +1
Query: 34 FDLACIDHRGTPEHPARTCTLEEKEGEICWGAVYCVRG 71
F + +HR T EH AR L+ G + GAV CV G
Sbjct: 145 FAITLREHRTTLEHAARLPLLKHDSGIVVMGAV-CVVG 255
>TC227573 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis regulatory
protein homolog, partial (24%)
Length = 439
Score = 26.9 bits (58), Expect = 7.4
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = +2
Query: 176 HEVRKVLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEP 219
H V+ + +LP+ + L GP+ P +P + LP P+P
Sbjct: 218 HLVQAIADALFILPSTEDLDGPLVTLPPPLTKLPREKHLPKPKP 349
>TC210251 similar to UP|Q9FME7 (Q9FME7) Kinesin-like protein, partial (4%)
Length = 1071
Score = 26.6 bits (57), Expect = 9.7
Identities = 17/59 (28%), Positives = 25/59 (41%), Gaps = 5/59 (8%)
Frame = -1
Query: 9 GSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPE-----HPARTCTLEEKEGEIC 62
G L+W F+Y + V DH G + HP T LE++EG++C
Sbjct: 396 GHLIWPVYFQYQKVVF------------LAHDHCGPQQAPQSVHPEATQHLEQQEGDLC 256
>BE800716
Length = 419
Score = 26.6 bits (57), Expect = 9.7
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = +3
Query: 191 DKKLSGPVQIQHSPHAPIPALQLLPMPEP 219
+++L+G + HS HAP L++LP +P
Sbjct: 171 EEELNGTNVLFHSVHAPFVNLEMLPSIQP 257
>TC207098 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (8%)
Length = 733
Score = 26.6 bits (57), Expect = 9.7
Identities = 23/88 (26%), Positives = 39/88 (44%), Gaps = 11/88 (12%)
Frame = +2
Query: 42 RGTPEHPARTCTLEEKEGEICWGAVYCVRG-----------GPEKEKLVMQYLERRECEY 90
RG P PAR T ++E ++ W AV +G GP +E Q++ E
Sbjct: 86 RGGP--PARARTSAQREAQVRWRAVQDEKGWAGWRVGFGALGP-RESAQFQWMNGYE--- 247
Query: 91 DHKTLVDFYKEGDSSHPALTGVIVFTST 118
V+F+ E + H L +++ T++
Sbjct: 248 -----VEFFSEDEDKHILLQRLVLNTNS 316
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,679,795
Number of Sequences: 63676
Number of extensions: 172284
Number of successful extensions: 914
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 903
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 913
length of query: 224
length of database: 12,639,632
effective HSP length: 94
effective length of query: 130
effective length of database: 6,654,088
effective search space: 865031440
effective search space used: 865031440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0372b.7


