

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
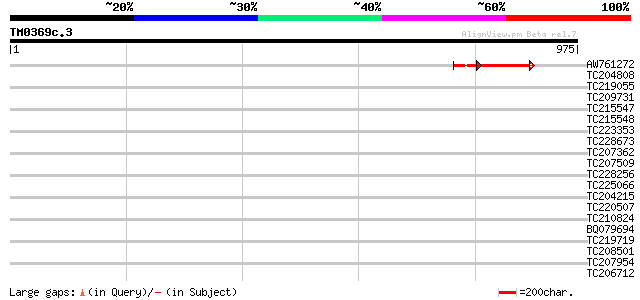
Query= TM0369c.3
(975 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW761272 131 7e-48
TC204808 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone g... 34 0.27
TC219055 similar to UP|PFTB_PEA (Q04903) Protein farnesyltransfe... 34 0.36
TC209731 similar to UP|Q8W3M8 (Q8W3M8) 6b-interacting protein 1,... 32 1.4
TC215547 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partia... 31 2.3
TC215548 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partia... 31 3.0
TC223353 weakly similar to UP|Q9SEW3 (Q9SEW3) Receptor-like prot... 31 3.0
TC228673 similar to UP|Q8L8M8 (Q8L8M8) Serine-rich protein, part... 31 3.0
TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protei... 30 4.0
TC207509 30 4.0
TC228256 weakly similar to UP|Q9ATY4 (Q9ATY4) MAP kinase phospha... 30 6.8
TC225066 homologue to UP|Q9M2F1 (Q9M2F1) Ribosomal protein S27 (... 30 6.8
TC204215 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin ... 30 6.8
TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), par... 30 6.8
TC210824 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper pro... 29 8.8
BQ079694 29 8.8
TC219719 homologue to UP|Q8GZD2 (Q8GZD2) Expansin, partial (97%) 29 8.8
TC208501 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F2... 29 8.8
TC207954 similar to UP|Q9M9V0 (Q9M9V0) F6A14.10 protein, partial... 29 8.8
TC206712 similar to GB|AAT41855.1|48310625|BT014872 At2g19340 {A... 29 8.8
>AW761272
Length = 414
Score = 131 bits (329), Expect(2) = 7e-48
Identities = 69/98 (70%), Positives = 81/98 (82%)
Frame = +1
Query: 805 FLLNARHLNELGSSYSLLEESCISSSMEPASINQAYHSRVSKNASWAKVHQSSNEIPPAS 864
F LNA HL ELGS+YSL EES ISS MEPA NQA+H++VSKNASW KVH+SSNEI A
Sbjct: 127 FFLNAEHLTELGSAYSLSEESHISS-MEPAGRNQAFHTKVSKNASWNKVHRSSNEILSAI 303
Query: 865 TNKERDADKSNEMMPNGDLNSDGLLYTSIPLKEQNLTS 902
+NKERD +KSN+M+PN D+NSDGL++TSIPL NLTS
Sbjct: 304 SNKERDVNKSNQMIPNRDMNSDGLIHTSIPL--HNLTS 411
Score = 79.3 bits (194), Expect(2) = 7e-48
Identities = 37/48 (77%), Positives = 41/48 (85%)
Frame = +3
Query: 763 LRIGSNVVPVYLSGGERSKPHSSLKHVIVYIGFEHECPRGHRFLLNAR 810
LRIGSNVVPV+L+GGER+ HS LKH IVY+GFEHECPRGHRFL R
Sbjct: 3 LRIGSNVVPVFLNGGERNISHS-LKHAIVYLGFEHECPRGHRFLFECR 143
>TC204808 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone
glucosyltransferase (Arbutin synthase) , partial (25%)
Length = 431
Score = 34.3 bits (77), Expect = 0.27
Identities = 25/87 (28%), Positives = 35/87 (39%)
Frame = +3
Query: 41 HAMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSY 100
H +P + S + PP PS S + PTSS P + S S S
Sbjct: 66 HTPSPPSSSTSSPPTPSTSPPNSMPPPTSSTPPQPPSSPSPSTSPPSTSRSSASSETSRN 245
Query: 101 ASLFPGQCIPVTLFVFVDDFSNLSNSS 127
S PG CIP+ + F+D +N +
Sbjct: 246 RSPIPG-CIPLPVKDFLDPVLERTNEA 323
>TC219055 similar to UP|PFTB_PEA (Q04903) Protein farnesyltransferase beta
subunit (CAAX farnesyltransferase beta subunit) (RAS
proteins prenyltransferase beta) (FTase-beta) , partial
(30%)
Length = 904
Score = 33.9 bits (76), Expect = 0.36
Identities = 24/80 (30%), Positives = 35/80 (43%), Gaps = 4/80 (5%)
Frame = -2
Query: 737 KSQDILLTEGSLDGSGNNSCRDDDPFLRIGSNVVPVYLSGGERSKPHSSLKHVIVYIGFE 796
K D ++ E +DG S +DP GS V G E +K + HVI+ + +
Sbjct: 366 KKIDSVIFEFIIDGFSRQSNGVEDPVAEPGSISVQNGKCGTEMAKAFGDILHVIIALKLQ 187
Query: 797 HE----CPRGHRFLLNARHL 812
HE G L+N +HL
Sbjct: 186 HEGLGVARNGGEELVNLKHL 127
>TC209731 similar to UP|Q8W3M8 (Q8W3M8) 6b-interacting protein 1, partial
(20%)
Length = 581
Score = 32.0 bits (71), Expect = 1.4
Identities = 17/27 (62%), Positives = 19/27 (69%)
Frame = +3
Query: 49 SQTMPPLPSRSHSSLSSRPTSSANNSS 75
S T PLPS SHSS SS SSA++SS
Sbjct: 498 SSTSLPLPSSSHSSSSSSSKSSASSSS 578
>TC215547 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial (97%)
Length = 797
Score = 31.2 bits (69), Expect = 2.3
Identities = 18/48 (37%), Positives = 27/48 (55%)
Frame = +2
Query: 124 SNSSTNGEESSDVSSLSQSNSMSSAAKANLPAKVSGSVVMLARPASRS 171
S SST +S S S ++S +SA ++ P+ SGSV + P +RS
Sbjct: 320 SRSSTRTPPASSPSPSSATSSPTSARSSSPPSSTSGSVKSTSDPTARS 463
Score = 29.3 bits (64), Expect = 8.8
Identities = 15/41 (36%), Positives = 21/41 (50%)
Frame = +2
Query: 41 HAMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGG 81
++ P S PP S S SS +S PTS+ ++S P G
Sbjct: 305 NSATPSRSSTRTPPASSPSPSSATSSPTSARSSSPPSSTSG 427
>TC215548 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial (97%)
Length = 771
Score = 30.8 bits (68), Expect = 3.0
Identities = 18/48 (37%), Positives = 27/48 (55%)
Frame = +3
Query: 124 SNSSTNGEESSDVSSLSQSNSMSSAAKANLPAKVSGSVVMLARPASRS 171
S SST +S S S ++S +SA ++ P+ SGSV + P +RS
Sbjct: 333 SRSSTRTPPASSPSPSSATSSPTSARSSSPPSSTSGSVKSTSVPTARS 476
Score = 29.3 bits (64), Expect = 8.8
Identities = 15/41 (36%), Positives = 21/41 (50%)
Frame = +3
Query: 41 HAMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGG 81
++ P S PP S S SS +S PTS+ ++S P G
Sbjct: 318 NSATPSRSSTRTPPASSPSPSSATSSPTSARSSSPPSSTSG 440
>TC223353 weakly similar to UP|Q9SEW3 (Q9SEW3) Receptor-like protein kinase
(Fragment), partial (17%)
Length = 714
Score = 30.8 bits (68), Expect = 3.0
Identities = 21/64 (32%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Frame = -1
Query: 722 NGERVNAAVDYEISQKSQDILLTEGSLDGSGNNSCRDDDPFLRI-GSN--VVPVYLSGGE 778
NGE++NA VDYE S K ++ L++ + D F +I G+N + + S G
Sbjct: 198 NGEKLNAWVDYEASSKVLEVRLSKWGAQKPSDPIVSHDIDFSKIWGANPVIAGISSSNGA 19
Query: 779 RSKP 782
S P
Sbjct: 18 HSVP 7
>TC228673 similar to UP|Q8L8M8 (Q8L8M8) Serine-rich protein, partial (43%)
Length = 1083
Score = 30.8 bits (68), Expect = 3.0
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 4/55 (7%)
Frame = +1
Query: 104 FPGQCIPVTLFVFVDDFSNLSNSSTNG----EESSDVSSLSQSNSMSSAAKANLP 154
FP + ++L VF+ ++ S + ++G SS SSLS SNS S + +N P
Sbjct: 67 FPSLSLSLSLSVFLFVMASSSRTKSSGPVLRSSSSSSSSLSHSNSFCSYSTSNAP 231
>TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protein homolog,
partial (51%)
Length = 1959
Score = 30.4 bits (67), Expect = 4.0
Identities = 26/92 (28%), Positives = 39/92 (42%), Gaps = 4/92 (4%)
Frame = +2
Query: 121 SNLSNSSTNGEESSDVSSLSQSNS---MSSAAKANLPAKVSGSVVMLARPASRSEGGFRK 177
SN SN S NG+ + S M SAAK + G L++ +S G +++
Sbjct: 26 SNGSNGSPNGDGIQAIGRSSHKMIEAVMQSAAKGKVQTIRQG---YLSKRSSNLRGDWKR 196
Query: 178 KLQA-SLEAQIRFLIKKCHTLSGSEITHSGVR 208
+ + + K+C SGS HSG R
Sbjct: 197 RFFVLDSRGMLYYYRKQCSKSSGSSXQHSGXR 292
>TC207509
Length = 641
Score = 30.4 bits (67), Expect = 4.0
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = -3
Query: 44 APFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNV 84
AP +++ PP S + S S P +S +SP R GGG V
Sbjct: 399 APRRRARRTPPARSTARPSRSGCPRASKRVASPARKGGGIV 277
>TC228256 weakly similar to UP|Q9ATY4 (Q9ATY4) MAP kinase phosphatase
(Fragment), partial (10%)
Length = 1848
Score = 29.6 bits (65), Expect = 6.8
Identities = 33/105 (31%), Positives = 54/105 (51%), Gaps = 5/105 (4%)
Frame = +3
Query: 61 SSLSSRPTSSANNSSPGRGGGGNVSRNGSAMS-LMSGLGSYASLFPGQCIPVTLFVFVDD 119
SSLSS +S +++SSP ++S + S S +S + +S IPV+ + +
Sbjct: 393 SSLSSSSSSVSSSSSPSYVSPDSISSDSSPHSKSLSEVSPDSSSIVLTSIPVS--SCLSN 566
Query: 120 FSNLSNSSTNG----EESSDVSSLSQSNSMSSAAKANLPAKVSGS 160
FSNLS +S + S+DV + S+ S + A+LP K S +
Sbjct: 567 FSNLSITSYSTPKTVSNSTDVLGVKLSHPWSQS--ASLPLKKSST 695
>TC225066 homologue to UP|Q9M2F1 (Q9M2F1) Ribosomal protein S27 (Fragment),
partial (94%)
Length = 669
Score = 29.6 bits (65), Expect = 6.8
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +2
Query: 513 FFLHACACGRSRQLRPDPFDFESADASCFSDC 544
F++ C C R+ R +PF F S D + FS C
Sbjct: 74 FYVIFCCCVSPRKERSEPFVFLSDDDAIFSSC 169
>TC204215 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin response
modulator, partial (45%)
Length = 1589
Score = 29.6 bits (65), Expect = 6.8
Identities = 18/46 (39%), Positives = 27/46 (58%)
Frame = +2
Query: 124 SNSSTNGEESSDVSSLSQSNSMSSAAKANLPAKVSGSVVMLARPAS 169
SNS+ + +S +S+ + S S K +L + S S+VMLARP S
Sbjct: 1229 SNSAASFATASRIST--RKCSRSDPEKPSLSTRFSSSIVMLARPGS 1360
>TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), partial (3%)
Length = 829
Score = 29.6 bits (65), Expect = 6.8
Identities = 17/51 (33%), Positives = 25/51 (48%)
Frame = -1
Query: 25 FDTRVLRNFRVLQAAKHAMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSS 75
F+ + NF +L + + S + P PS S S S PTSS++ SS
Sbjct: 529 FENALFPNFLILCLSNLLLTFHGTSSSSKPYPSSSSPSSLSPPTSSSSTSS 377
>TC210824 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper protein-like
(AT5g61010/maf19_10), partial (13%)
Length = 1027
Score = 29.3 bits (64), Expect = 8.8
Identities = 19/57 (33%), Positives = 27/57 (47%)
Frame = -1
Query: 55 LPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYASLFPGQCIPV 111
L S +S L+ PTSS N +S S+ S S + S ++F G CIP+
Sbjct: 190 LRSLCNSPLTKEPTSSKNKTSTST----------SSFSKTSYMSSKRNIFSGVCIPI 50
>BQ079694
Length = 426
Score = 29.3 bits (64), Expect = 8.8
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = +1
Query: 44 APFAKSQTMPPLPSRSHSSLSSRPTSSANNSSP 76
AP + PP P+ S SSL P+SSAN +P
Sbjct: 82 APAPLRRASPPSPTTSPSSLPPPPSSSANAFAP 180
>TC219719 homologue to UP|Q8GZD2 (Q8GZD2) Expansin, partial (97%)
Length = 1007
Score = 29.3 bits (64), Expect = 8.8
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +2
Query: 47 AKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSG 96
A T+P S S SSR S+A SSP G + R G++ ++G
Sbjct: 407 ADGATLPARTSTSPCPCSSRSPSTAPESSPSLTAGCHAERQGASRFTING 556
>TC208501 similar to GB|AAN31116.1|23506209|AY149962 At3g06130/F28L1_7
{Arabidopsis thaliana;} , partial (26%)
Length = 1081
Score = 29.3 bits (64), Expect = 8.8
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = -1
Query: 59 SHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYASL 103
S S+ SS +S+++NSS + G ++ GLGS+ SL
Sbjct: 187 SSSNSSSNSSSNSSNSSSDKSSSGKLNLTAFGFGFGFGLGSFMSL 53
>TC207954 similar to UP|Q9M9V0 (Q9M9V0) F6A14.10 protein, partial (33%)
Length = 742
Score = 29.3 bits (64), Expect = 8.8
Identities = 21/68 (30%), Positives = 32/68 (46%), Gaps = 12/68 (17%)
Frame = -1
Query: 57 SRSHSSLSSRPTSSANNSSPGRGGGGNVSR------------NGSAMSLMSGLGSYASLF 104
S S S SS P+SSA++SSP G ++ + N + S MS + S+ S+
Sbjct: 355 SSSSSDSSSFPSSSASSSSPSSSSGSSLLK*VSGFGHKSSLINSATSS*MSSISSFCSVS 176
Query: 105 PGQCIPVT 112
Q +T
Sbjct: 175 LNQLKKLT 152
>TC206712 similar to GB|AAT41855.1|48310625|BT014872 At2g19340 {Arabidopsis
thaliana;} , partial (74%)
Length = 741
Score = 29.3 bits (64), Expect = 8.8
Identities = 30/113 (26%), Positives = 44/113 (38%)
Frame = +2
Query: 44 APFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYASL 103
AP A + + PP PS + S +S + SSP + S SA +
Sbjct: 191 APLASASSFPPSPSPPRTPSSPSSSSPTSWSSPALFTTSSSSPPASA--------PLRTP 346
Query: 104 FPGQCIPVTLFVFVDDFSNLSNSSTNGEESSDVSSLSQSNSMSSAAKANLPAK 156
P P + S S +S SS V+S S S++ S A A P++
Sbjct: 347 TPAPSAPSSSCPAASTASTSSRASPPDSCSSSVASASFSSTSPSIATARNPSR 505
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,066,158
Number of Sequences: 63676
Number of extensions: 639798
Number of successful extensions: 3455
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 3321
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3425
length of query: 975
length of database: 12,639,632
effective HSP length: 106
effective length of query: 869
effective length of database: 5,889,976
effective search space: 5118389144
effective search space used: 5118389144
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0369c.3


