

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.1
(573 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
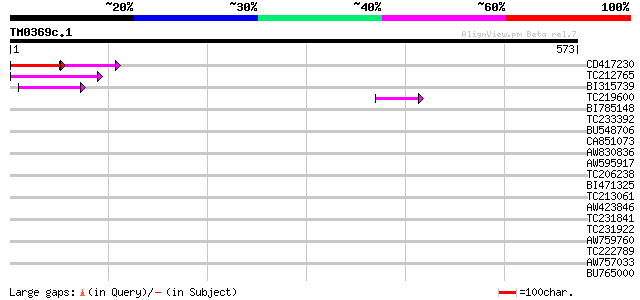
Sequences producing significant alignments: (bits) Value
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 43 1e-07
TC212765 47 3e-05
BI315739 45 7e-05
TC219600 weakly similar to UP|Q9SGD7 (Q9SGD7) T23G18.9, partial ... 42 6e-04
BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevi... 37 0.024
TC233392 35 0.091
BU548706 35 0.091
CA851073 34 0.16
AW830836 33 0.27
AW595917 32 0.59
TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragmen... 32 0.59
BI471325 32 0.77
TC213061 32 0.77
AW423846 31 1.3
TC231841 31 1.3
TC231922 30 2.2
AW759760 30 2.9
TC222789 30 2.9
AW757033 30 3.8
BU765000 30 3.8
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 43.1 bits (100), Expect(2) = 1e-07
Identities = 22/56 (39%), Positives = 36/56 (64%)
Frame = -3
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMF 56
I + + G+ ISHL VD ++LF + Q+ +I TL+ F ++SG KVN+ K++ F
Sbjct: 494 IQVNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTKSF 327
Score = 30.8 bits (68), Expect(2) = 1e-07
Identities = 19/62 (30%), Positives = 28/62 (44%)
Frame = -2
Query: 51 QKSRMFCSNVVDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
Q + S V+ ++++S +G+ LGKYL +P T F IIDK N
Sbjct: 342 QNQKFSFSKNVNWWVEEDISSGLGLQSTDDLGKYLGVPIFHKNPTSHTFDFIIDKVNKRF 163
Query: 111 GR 112
R
Sbjct: 162 SR 157
>TC212765
Length = 637
Score = 46.6 bits (109), Expect = 3e-05
Identities = 30/93 (32%), Positives = 46/93 (49%)
Frame = +3
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I ++ + + +L VD + LF L + TL FF+ G+K+NL KS M+CS
Sbjct: 93 ILVSDDNRLVPYLMLVDDIFLFINAIGD*AYLASLTL*NFFQTFGLKLNLDKSSMYCS*R 272
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKAR 93
+ + +S + I A LGKYL L + R
Sbjct: 273 MAPVLRDLISNTLNIVHAPSLGKYLCLKFIHGR 371
>BI315739
Length = 442
Score = 45.4 bits (106), Expect = 7e-05
Identities = 26/67 (38%), Positives = 36/67 (52%)
Frame = +2
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
ISHLF D LLF ++ Q++L+ L++F G+K LQKSR V RK
Sbjct: 164 ISHLFFPDNCLLFVKATSSQVKLVRDVLQQFCLT*GLKAGLQKSRFLA*KNVSNRKIARF 343
Query: 70 SIIVGIT 76
S I+ I+
Sbjct: 344 SFIIVIS 364
>TC219600 weakly similar to UP|Q9SGD7 (Q9SGD7) T23G18.9, partial (21%)
Length = 785
Score = 42.4 bits (98), Expect = 6e-04
Identities = 18/49 (36%), Positives = 23/49 (46%)
Frame = +3
Query: 370 EKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
EK+ F W + L TN +R+H L D C C + E H LR C
Sbjct: 24 EKIRLFYWTVAKGGLKTNNRRHHYSLA*DDMCIMCSGAAETYLHLLRAC 170
>BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevipalpis},
partial (2%)
Length = 422
Score = 37.0 bits (84), Expect = 0.024
Identities = 22/68 (32%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Frame = -2
Query: 370 EKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS--VWVRSR 427
+K+ WL ++L N R+H +LV A C ED H LR P + +W+
Sbjct: 415 KKIRIMFWLILHNSLSKNSTRFHRNLVNSSAFPHCSNPREDSLHCLRDFPHAKEIWLGFG 236
Query: 428 FAKALPGL 435
FA L L
Sbjct: 235 FAPHLSSL 212
>TC233392
Length = 1145
Score = 35.0 bits (79), Expect = 0.091
Identities = 22/73 (30%), Positives = 26/73 (35%), Gaps = 9/73 (12%)
Frame = -1
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDA-CFRCGASVEDVEHALRRC--- 418
+WK K F W RD LPT + +V D+ C C ED H C
Sbjct: 545 LWKLKIPAKASIFAWRLIRDRLPTKSNLHRRQVVLEDSLCPFCRIREEDASHIFLECNKI 366
Query: 419 -----PGSVWVRS 426
WVRS
Sbjct: 365 RPLWWESQTWVRS 327
>BU548706
Length = 647
Score = 35.0 bits (79), Expect = 0.091
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = +2
Query: 359 VMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYH 392
+ K IW ++V FLW + DALPTN +H
Sbjct: 413 LFKSIWDRPGPKRVKLFLWKSAHDALPTNYLXHH 514
>CA851073
Length = 633
Score = 34.3 bits (77), Expect = 0.16
Identities = 26/104 (25%), Positives = 37/104 (35%), Gaps = 19/104 (18%)
Frame = +1
Query: 359 VMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHL-VKFDACFRCGASVEDVEHALRR 417
+ K IWK + F W +D LPT ++ ++ + C CG E+ +H
Sbjct: 127 IFKTIWKLKIPPRAAVFSWRLIKDRLPTRHNLLRRNVSIQENECPLCGYEQEEADHLFFN 306
Query: 418 C------------------PGSVWVRSRFAKALPGLSAGALMTR 443
C P SV S F + G AG TR
Sbjct: 307 CKMTRGLWWESMRWNQIVGPLSVSPASHFVQFCDGFGAGRNHTR 438
>AW830836
Length = 294
Score = 33.5 bits (75), Expect = 0.27
Identities = 20/73 (27%), Positives = 31/73 (42%), Gaps = 5/73 (6%)
Frame = -2
Query: 356 KFAVMKLIWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFD-ACFRCGASVEDVE 412
K M +W+ V F+W RD + T +K+ L D +C C E ++
Sbjct: 227 KLPSMSRLWEIPITSNVSIFVWKVIRDKIQTRGKLKKRQIKLQAEDYSCALCHGEEETIQ 48
Query: 413 HALRRCP--GSVW 423
H L CP ++W
Sbjct: 47 HLLFTCPFASTIW 9
>AW595917
Length = 404
Score = 32.3 bits (72), Expect = 0.59
Identities = 16/56 (28%), Positives = 19/56 (33%)
Frame = +1
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
+WK K F W +D LPT + C C S ED H C
Sbjct: 7 LWKLKIPPKAAVFTWRLIKDQLPTKANLRRQMEISDSLCPLCNNSKEDATHLFFNC 174
>TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragment), partial
(9%)
Length = 1259
Score = 32.3 bits (72), Expect = 0.59
Identities = 19/50 (38%), Positives = 24/50 (48%)
Frame = +3
Query: 370 EKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
E+++ L A AL TN R + H +C RC E HALR CP
Sbjct: 132 ERIINLLGNAADGALFTNPLRRYRHTSMDSSCPRCPELEETCLHALRDCP 281
>BI471325
Length = 421
Score = 32.0 bits (71), Expect = 0.77
Identities = 19/67 (28%), Positives = 33/67 (48%)
Frame = +2
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
+SHL VD LF + + ++ L F +SG+ VNL+KS + S Q + +
Sbjct: 44 LSHLLSVDFYFLFCRTIDKEADVLKNILTTFEASSGLVVNLRKSEICFSRNTSQIARDSI 223
Query: 70 SIIVGIT 76
+G++
Sbjct: 224 FFHLGMS 244
>TC213061
Length = 823
Score = 32.0 bits (71), Expect = 0.77
Identities = 19/60 (31%), Positives = 26/60 (42%), Gaps = 2/60 (3%)
Frame = +2
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
K +WK KV F W +D LPT N++R L ++ C C + E H C
Sbjct: 233 KELWKLKVPMKVTMFAWRLLKDILPTRDNLRRKRVELHEY-VCPLCRSMDESASHLFFHC 409
>AW423846
Length = 385
Score = 31.2 bits (69), Expect = 1.3
Identities = 17/49 (34%), Positives = 20/49 (40%), Gaps = 1/49 (2%)
Frame = +3
Query: 371 KVMFFLWLAGRDALPTNMKRYHSHLVKFD-ACFRCGASVEDVEHALRRC 418
K FF+W RD LPT + H+ D C C ED H C
Sbjct: 21 KAAFFVWRLIRDRLPTKTNLHRRHVEIHDLTCPFCKNKEEDAAHLFFTC 167
>TC231841
Length = 791
Score = 31.2 bits (69), Expect = 1.3
Identities = 22/72 (30%), Positives = 29/72 (39%), Gaps = 4/72 (5%)
Frame = +3
Query: 359 VMKLIWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVEHALR 416
V + +WK K+ F W RD LPT N++R V C C ++ E H
Sbjct: 162 VFEELWKLKLPSKITIFAWRLIRDRLPTRSNLRRKQIE-VDDPRCPFCRSAEESAAHLFF 338
Query: 417 RCP--GSVWVRS 426
C VW S
Sbjct: 339 HCSRIAPVWWES 374
>TC231922
Length = 803
Score = 30.4 bits (67), Expect = 2.2
Identities = 22/88 (25%), Positives = 34/88 (38%), Gaps = 5/88 (5%)
Frame = -3
Query: 355 PKFAVMKLIWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVE 412
P + +++W + F W D LPT N+ R S ++ C CG E+
Sbjct: 357 PSRRIFQILWHLKIPPRAAVFSWRLFLDRLPTRGNLSR-RSIPIQDIMCPLCGCQHEEAG 181
Query: 413 HALRRC---PGSVWVRSRFAKALPGLSA 437
H C G W R+ + + L A
Sbjct: 180 HLFFHCKMTKGLWWESMRWIRVIGALPA 97
>AW759760
Length = 404
Score = 30.0 bits (66), Expect = 2.9
Identities = 24/84 (28%), Positives = 45/84 (53%), Gaps = 2/84 (2%)
Frame = +1
Query: 10 ISHL-FCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRM-FCSNVVDQRKQQ 67
++HL +C ++L F V *Q L+ F + N K R+ FC+N+ +++++Q
Sbjct: 133 LTHLIWCT*MIIL*FGVVA*Q*ALLFSYFHYFLDKFLKTRNKVKMRLRFCNNL*NRKQKQ 312
Query: 68 ELSIIVGITRASYLGKYLDLPQLK 91
++ + T Y+G +L LP L+
Sbjct: 313 KIWCKL-FTLVGYVGNFLRLPSLR 381
>TC222789
Length = 954
Score = 30.0 bits (66), Expect = 2.9
Identities = 17/57 (29%), Positives = 22/57 (37%), Gaps = 1/57 (1%)
Frame = +2
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFD-ACFRCGASVEDVEHALRRC 418
+WK K F W RD LPT + + + D +C C ED H C
Sbjct: 362 LWKLKVPIKYEVFAWRLLRDRLPTKVNLHRRQIQVMDRSCPFCRNVEEDAGHLFFHC 532
>AW757033
Length = 441
Score = 29.6 bits (65), Expect = 3.8
Identities = 26/94 (27%), Positives = 44/94 (46%)
Frame = -2
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
+S L D + F + +++I L+ F ASG+K+N KSR +V D + +E
Sbjct: 284 VSILQYADDTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WR-KEA 108
Query: 70 SIIVGITRASYLGKYLDLPQLKARVTKEHFAPII 103
+ + + S YL +P +E + PII
Sbjct: 107 AEFMNCSLLSLPFSYLGIPIGANPRRRETWDPII 6
>BU765000
Length = 425
Score = 29.6 bits (65), Expect = 3.8
Identities = 19/60 (31%), Positives = 25/60 (41%), Gaps = 2/60 (3%)
Frame = +2
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
K +WK KV F W +D LPT N++R L ++ C C E H C
Sbjct: 53 KELWKLKVPPKVAVFAWRLLKDRLPTRDNLRRKQVELHEY-LCPFCRTMEESACHLFFHC 229
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.339 0.147 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,175,302
Number of Sequences: 63676
Number of extensions: 479043
Number of successful extensions: 3632
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 3603
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3631
length of query: 573
length of database: 12,639,632
effective HSP length: 102
effective length of query: 471
effective length of database: 6,144,680
effective search space: 2894144280
effective search space used: 2894144280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0369c.1


