

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369b.1
(266 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
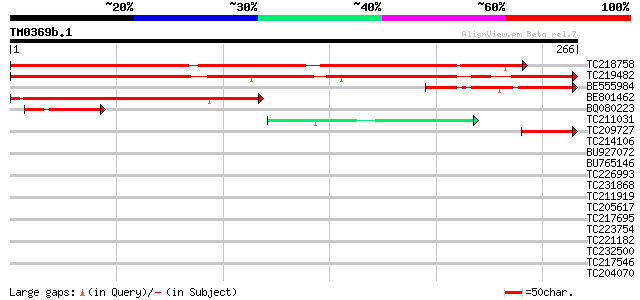
Score E
Sequences producing significant alignments: (bits) Value
TC218758 similar to UP|Q9FFB5 (Q9FFB5) Emb|CAB89309.1, partial (... 363 e-101
TC219482 similar to UP|Q9FFB5 (Q9FFB5) Emb|CAB89309.1, partial (... 354 3e-98
BE555984 104 5e-23
BE801462 94 8e-20
BQ080223 50 1e-06
TC211031 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 44 1e-04
TC209727 44 1e-04
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 40 0.001
BU927072 40 0.001
BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvo... 40 0.001
TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 deh... 40 0.001
TC231868 similar to UP|Q93424 (Q93424) C. elegans GRL-23 protein... 38 0.004
TC211919 weakly similar to UP|K2C1_HUMAN (P04264) Keratin, type ... 38 0.005
TC205617 38 0.005
TC217695 weakly similar to UP|Q9LUI1 (Q9LUI1) Extensin protein-l... 37 0.007
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 37 0.009
TC221182 weakly similar to UP|ENAH_MOUSE (Q03173) Enabled protei... 37 0.009
TC232500 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 37 0.012
TC217546 weakly similar to UP|Q9GZC7 (Q9GZC7) RNA binding protei... 37 0.012
TC204070 similar to UP|Q39835 (Q39835) Extensin, partial (34%) 37 0.012
>TC218758 similar to UP|Q9FFB5 (Q9FFB5) Emb|CAB89309.1, partial (59%)
Length = 831
Score = 363 bits (931), Expect = e-101
Identities = 189/244 (77%), Positives = 209/244 (85%), Gaps = 1/244 (0%)
Frame = +1
Query: 1 MDYQRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLN 60
MD QR+ LL+ AY+FQGKSMEE+RQSL+YTTLELEQTR AVQEEL+KRD+QLLNLKDLL+
Sbjct: 133 MDNQRSPLLSWAYYFQGKSMEELRQSLIYTTLELEQTRVAVQEELKKRDEQLLNLKDLLS 312
Query: 61 NTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSS 120
TIRERDEAQEKCQRLLLEK +FQ QQ Q A P SG+SSIEDEPRRGIDSNNG S S
Sbjct: 313 KTIRERDEAQEKCQRLLLEKLVFQ----QQLQHAAPVSGISSIEDEPRRGIDSNNGHSLS 480
Query: 121 DCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTL 180
DCEESIVSSPV D++PQPQ QSMIELTP+KPLPEKGKLLQAVMKAGPLLQTL
Sbjct: 481 DCEESIVSSPVIDHLPQPQ------PQTQSMIELTPDKPLPEKGKLLQAVMKAGPLLQTL 642
Query: 181 LLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSS-STSTHSQC 239
LLAGPLPQWRHPPPPLESFEIPPVTIPSPP PPQ PQL HQ++FFN ++ S+ST + C
Sbjct: 643 LLAGPLPQWRHPPPPLESFEIPPVTIPSPP-PPPQPQPQLFHQDTFFNNTNGSSSTTTNC 819
Query: 240 GRVS 243
GR+S
Sbjct: 820 GRLS 831
>TC219482 similar to UP|Q9FFB5 (Q9FFB5) Emb|CAB89309.1, partial (49%)
Length = 1096
Score = 354 bits (908), Expect = 3e-98
Identities = 190/269 (70%), Positives = 214/269 (78%), Gaps = 3/269 (1%)
Frame = +2
Query: 1 MDYQRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLN 60
MD QR+ LL+ A++ GKSMEE++QSL+YTTLELEQTR VQEELRKRDDQLL LK+LLN
Sbjct: 107 MDNQRSPLLSWAFYCHGKSMEELKQSLMYTTLELEQTRATVQEELRKRDDQLLTLKELLN 286
Query: 61 NTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGID--SNNGLS 118
IRERDEAQEKCQRL+LEK +FQH Q P SGVSSIEDEPRRGI+ SNNGL
Sbjct: 287 KVIRERDEAQEKCQRLVLEKMVFQH-------QTAPASGVSSIEDEPRRGINDSSNNGLX 445
Query: 119 SSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIEL-TPEKPLPEKGKLLQAVMKAGPLL 177
SSDCEESIVSSPV +++P Q LP +SMIEL +P+KPLPEKGKLLQAVMKAGPLL
Sbjct: 446 SSDCEESIVSSPVMEHLPPQQQQLP-----ESMIELISPDKPLPEKGKLLQAVMKAGPLL 610
Query: 178 QTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHS 237
QTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPP PQL HQ+SF S
Sbjct: 611 QTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPP------QPQLPHQDSF---------TS 745
Query: 238 QCGRVSRKRIFCDGSDSPSETKFQRVVLH 266
CGRVSRKR+FC+G+DSP++ KFQR+VLH
Sbjct: 746 NCGRVSRKRVFCEGTDSPTQNKFQRIVLH 832
>BE555984
Length = 289
Score = 104 bits (259), Expect = 5e-23
Identities = 55/74 (74%), Positives = 64/74 (86%), Gaps = 3/74 (4%)
Frame = +3
Query: 196 LESFEIPPVTIPSPPLQPPQQPPQLLHQESFFN---GSSSTSTHSQCGRVSRKRIFCDGS 252
LESFEIPPVTIPSPP PP P QLLHQ++FFN GSSST+T+ CGR+SRKR+FCDGS
Sbjct: 6 LESFEIPPVTIPSPP--PP--PLQLLHQDTFFNKTNGSSSTTTN--CGRLSRKRVFCDGS 167
Query: 253 DSPSETKFQRVVLH 266
DSP+ETK+QR+VLH
Sbjct: 168 DSPTETKYQRLVLH 209
>BE801462
Length = 424
Score = 93.6 bits (231), Expect = 8e-20
Identities = 49/122 (40%), Positives = 78/122 (63%), Gaps = 3/122 (2%)
Frame = +1
Query: 1 MDYQRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLN 60
M+Y+R S L ++ Q + +E++R SL+YTTLEL+ T + +EE+ +R+ +L+++ DLL+
Sbjct: 58 MEYER-SPLGWEFYHQEEGLEDLRHSLLYTTLELDATIASAKEEITRRECELIHVNDLLS 234
Query: 61 NTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQ---QADPNSGVSSIEDEPRRGIDSNNGL 117
I+ERDEAQ KCQ+L+LEK Q +QQ +Q Q +S E+E + G +
Sbjct: 235 RVIKERDEAQAKCQKLMLEKLELQQKQQLEQHQFVQTHQRDTISQSEEEQQEGFSEKHSA 414
Query: 118 SS 119
SS
Sbjct: 415 SS 420
>BQ080223
Length = 426
Score = 49.7 bits (117), Expect = 1e-06
Identities = 28/38 (73%), Positives = 31/38 (80%)
Frame = +3
Query: 8 LLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEEL 45
LLN + Q SMEE+RQSL+YTTLELEQTR AVQEEL
Sbjct: 318 LLNSINWVQ--SMEELRQSLIYTTLELEQTRDAVQEEL 425
Score = 28.5 bits (62), Expect = 3.3
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +2
Query: 1 MDYQRTSLLNCAYFFQGK 18
MD QR+ LL+ AY+FQGK
Sbjct: 104 MDNQRSPLLSWAYYFQGK 157
>TC211031 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (11%)
Length = 678
Score = 43.5 bits (101), Expect = 1e-04
Identities = 33/103 (32%), Positives = 39/103 (37%), Gaps = 4/103 (3%)
Frame = +3
Query: 122 CEESIVSSPVFDNMPQPQFPL----PLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLL 177
C+E + P P P PL PLP + +P PLP PL
Sbjct: 132 CKECELPPPPPKECPPPPIPLAPPPPLPPSPPPPLPPSPPPPLPPS--------PPPPLP 287
Query: 178 QTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQL 220
+ L+ P P PPPPL PP PPL PP P L
Sbjct: 288 PPIPLSTPPPLPPSPPPPLPPSPSPPNLPSPPPLAPPPPLPPL 416
>TC209727
Length = 752
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/26 (73%), Positives = 23/26 (88%)
Frame = +1
Query: 241 RVSRKRIFCDGSDSPSETKFQRVVLH 266
R+SRKR+FCDGSD +ETK QR+VLH
Sbjct: 22 RLSRKRVFCDGSDFITETKSQRLVLH 99
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 40.0 bits (92), Expect = 0.001
Identities = 31/95 (32%), Positives = 34/95 (35%), Gaps = 6/95 (6%)
Frame = +1
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV+ P P P P P P P V P +Q + P PQ
Sbjct: 43 PVYSPPPPPPSPPPPSPTYCVRSPPPPSPPPPSPPPPPPPVFSPPPPVQYYYSSPPPPQH 222
Query: 190 RHPPPPLESFEIPPVTIPSPP------LQPPQQPP 218
PPPP S PP PSPP L PP PP
Sbjct: 223 SPPPPPPHS---PPPPPPSPPPPVYPYLSPPPPPP 318
Score = 33.9 bits (76), Expect = 0.078
Identities = 29/91 (31%), Positives = 35/91 (37%), Gaps = 2/91 (2%)
Frame = +1
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PVF P Q+ P Q +P P P P + L P P
Sbjct: 160 PVFSPPPPVQYYYSSPPPPQH----SPPPPPPHSPP--PPPPSPPPPVYPYLSPPPPPPV 321
Query: 190 RHPPPPLESFEIPPVTIPSPP--LQPPQQPP 218
PPPP+ S PP PSPP ++PP PP
Sbjct: 322 HSPPPPVYS---PPPPPPSPPPCIEPPPPPP 405
Score = 31.2 bits (69), Expect = 0.50
Identities = 30/103 (29%), Positives = 33/103 (31%), Gaps = 7/103 (6%)
Frame = +1
Query: 130 PVFDNMPQPQFPLPL---PSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
PV+ P P P P P IE P P P + P
Sbjct: 337 PVYSPPPPPPSPPPCIEPPPPPPPCIEPPPPPPSPPPCEE----------------HSPP 468
Query: 187 PQWRHP----PPPLESFEIPPVTIPSPPLQPPQQPPQLLHQES 225
P HP PPP S PPV SPP P P + H S
Sbjct: 469 PPSPHPAPYHPPPSPSPPPPPVQYNSPPPPSPPPPTPVYHYNS 597
Score = 31.2 bits (69), Expect = 0.50
Identities = 13/27 (48%), Positives = 14/27 (51%)
Frame = +1
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPPP PP + SPP PP PP
Sbjct: 4 PPPPSPPPPSPPPPVYSPPPPPPSPPP 84
Score = 29.3 bits (64), Expect = 1.9
Identities = 17/45 (37%), Positives = 18/45 (39%), Gaps = 11/45 (24%)
Frame = +1
Query: 185 PLPQWRHPPPPLESFEIPP-----------VTIPSPPLQPPQQPP 218
P P PPPP+ S PP V P PP PP PP
Sbjct: 13 PSPPPPSPPPPVYSPPPPPPSPPPPSPTYCVRSPPPPSPPPPSPP 147
>BU927072
Length = 314
Score = 40.0 bits (92), Expect = 0.001
Identities = 29/73 (39%), Positives = 32/73 (43%), Gaps = 8/73 (10%)
Frame = +3
Query: 156 PEKPLPEKGKLLQAVMKAGPLLQTLL-------LAGPLPQ-WRHPPPPLESFEIPPVTIP 207
P P P +L + PLLQ LL L PLP +R P PL PP P
Sbjct: 66 PPPPPPNPLRLRHRLPPPKPLLQLLLPLHHPPPLLHPLPPPFRKPLRPLLRRPPPPPPAP 245
Query: 208 SPPLQPPQQPPQL 220
PPL PP PP L
Sbjct: 246 PPPLPPPTPPPVL 284
Score = 35.0 bits (79), Expect = 0.035
Identities = 30/83 (36%), Positives = 34/83 (40%), Gaps = 10/83 (12%)
Frame = +3
Query: 141 PLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTL----------LLAGPLPQWR 190
P P P+ + L P KPL + LL + PLL L LL P P
Sbjct: 69 PPPPPNPLRLRHRLPPPKPLLQ---LLLPLHHPPPLLHPLPPPFRKPLRPLLRRPPPPPP 239
Query: 191 HPPPPLESFEIPPVTIPSPPLQP 213
PPPPL PPV P P L P
Sbjct: 240 APPPPLPPPTPPPVLPPPPXLPP 308
>BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (25%)
Length = 407
Score = 39.7 bits (91), Expect = 0.001
Identities = 29/83 (34%), Positives = 30/83 (35%)
Frame = -2
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P P P P PS L P+KP P P PLP PPPP
Sbjct: 283 PPPPPPPPPPSPPPPXXSLPPKKPPPPPPPPPPPPPPPPP--------PPLPPPPPPPPP 128
Query: 196 LESFEIPPVTIPSPPLQPPQQPP 218
PP PSPP PP PP
Sbjct: 127 PPPPPPPPPPPPSPPPPPPPPPP 59
Score = 35.8 bits (81), Expect = 0.020
Identities = 28/85 (32%), Positives = 29/85 (33%)
Frame = -3
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P P P P P P PLP L + P P P PPPP
Sbjct: 324 PSPPPPPPPPPPPPPPPPPPPPPPLPPLPXSLSPLKNPPP-------PPPPPPPPPPPPP 166
Query: 196 LESFEIPPVTIPSPPLQPPQQPPQL 220
F PP P PP PP PP L
Sbjct: 165 PPPFPPPP---PPPPPPPPPPPPPL 100
Score = 34.7 bits (78), Expect = 0.046
Identities = 25/83 (30%), Positives = 30/83 (36%)
Frame = -1
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P P P P P ++ + L+P P P P P PPPP
Sbjct: 287 PPPPPPPPPPPLSPPSLXLSPP*KTPPPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPP 108
Query: 196 LESFEIPPVTIPSPPLQPPQQPP 218
S +PP P PP PP PP
Sbjct: 107 PPSPPLPPPPPPPPPPPPPPPPP 39
Score = 34.3 bits (77), Expect = 0.060
Identities = 27/83 (32%), Positives = 29/83 (34%)
Frame = -3
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P P P PLP + S+ L P P P P P PPPP
Sbjct: 276 PPPPPPPPLPPLPXSLSPLKNPPPPPPPPPPPPPPPPPPPFPPPPPPPPPPPP--PPPPP 103
Query: 196 LESFEIPPVTIPSPPLQPPQQPP 218
L PP P PP PP PP
Sbjct: 102 LPPPPPPPPPPPPPPPPPPPPPP 34
Score = 33.9 bits (76), Expect = 0.078
Identities = 18/41 (43%), Positives = 18/41 (43%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQES 225
P P PPPPL PP P PP PP PP L S
Sbjct: 362 PPPPPPPPPPPLPPPPPPPPXPPPPPPPPPPPPPPPLSPPS 240
Score = 33.5 bits (75), Expect = 0.10
Identities = 17/37 (45%), Positives = 17/37 (45%)
Frame = -3
Query: 182 LAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
L PLP PPPP PP P PP PP PP
Sbjct: 387 LLPPLPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPP 277
Score = 33.5 bits (75), Expect = 0.10
Identities = 14/27 (51%), Positives = 14/27 (51%)
Frame = -1
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPPP S PP P PP PP PP
Sbjct: 401 PPPPPSSLPSPPPPPPPPPPPPPPLPP 321
Score = 33.5 bits (75), Expect = 0.10
Identities = 15/34 (44%), Positives = 16/34 (46%)
Frame = -2
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPP +PP P PP PP PP
Sbjct: 370 PPPPPPPPPPPPPPSPLPPPPPPXPPPPPPPPPP 269
Score = 33.5 bits (75), Expect = 0.10
Identities = 16/37 (43%), Positives = 16/37 (43%)
Frame = -1
Query: 182 LAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
L P P PPPP PP P PP PP PP
Sbjct: 380 LPSPPPPPPPPPPPPPPLPPPPPPPPXPPPPPPPPPP 270
Score = 32.7 bits (73), Expect = 0.17
Identities = 19/43 (44%), Positives = 19/43 (44%)
Frame = -3
Query: 176 LLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
LL L P P PPPP S PP P PP PP PP
Sbjct: 387 LLPPLPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPPP 259
Score = 32.3 bits (72), Expect = 0.23
Identities = 15/34 (44%), Positives = 15/34 (44%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPP PP P PP PP PP
Sbjct: 134 PPPPPPPPPPPSPPLPPPPPPPPPPPPPPPPPPP 33
Score = 32.0 bits (71), Expect = 0.30
Identities = 16/35 (45%), Positives = 17/35 (47%)
Frame = -3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQ 219
PLP PPPP PP P PP PP PP+
Sbjct: 105 PLPPPPPPPPP------PPPPPPPPPPPPPPPPPR 19
Score = 31.2 bits (69), Expect = 0.50
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = -3
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPPP S +PP+ P PP PP PP
Sbjct: 405 PPPP--SSLLPPLPPPPPPPPPPPPPP 331
Score = 30.8 bits (68), Expect = 0.66
Identities = 15/34 (44%), Positives = 15/34 (44%)
Frame = -2
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPP PP P PP PP PP
Sbjct: 124 PPPPPPPPPPPSPPPPPPPPPPPPPPPPPPPPPP 23
Score = 30.0 bits (66), Expect = 1.1
Identities = 13/27 (48%), Positives = 13/27 (48%)
Frame = -1
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPPP PP P PP PP PP
Sbjct: 398 PPPPSSLPSPPPPPPPPPPPPPPLPPP 318
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/38 (42%), Positives = 16/38 (42%)
Frame = -2
Query: 181 LLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
LL P P PPPP P P PP PP PP
Sbjct: 394 LLPPPSPPPPPPPPPPPPPPPPSPLPPPPPPXPPPPPP 281
Score = 28.5 bits (62), Expect = 3.3
Identities = 14/33 (42%), Positives = 15/33 (45%)
Frame = -2
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
P P PPPP PP P PP PP +P
Sbjct: 109 PPPPPPSPPPPPPPPPPPPPPPPPPPPPPPGKP 11
Score = 27.7 bits (60), Expect = 5.6
Identities = 15/37 (40%), Positives = 15/37 (40%), Gaps = 3/37 (8%)
Frame = -2
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQ---PPQQPP 218
P P PPPP PP P PP PP PP
Sbjct: 364 PPPPPPPPPPPPSPLPPPPPPXPPPPPPPPPPPPPPP 254
Score = 27.7 bits (60), Expect = 5.6
Identities = 15/35 (42%), Positives = 15/35 (42%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQ 219
P P PPPP PP P PP PP P Q
Sbjct: 104 PSPPLPPPPPPP-----PPPPPPPPPPPPPPPPGQ 15
Score = 27.3 bits (59), Expect = 7.3
Identities = 13/28 (46%), Positives = 13/28 (46%)
Frame = -2
Query: 193 PPPLESFEIPPVTIPSPPLQPPQQPPQL 220
PPPL PP P PP PP P L
Sbjct: 403 PPPLLPPPSPPPPPPPPPPPPPPPPSPL 320
>TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 dehydratase
(GDP-D-mannose dehydratase) (GMD) , partial (94%)
Length = 1388
Score = 39.7 bits (91), Expect = 0.001
Identities = 34/113 (30%), Positives = 40/113 (35%), Gaps = 10/113 (8%)
Frame = +2
Query: 141 PLPLPSVAQSMIELTP---EKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLE 197
PL L S + S TP P P P T P P R PPL
Sbjct: 395 PLSLTSPSPSRYPTTPPTSSPPAPSASSRPSVPTSPPPAAPTSATTKPAPP-RCSAPPLR 571
Query: 198 SFEIPPVTIPSPPLQPPQQPP-------QLLHQESFFNGSSSTSTHSQCGRVS 243
PP + P+PP PP PP L S SSST++ R+S
Sbjct: 572 RSPKPPPSTPAPPTPPPNAPPTGTP*TTARLTPSSPATASSSTTSPPAAARIS 730
>TC231868 similar to UP|Q93424 (Q93424) C. elegans GRL-23 protein
(Corresponding sequence E02A10.2), partial (23%)
Length = 790
Score = 38.1 bits (87), Expect = 0.004
Identities = 31/95 (32%), Positives = 37/95 (38%), Gaps = 11/95 (11%)
Frame = -3
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHP-PP 194
PQP P P ++ P+ P P Q + PLL P P + P PP
Sbjct: 458 PQPPPQPPQPLLSNLFPHPPPQPPQPPPQPPPQPAPQP-PLLIKFPQPPPQPPPQPPQPP 282
Query: 195 PLESFEIPP----------VTIPSPPLQPPQQPPQ 219
PL F PP + P PP QP QPPQ
Sbjct: 281 PLIKFPQPPPQPEPQPPLLIIFPQPPPQPAPQPPQ 177
Score = 30.4 bits (67), Expect = 0.86
Identities = 30/101 (29%), Positives = 39/101 (37%), Gaps = 13/101 (12%)
Frame = -3
Query: 130 PVFDNM---PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVM----------KAGPL 176
P+ N+ P PQ P P P P +P P+ L++ + PL
Sbjct: 431 PLLSNLFPHPPPQPPQPPPQ--------PPPQPAPQPPLLIKFPQPPPQPPPQPPQPPPL 276
Query: 177 LQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
++ P PQ PP L F PP P P QPPQ P
Sbjct: 275 IK---FPQPPPQPEPQPPLLIIFPQPP---PQPAPQPPQPP 171
>TC211919 weakly similar to UP|K2C1_HUMAN (P04264) Keratin, type II
cytoskeletal 1 (Cytokeratin 1) (K1) (CK 1) (67 kDa
cytokeratin) (Hair alpha protein), partial (5%)
Length = 590
Score = 37.7 bits (86), Expect = 0.005
Identities = 28/99 (28%), Positives = 38/99 (38%)
Frame = +3
Query: 123 EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLL 182
+ES + SP P P F P P TP+ P L + P L
Sbjct: 201 DESCIPSPT--TFPNPNFNTPTPPSPIDNNPTTPQTPQ------LPNPIPPPPYSNNPSL 356
Query: 183 AGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLL 221
P P + PPPP E+ P ++ +PP+ P LL
Sbjct: 357 PPPPPILQSPPPPEETTPPTPPSVAAPPMNTPTPGSALL 473
>TC205617
Length = 919
Score = 37.7 bits (86), Expect = 0.005
Identities = 32/87 (36%), Positives = 39/87 (44%), Gaps = 6/87 (6%)
Frame = +1
Query: 159 PLPEKGKLL-QAVMKAGPLLQTLLLAGPLPQWRHPP---PPLESFEIP--PVTIPSPPLQ 212
PL KLL QA L+ LLL +PQW PP PL S P +TI P
Sbjct: 625 PLKHPPKLLIQASYPRLILVVKLLLPWLIPQWIAPPLTSSPLSSATSPRWKITIKGPH*S 804
Query: 213 PPQQPPQLLHQESFFNGSSSTSTHSQC 239
QPP LH+++ ST S+C
Sbjct: 805 SKIQPP*KLHKQASIKWYQSTRASSRC 885
>TC217695 weakly similar to UP|Q9LUI1 (Q9LUI1) Extensin protein-like, partial
(8%)
Length = 822
Score = 37.4 bits (85), Expect = 0.007
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Frame = +2
Query: 185 PLPQWRHPPPPLESFE-IPPVTIPSPPLQPPQQPP 218
P ++ PPPP+E F PP+ PSPP P++PP
Sbjct: 341 PPIEYLSPPPPIEYFSPPPPIVYPSPPPPSPKKPP 445
Score = 26.9 bits (58), Expect = 9.5
Identities = 21/69 (30%), Positives = 30/69 (43%), Gaps = 13/69 (18%)
Frame = +2
Query: 182 LAGPLP-QWRHPPPPLESFEIPPVTIPSPPLQ----PPQQ--------PPQLLHQESFFN 228
L+ P P ++ PPPP+ PP + PP + PP P L + F+
Sbjct: 356 LSPPPPIEYFSPPPPIVYPSPPPPSPKKPPTKYCPPPPSSAYLYMTGPPGNLYPVDENFS 535
Query: 229 GSSSTSTHS 237
G+SS HS
Sbjct: 536 GASSNRHHS 562
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 37.0 bits (84), Expect = 0.009
Identities = 30/105 (28%), Positives = 42/105 (39%), Gaps = 19/105 (18%)
Frame = -2
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGK----LLQAVMKAGPLLQTLLLAGPLPQWRH 191
P P P P P++ SM + + P K L M P+ Q+L + P +
Sbjct: 408 PPPPLPPPPPNLPPSMPQSLAK*PTSPHRKQPPPLPPPPMLPFPVAQSLAMCPNPPHLKQ 229
Query: 192 ----PPPPLESFEIPPVT-----------IPSPPLQPPQQPPQLL 221
PPPP+ + PP T + PP PP PP+LL
Sbjct: 228 DPPPPPPPMPTLPPPPRTQSLARWPNPPQLKQPPPLPPPPPPELL 94
Score = 28.1 bits (61), Expect = 4.3
Identities = 24/71 (33%), Positives = 29/71 (40%), Gaps = 10/71 (14%)
Frame = -2
Query: 136 PQPQFPLPLPSVAQSM--------IELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLP 187
P P LP P QS+ ++ P P P +LL LLQ+L P
Sbjct: 213 PPPMPTLPPPPRTQSLARWPNPPQLKQPPPLPPPPPPELLL-------LLQSLARCPNSP 55
Query: 188 QWRH--PPPPL 196
W H PPPPL
Sbjct: 54 HW*HAAPPPPL 22
Score = 28.1 bits (61), Expect = 4.3
Identities = 26/88 (29%), Positives = 30/88 (33%)
Frame = -2
Query: 133 DNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHP 192
+NM F LP +V P P PE A P L+ +L P
Sbjct: 543 ENMYYSTFFLPSQNVTN------PSSPQPEASTHSLAKCPGFPQLKQVLFP--------P 406
Query: 193 PPPLESFEIPPVTIPSPPLQPPQQPPQL 220
PPPL P PP PP P L
Sbjct: 405 PPPLP---------PPPPNLPPSMPQSL 349
>TC221182 weakly similar to UP|ENAH_MOUSE (Q03173) Enabled protein homolog
(NPC derived proline-rich protein 1) (NDPP-1), partial
(5%)
Length = 446
Score = 37.0 bits (84), Expect = 0.009
Identities = 20/39 (51%), Positives = 21/39 (53%)
Frame = +3
Query: 180 LLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
LLL PLP HPPPPL PP +P P PP PP
Sbjct: 87 LLLPPPLPLPHHPPPPL-----PP--LPPKPATPPPTPP 182
>TC232500 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (3%)
Length = 425
Score = 36.6 bits (83), Expect = 0.012
Identities = 32/96 (33%), Positives = 37/96 (38%), Gaps = 5/96 (5%)
Frame = -3
Query: 129 SPVFDN--MPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLL---A 183
SP+F +PQP P P I LTP PLP A T LL
Sbjct: 402 SPLFPLGFLPQPDPPWP----TNLNIRLTPTAPLPLPPS------SASTTTTTALLDSPT 253
Query: 184 GPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQ 219
P + RHPPP S +PP P P P P+
Sbjct: 252 APTRRRRHPPPTTTSLPLPP---PPPTSSSPNSAPK 154
>TC217546 weakly similar to UP|Q9GZC7 (Q9GZC7) RNA binding protein RGGm,
partial (10%)
Length = 1047
Score = 36.6 bits (83), Expect = 0.012
Identities = 26/85 (30%), Positives = 30/85 (34%)
Frame = -2
Query: 134 NMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP 193
+ P P+FP P P + P+ LP L GP P
Sbjct: 515 HQPPPEFPDPHPFPDAAFFVADPQPELPCSPSL-----------------GPAPPLACSQ 387
Query: 194 PPLESFEIPPVTIPSPPLQPPQQPP 218
PPL PP PPL PP QPP
Sbjct: 386 PPLSLLLAPP-----PPLPPPPQPP 327
>TC204070 similar to UP|Q39835 (Q39835) Extensin, partial (34%)
Length = 596
Score = 36.6 bits (83), Expect = 0.012
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = +2
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P+ +++ PPPP+ ++ PP P PP + P PP
Sbjct: 296 PVYKYKSPPPPVYKYKSPPPPPPPPPYKYPSPPP 397
Score = 28.1 bits (61), Expect = 4.3
Identities = 12/29 (41%), Positives = 16/29 (54%), Gaps = 3/29 (10%)
Frame = +2
Query: 192 PPPPLESFEIPPVTI---PSPPLQPPQQP 217
PPPP+ ++ PP + SPP PP P
Sbjct: 287 PPPPVYKYKSPPPPVYKYKSPPPPPPPPP 373
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,647,105
Number of Sequences: 63676
Number of extensions: 301686
Number of successful extensions: 7978
Number of sequences better than 10.0: 653
Number of HSP's better than 10.0 without gapping: 3683
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5806
length of query: 266
length of database: 12,639,632
effective HSP length: 96
effective length of query: 170
effective length of database: 6,526,736
effective search space: 1109545120
effective search space used: 1109545120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0369b.1


