

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
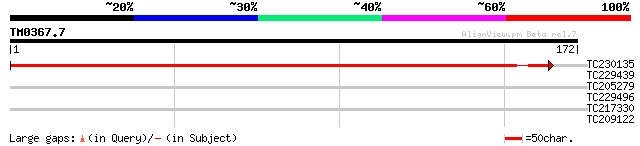
Query= TM0367.7
(172 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC230135 similar to UP|Q71F77 (Q71F77) TK1-like deoxyribonucleos... 250 3e-67
TC229439 weakly similar to UP|Q9LUG6 (Q9LUG6) Dual-specificity p... 30 0.56
TC205279 homologue to PRF|2206327A.0|1587206|2206327A T complex ... 29 0.96
TC229496 weakly similar to UP|Q26882 (Q26882) Surface coat glyco... 27 3.6
TC217330 similar to UP|Q9FH92 (Q9FH92) Emb|CAB62301.1 (At5g67210... 27 6.2
TC209122 similar to UP|Q93VB2 (Q93VB2) AT3g62770/F26K9_200, part... 26 8.1
>TC230135 similar to UP|Q71F77 (Q71F77) TK1-like deoxyribonucleoside kinase
, partial (78%)
Length = 865
Score = 250 bits (638), Expect = 3e-67
Identities = 121/165 (73%), Positives = 142/165 (85%)
Frame = +3
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NV ++KSSKDTRY +DS+VTHDG K PC AL +L SF++K G DAY++LDVIGIDEAQFF
Sbjct: 168 NVVLLKSSKDTRYAIDSVVTHDGIKFPCRALPDLLSFREKHGDDAYQKLDVIGIDEAQFF 347
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
+DLY+FC +AAD DGKTV+VAGLDG+YLRR+FGSVL IIPLADSVTKLTARCE+CGK AF
Sbjct: 348 EDLYEFCCKAADEDGKTVIVAGLDGDYLRRSFGSVLHIIPLADSVTKLTARCELCGKRAF 527
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLESQ 165
FTLRKT++ + ELIGG D+YMPVCR HY+N QVA R VLESQ
Sbjct: 528 FTLRKTEQRETELIGGADLYMPVCRLHYLNSQVA---ERSVLESQ 653
>TC229439 weakly similar to UP|Q9LUG6 (Q9LUG6) Dual-specificity protein
phosphatase-like protein, partial (72%)
Length = 824
Score = 30.0 bits (66), Expect = 0.56
Identities = 14/37 (37%), Positives = 23/37 (61%), Gaps = 3/37 (8%)
Frame = +1
Query: 46 YEQLDVIGIDEA---QFFDDLYDFCREAADHDGKTVV 79
Y+ +DV+ D+ Q+F++ +DF EA HDG +V
Sbjct: 310 YKIIDVVDKDDEDLKQYFNECFDFIDEAKRHDGGVLV 420
>TC205279 homologue to PRF|2206327A.0|1587206|2206327A T complex protein.
{Cucumis sativus;} , partial (52%)
Length = 838
Score = 29.3 bits (64), Expect = 0.96
Identities = 12/43 (27%), Positives = 23/43 (52%)
Frame = +3
Query: 38 KQKFGVDAYEQLDVIGIDEAQFFDDLYDFCREAADHDGKTVVV 80
K K +D E+ + + E ++FDD+ C++ G T+V+
Sbjct: 360 KHKVDIDTVEKFQTLRLQEQKYFDDMVQKCKDV----GATLVI 476
>TC229496 weakly similar to UP|Q26882 (Q26882) Surface coat glycoprotein
TES-120, partial (19%)
Length = 1079
Score = 27.3 bits (59), Expect = 3.6
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = -3
Query: 43 VDAYEQLDVIGIDEAQFFDDLYDFCREAADHDGKTVVVAG 82
VDA+E+L+ GI+E + D D E +D TVV G
Sbjct: 366 VDAFEELNDNGIEENEEEDSEADVMSEGSDGIPNTVVNVG 247
>TC217330 similar to UP|Q9FH92 (Q9FH92) Emb|CAB62301.1 (At5g67210), partial
(87%)
Length = 1091
Score = 26.6 bits (57), Expect = 6.2
Identities = 15/34 (44%), Positives = 18/34 (52%)
Frame = -1
Query: 49 LDVIGIDEAQFFDDLYDFCREAADHDGKTVVVAG 82
L VIGID FF ++ D DG +VVV G
Sbjct: 494 LHVIGIDFRVFFFEVRGVVPVFIDEDGSSVVVEG 393
>TC209122 similar to UP|Q93VB2 (Q93VB2) AT3g62770/F26K9_200, partial (28%)
Length = 692
Score = 26.2 bits (56), Expect = 8.1
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = +1
Query: 70 AADHDGKTVVVAGLDGNYLRRNFGS 94
A H TVV+ G+DG++ R F S
Sbjct: 319 AFGHQKNTVVILGMDGSFYRCQFDS 393
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,794,659
Number of Sequences: 63676
Number of extensions: 78351
Number of successful extensions: 327
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 327
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 327
length of query: 172
length of database: 12,639,632
effective HSP length: 91
effective length of query: 81
effective length of database: 6,845,116
effective search space: 554454396
effective search space used: 554454396
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0367.7


