

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0366.1
(1346 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
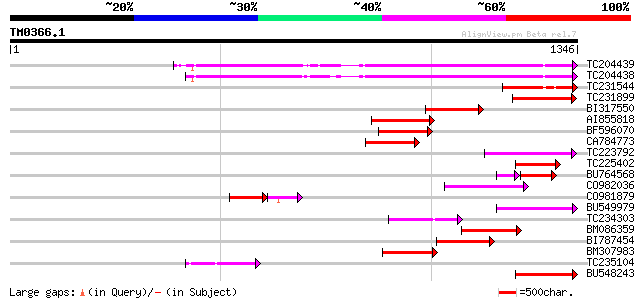
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 505 e-143
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 502 e-142
TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, ... 189 5e-48
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 171 2e-42
BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Ara... 171 2e-42
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 155 1e-37
BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vi... 154 2e-37
CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {V... 154 2e-37
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 147 2e-35
TC225402 weakly similar to UP|Q6I923 (Q6I923) Copia-like retrotr... 136 5e-32
BU764568 108 9e-31
CO982036 130 3e-30
CO981879 91 5e-30
BU549979 130 5e-30
TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, parti... 129 9e-30
BM086359 124 2e-28
BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberos... 124 4e-28
BM307983 120 3e-27
TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotei... 119 1e-26
BU548243 117 3e-26
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 505 bits (1301), Expect = e-143
Identities = 312/971 (32%), Positives = 499/971 (51%), Gaps = 13/971 (1%)
Frame = +1
Query: 389 SSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGHLSHDRLL-------ALHTVQNSI 441
+S SSTC S S E+ +WH R GHL H R + A+ + N +
Sbjct: 2023 TSYSSTCLS---SKEDEVR----------IWHQRFGHL-HLRGMKKIIDKGAVRGIPN-L 2157
Query: 442 SVSKSIVCDVCHLAKQ-KRKMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLD 500
+ + +C C + KQ K + + +L+HMD+ GP+ S+ +Y V+D
Sbjct: 2158 KIEEGRICGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVD 2337
Query: 501 DYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKAIRSDNGPEF---LLPAFYSAQG 557
D+SRF WV + K E + K ++ + ++K IRSD+G EF F +++G
Sbjct: 2338 DFSRFTWVNFIREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEG 2517
Query: 558 IVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKL 617
I H+ S TPQQNG VERK++ + AR +L +LP +W ++ A ++ NR+ +
Sbjct: 2518 ITHEFSAAITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRR 2697
Query: 618 LKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL 677
+ YE+ ++ +FGS C+ R K+DP++ IFLGY + Y +
Sbjct: 2698 GTPTTLYEIWKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRV 2877
Query: 678 MDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQAESTADKLNVASEIQSAKGDA 737
+ T+ + S +V+ + SPA D++ ++ D NVA +S G+
Sbjct: 2878 FNSRTRTVMESINVVVDDL--------SPARK-KDVEEDVRTSGD--NVADAAKS--GEN 3018
Query: 738 SVSTPCASIEDGV-QHEMPSGASIEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKR 796
+ ++ A+ E + Q + S I+ P + + P G T EVE+
Sbjct: 3019 AENSDSATDESNINQPDKRSSTRIQKMHPKELIIGD-PNRGVTTRSREVEI--------- 3168
Query: 797 PVHLANFQFSALPIRSSTAHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHW 856
++ + I EP EA D W
Sbjct: 3169 --------------------------------VSNSCFVSKI----EPKNVKEALTDEFW 3240
Query: 857 REAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQV 916
AMQ E++ ++N W+LV PEG IG KW+++ K N +G + R KARLVA+GY Q+
Sbjct: 3241 INAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQI 3420
Query: 917 EGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPS 976
EG+D+ +TF+PVA+L ++R++L +A + L+Q+DV +AFL+G L+E+VY+ P G
Sbjct: 3421 EGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFAD 3600
Query: 977 -VKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLV 1035
P+ V +L K+LYGLKQA R WYE+L+ L G+++ D +LFVK + +
Sbjct: 3601 PTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQI 3780
Query: 1036 YVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDL 1095
YVDD++ G + + + F + +G L YFLGL+V I L Q +Y ++
Sbjct: 3781 YVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNI 3960
Query: 1096 LQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQL 1155
+++ G + TP ++LS+++ D + YR ++G LLYLT +RPDI++A
Sbjct: 3961 VKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVC 4140
Query: 1156 SQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISG 1215
++Y +NP H +R+L+Y+ + G+++ S + GY DADWAG D R+S SG
Sbjct: 4141 ARYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSG 4320
Query: 1216 YCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLF 1275
CFY+G +L+SW +KKQ VS S+ EAEY A + +L W+ +L++ V L+
Sbjct: 4321 GCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVE-QDVMTLY 4497
Query: 1276 CDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPR 1335
CDN SA++I+ NPV H RTKH+DI H +R+ + ++ L + T Q+AD+FTKAL
Sbjct: 4498 CDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKALDAN 4677
Query: 1336 VFQGFATKLAM 1346
F+ KL +
Sbjct: 4678 QFEKLRGKLGI 4710
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 502 bits (1292), Expect = e-142
Identities = 300/944 (31%), Positives = 484/944 (50%), Gaps = 15/944 (1%)
Frame = +1
Query: 418 LWHFRLGHLSHDRLL-------ALHTVQNSISVSKSIVCDVCHLAKQ-KRKMFTVSVSKA 469
+WH R GHL H R + A+ + N + + + +C C + KQ K +
Sbjct: 2074 IWHQRFGHL-HLRGMKKIIDKGAVRGIPN-LKIEEGRICGECQIGKQVKMSHQKLQHQTT 2247
Query: 470 QKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVK 529
+ +L+HMD+ GP+ S+ +Y V+DD+SRF WV + K + + K ++
Sbjct: 2248 SRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSDTFEVFKELSLRLQ 2427
Query: 530 TQFGQIVKAIRSDNGPEF---LLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIAR 586
+ ++K IRSD+G EF F +++GI H+ S TPQQNG VERK++ + AR
Sbjct: 2428 REKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVERKNRTLQEAAR 2607
Query: 587 ALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFA 646
+L +LP +W ++ A ++ NR+ + + YE+ ++ +FGS C+
Sbjct: 2608 VMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKHFHIFGSPCYI 2787
Query: 647 TTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSP 706
R K+DP++ IFLGY + Y + + T+ + S +V+ + + P + +
Sbjct: 2788 LADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRTVMESINVVVDD-LTPARKKDVE 2964
Query: 707 AWTCLDLQTQAESTADKLNVASEIQ---SAKGDASVSTPCASIEDGVQHEMPSGASIEDE 763
D++T ++ AD A + SA + +++ P +Q P I D
Sbjct: 2965 E----DVRTSGDNVADTAKSAENAENSDSATDEPNINQPDKRPSIRIQKMHPKELIIGD- 3129
Query: 764 IPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYS 823
P G T E+E+
Sbjct: 3130 ----------PNRGVTTRSREIEI------------------------------------ 3171
Query: 824 DERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVK 883
++ + I EP EA D W AMQ E++ ++N W+LV PEG
Sbjct: 3172 -----VSNSCFVSKI----EPKNVKEALTDEFWINAMQEELEQFKRNEVWELVPRPEGTN 3324
Query: 884 PIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAAS 943
IG KW+++ K N +G + R KARLVA+GY Q+EG+D+ +TF+PVA+L ++R++L +A
Sbjct: 3325 VIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPVARLESIRLLLGVACI 3504
Query: 944 QNWHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYEK 1002
+ L+Q+DV +AFL+G L+E+ Y+ P G V P+ V +L K+LYGLKQA R WYE+
Sbjct: 3505 LKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHPDHVYRLKKALYGLKQAPRAWYER 3684
Query: 1003 LSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFG 1062
L+ L G+++ D +LFVK + +YVDD++ G + + + F
Sbjct: 3685 LTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFE 3864
Query: 1063 IKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQG 1122
+ +G L YFLGL+V I L Q KY +++++ G + TP ++LS+++
Sbjct: 3865 MSLVGELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEA 4044
Query: 1123 KPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPG 1182
D + YR ++G LLYLT +RPDI++A ++Y +NP H +R+L+Y+ +
Sbjct: 4045 GTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISHLNQVKRILKYVNGTSD 4224
Query: 1183 LGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEA 1242
G+++ S + GY DADWAG D R+S SG CFY+G +L+SW +KKQ VS S+ EA
Sbjct: 4225 YGIMYCHCSDSMLVGYCDADWAGSADDRKSTSGGCFYLGTNLISWFSKKQNCVSLSTAEA 4404
Query: 1243 EYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCH 1302
EY A + +L W+ +L++ V L+CDN SA++I+ NPV H RTKH+DI H
Sbjct: 4405 EYIAAGSSCSQLVWMKQMLKEYNVE-QDVMTLYCDNMSAINISKNPVQHSRTKHIDIRHH 4581
Query: 1303 VVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
+R+ + ++ L + T Q+AD+FTKAL F+ KL +
Sbjct: 4582 YIRDLVDDKVITLEHVDTEEQIADIFTKALDANQFEKLRGKLGI 4713
>TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, partial (16%)
Length = 662
Score = 189 bits (481), Expect = 5e-48
Identities = 100/182 (54%), Positives = 131/182 (71%), Gaps = 4/182 (2%)
Frame = +3
Query: 1169 AAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWK 1228
AA RVL+YLK P GL F R S I I G+SDADWA C D+ +SI+ YCF++G SL+SWK
Sbjct: 18 AATRVLKYLKGCPRKGLSFSRESPIQILGFSDADWATCIDSSKSITWYCFFLGSSLISWK 197
Query: 1229 AKKQTTVSR--SSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALH-IA 1285
AKKQ TVSR SS+EA+YRAL TCELQWL YLL+DL VT +++CDNQSAL +
Sbjct: 198 AKKQNTVSRSSSSSEAKYRALTSTTCELQWLTYLLKDLHVT-----LIYCDNQSALQ*LP 362
Query: 1286 ANPVFHERTKHLDIDCHVVREKLQAGILK-LLPIPTTLQVADVFTKALQPRVFQGFATKL 1344
++H + L+IDCH+VREK Q G++ LLP+ ++ Q+AD+FTKAL P++F +KL
Sbjct: 363 IKVIYHGQ---LEIDCHIVREKTQQGLMHCLLPVSSSNQLADIFTKALSPKLFSSNLSKL 533
Query: 1345 AM 1346
+
Sbjct: 534 GL 539
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment), partial
(30%)
Length = 687
Score = 171 bits (434), Expect = 2e-42
Identities = 83/150 (55%), Positives = 106/150 (70%)
Frame = +2
Query: 1195 IQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCEL 1254
+ GY DADWAGCP RRS SGYC +IG +LVSWK+KKQT V+RSS EAEYR++A TCEL
Sbjct: 17 LSGYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCEL 196
Query: 1255 QWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILK 1314
W+ LQ+L+ L+CDNQ+ALHIA+NPVFHERTKH++IDCH +REKL + +
Sbjct: 197 MWIKQFLQELRFCEELQMKLYCDNQAALHIASNPVFHERTKHIEIDCHFIREKLLSKEIV 376
Query: 1315 LLPIPTTLQVADVFTKALQPRVFQGFATKL 1344
I + Q D+ TK+L+ Q +KL
Sbjct: 377 TEFIGSNDQPVDILTKSLRGPKIQIVCSKL 466
>BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Arabidopsis
thaliana}, partial (18%)
Length = 421
Score = 171 bits (434), Expect = 2e-42
Identities = 86/139 (61%), Positives = 107/139 (76%)
Frame = -2
Query: 987 KSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNT 1046
KSLYGLKQASRKWYEKL+ L G+ Q+ SD+SLF +G++FT LLVYVDD+IL G++
Sbjct: 420 KSLYGLKQASRKWYEKLTNLLLKEGYIQSISDYSLFTLTKGNTFTALLVYVDDIILAGDS 241
Query: 1047 VTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKP 1106
+ EF +K+ L AF IK+LG LKYFLGLEVAHS GI++ QRKYCLDLL+++G LG KP
Sbjct: 240 IDEFDRIKNVLDLAFKIKNLGKLKYFLGLEVAHSRLGITISQRKYCLDLLKDSGLLGCKP 61
Query: 1107 VATPLDPSIRLSQEQGKPY 1125
+TPLD SI+L G PY
Sbjct: 60 ASTPLDTSIKLHSAAGTPY 4
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 155 bits (391), Expect = 1e-37
Identities = 78/151 (51%), Positives = 105/151 (68%), Gaps = 2/151 (1%)
Frame = -3
Query: 859 AMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEG 918
AMQ E+ E+N WKLV+ PE IG KWV+R K + G + R KARLVAKGYNQ EG
Sbjct: 458 AMQEELNQFERNNVWKLVEKPENYPVIGTKWVFRNKLDEHGIIIRNKARLVAKGYNQEEG 279
Query: 919 LDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAG--VPS 976
+DY +T++PVA+L +R++LA + N+ L+Q+DV +AFL+G + E+VY+ P G +P
Sbjct: 278 IDYEETYAPVARLEVIRMLLAYVSIMNFKLYQMDVKSAFLNGLIQEEVYVEQPPGFEIPD 99
Query: 977 VKPNQVCKLLKSLYGLKQASRKWYEKLSAHL 1007
KP V KL K+LYGLKQA R WYE++S L
Sbjct: 98 -KPTHVYKLQKALYGLKQAPRAWYERISNFL 9
>BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vinifera},
partial (34%)
Length = 407
Score = 154 bits (390), Expect = 2e-37
Identities = 79/130 (60%), Positives = 94/130 (71%), Gaps = 1/130 (0%)
Frame = -2
Query: 876 VDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVR 935
V LP G P+G +WVY VK G + R KARLVAKGY QV G+DY DTFSPVAKLTTVR
Sbjct: 406 VPLPPGKTPVGCRWVYTVKVGPTGEVDRLKARLVAKGYTQVYGIDYCDTFSPVAKLTTVR 227
Query: 936 VILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQ 994
+ LA+AA +W LHQLD+ NAFLHG+L+ED+YM P G V + VCKL +SLYGLKQ
Sbjct: 226 LFLAMAAICHWPLHQLDIKNAFLHGDLEEDIYMEQPPGFVAQGEYGLVCKLHRSLYGLKQ 47
Query: 995 ASRKWYEKLS 1004
+ R W+ K S
Sbjct: 46 SPRAWFGKFS 17
>CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {Vitis
vinifera}, partial (34%)
Length = 409
Score = 154 bits (390), Expect = 2e-37
Identities = 76/128 (59%), Positives = 94/128 (73%)
Frame = +3
Query: 844 PSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLAR 903
PST EA WR+AM E++ALE NGTW+LV LP G +G +WVY VK +G + R
Sbjct: 21 PSTIREALDHPGWRQAMVDEMQALENNGTWELVPLPPGKTTVGCRWVYTVKVGPNGKVDR 200
Query: 904 YKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLD 963
KARLVAKGY QV G++Y DTFSPV LTTVR+ LA+AA ++W LHQLD+ NAFLHG+L+
Sbjct: 201 LKARLVAKGYTQVYGIEYCDTFSPVFFLTTVRLFLAMAAIRHWPLHQLDIKNAFLHGDLE 380
Query: 964 EDVYMTIP 971
ED+YM P
Sbjct: 381 EDIYMEQP 404
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein, partial
(7%)
Length = 804
Score = 147 bits (372), Expect = 2e-35
Identities = 76/221 (34%), Positives = 130/221 (58%), Gaps = 3/221 (1%)
Frame = +1
Query: 1127 DPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLL 1186
D +RRL+G L YL +RP+I A +S++M P H +AA+RVLR +K + G G+L
Sbjct: 10 DVTEFRRLIGSLRYLCNSRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVL 189
Query: 1187 FP---RNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAE 1243
FP ++ ++ GY+D+DW P+ +S GY F + V+ +KKQ ++ S+ EAE
Sbjct: 190 FPFKAKSGKPDLLGYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAE 369
Query: 1244 YRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHV 1303
Y A + C+ W++ LL++LK+ L DN+SA+++A +P H R+KH+++ H
Sbjct: 370 YVAASLGACQAVWMMNLLEELKLRERKPVNLLIDNKSAINLAKHPTLHGRSKHIELRFHY 549
Query: 1304 VREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKL 1344
+R+++ G + + Q+AD+ TK +Q F+ ++L
Sbjct: 550 IRDQVSKGNVTVEYCKAEEQLADLMTKPIQVSRFKQICSEL 672
>TC225402 weakly similar to UP|Q6I923 (Q6I923) Copia-like retrotransposon
Hopscotch polyprotein, partial (7%)
Length = 1446
Score = 136 bits (343), Expect = 5e-32
Identities = 66/109 (60%), Positives = 79/109 (71%)
Frame = +2
Query: 1200 DADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLY 1259
DA+WA P R S GYC IG +LV WK+ K V+RSS EAEY+A+ ATCEL W+
Sbjct: 8 DANWAVSPIDRGSTLGYCVSIGENLVLWKSNK*NVVARSSAEAEYKAMTVATCELIWIKQ 187
Query: 1260 LLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKL 1308
LLQ+LK T L CDNQ+ALHIA+NPVFHERTKH++IDCH VREK+
Sbjct: 188 LLQELKFGSTQQMKLCCDNQAALHIASNPVFHERTKHIEIDCHFVREKV 334
>BU764568
Length = 420
Score = 108 bits (269), Expect(2) = 9e-31
Identities = 51/84 (60%), Positives = 63/84 (74%)
Frame = +3
Query: 1214 SGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPV 1273
SGYC IG +L+SWK+KKQ+ V++SS EAEYRA+A TCEL WL LL +LK
Sbjct: 168 SGYCVLIGGNLISWKSKKQSVVAKSSAEAEYRAMALVTCELIWLKQLL*ELKFEEDTQMT 347
Query: 1274 LFCDNQSALHIAANPVFHERTKHL 1297
L CDNQ+ALHIA+NP+FH RTKH+
Sbjct: 348 LICDNQAALHIASNPIFH*RTKHI 419
Score = 45.4 bits (106), Expect(2) = 9e-31
Identities = 21/55 (38%), Positives = 32/55 (58%)
Frame = +1
Query: 1156 SQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTR 1210
SQ++++P H+ A +L+ K++PG GL++ I GYSDAD G P R
Sbjct: 1 SQFLNSPCQDHWNAVS*ILK*TKSAPGKGLIYEDKGHSQIIGYSDAD*VGSPSDR 165
>CO982036
Length = 674
Score = 130 bits (328), Expect = 3e-30
Identities = 76/203 (37%), Positives = 117/203 (57%), Gaps = 3/203 (1%)
Frame = -2
Query: 1033 LLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYC 1092
LLVYVD +I+ G++ T Q + L+ +F +K LG L YF+ +EV S + R
Sbjct: 646 LLVYVD-IIITGSSCTLIQNLTSKLNSSFPLKLLGKLDYFVEIEVK-SMPDLLFSLRTSI 473
Query: 1093 LDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHAT 1152
++ ++P+++P+ + +LS+ + P YR +VG L Y T RP+IS A
Sbjct: 472 FEIFCRKPR*QAQPISSPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTVIRPEISFAV 293
Query: 1153 QQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGL-LFPRNST--INIQGYSDADWAGCPDT 1209
++ Q+MSNP+D H+ +R+LRYLK S GL L P S+ + I+G+ DADWA D
Sbjct: 292 NKVCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAISSQPLPIRGFCDADWASAVDD 113
Query: 1210 RRSISGYCFYIGRSLVSWKAKKQ 1232
+RS SG ++G +L+SW KQ
Sbjct: 112 KRSTSGAAVFLGPNLISWWXXKQ 44
>CO981879
Length = 576
Score = 90.9 bits (224), Expect(2) = 5e-30
Identities = 44/94 (46%), Positives = 59/94 (61%), Gaps = 3/94 (3%)
Frame = -1
Query: 522 KNFITLVKTQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKH 578
K F +++TQF +K RSDNG E+ L GI+HQ SCV TPQQNG ERK+
Sbjct: 564 KTFFQMIQTQFQVKIKVFRSDNGREYFNKHLSKXXLENGIIHQSSCVDTPQQNGVAERKN 385
Query: 579 QHILNIARALLFQSKLPKKMWCYSVLHAVFLMNR 612
+H+ +ARALLFQ+K PK W ++L +L N+
Sbjct: 384 RHLXEVARALLFQNKAPKYXWGEAILTGTYLKNK 283
Score = 60.5 bits (145), Expect(2) = 5e-30
Identities = 30/89 (33%), Positives = 50/89 (55%), Gaps = 5/89 (5%)
Frame = -2
Query: 612 RIPSKLLKNKSPYELLYREAVDLEM-----MKVFGSLCFATTLTNNRSKLDPRARKCIFL 666
R+PSK+L ++P ++ + + +K+FG F N+ KL+PRA+KC+F+
Sbjct: 284 RMPSKILNFRTPLDVFTSAFPNNRLSCTLPLKIFGCTVFVHIHEPNQGKLEPRAKKCVFV 105
Query: 667 GYKQGMKGYVLMDLITQEIFISRDVIFHE 695
GY KGY D +++ F++ DV F E
Sbjct: 104 GYAPNQKGYKCFDPTSKKTFVTIDVTFFE 18
>BU549979
Length = 615
Score = 130 bits (326), Expect = 5e-30
Identities = 67/194 (34%), Positives = 115/194 (58%), Gaps = 2/194 (1%)
Frame = -1
Query: 1155 LSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSIS 1214
L +Y SNP H+K A++V+RYL+ + L++ + + + + GYSD+D+AGC D+RRS S
Sbjct: 612 LGRYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYKQTNCLEVIGYSDSDFAGCVDSRRSTS 433
Query: 1215 GYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVT-CTATPV 1273
GY F + +VSW++ KQT ++ S+ E E+ AT WL + L+V + P+
Sbjct: 432 GYIFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSLRVVDSISRPL 253
Query: 1274 -LFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKAL 1332
L+CDN +A+ +A N R+KH+DI V+RE+++ + + + T L + D TK +
Sbjct: 252 KLYCDNFAAVFMAKNNKSGNRSKHIDIKYLVIRERVKEKKVVIEHVNTELMIVDPLTKGM 73
Query: 1333 QPRVFQGFATKLAM 1346
P+ F+ ++ +
Sbjct: 72 TPKNFKDHVVRMEL 31
>TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, partial (10%)
Length = 558
Score = 129 bits (324), Expect = 9e-30
Identities = 76/179 (42%), Positives = 101/179 (55%), Gaps = 2/179 (1%)
Frame = +1
Query: 899 GTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFL 958
GT+ ++KARLVAK Y QV G DY TFSPVAK+ V ++ ++A +W L LD NAFL
Sbjct: 28 GTIDQFKARLVAKSYTQVYGQDYTGTFSPVAKMAYVHLLWSMAVVCHWPLF*LDAKNAFL 207
Query: 959 HGNLDEDVYMTIPAG--VPSVKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTA 1016
HG L+E+VYM P G N VC+L +S YGLKQ+ R W + +
Sbjct: 208 HGYLEEEVYMEQPLGFVAQGESSNMVCQLCRSFYGLKQSPRAWPFLYCG--AAIWYDSHE 381
Query: 1017 SDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGL 1075
+DHS+F L+VYVDD+ + G+ +K L F KDLG L+YFLG+
Sbjct: 382 ADHSVFYCHSPQGCIYLIVYVDDIGITGSDQHGIT*LK*XLCCQFQTKDLGKLRYFLGI 558
>BM086359
Length = 427
Score = 124 bits (312), Expect = 2e-28
Identities = 65/142 (45%), Positives = 87/142 (60%)
Frame = +1
Query: 1073 LGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYR 1132
LG++VA S+ GI + Q KY LD+L ETG L P TP+DP+++L QG+ +DP
Sbjct: 1 LGIDVAQSSYGIVISQWKYALDILTETGMLDCLPSNTPMDPNVKLLSGQGEALEDPGR*C 180
Query: 1133 RLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNST 1192
LVGRL YLT TR DI+ A LSQ++ +P D + A R+LRY+K +PG GLL+
Sbjct: 181 CLVGRLNYLTVTRLDITFAVGVLSQFLKDPTDSQWNATIRILRYIKNAPGPGLLYEDKGN 360
Query: 1193 INIQGYSDADWAGCPDTRRSIS 1214
+ Y DADW G P + S S
Sbjct: 361 GKVVCYFDADWPGSPSDKSSTS 426
>BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial
(21%)
Length = 421
Score = 124 bits (310), Expect = 4e-28
Identities = 65/140 (46%), Positives = 89/140 (63%), Gaps = 1/140 (0%)
Frame = +2
Query: 1013 KQTASDHSLFVKFQG-SSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKY 1071
K + +DHS+F L+VYVDD+++ T+ +K+ L F KDL LKY
Sbjct: 2 K*SEADHSVFYCHTSPGKCVYLMVYVDDIMITKKDATKIVQLKEHLFNHFQTKDLRYLKY 181
Query: 1072 FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAY 1131
FLG+EVA S G+ + QRKY LD+L+ETG + V +P+DP+++L Q + Y DP Y
Sbjct: 182 FLGIEVAQSGDGVVISQRKYALDILEETGMQNCRLVDSPMDPNLKLMAYQSEVYPDPERY 361
Query: 1132 RRLVGRLLYLTTTRPDISHA 1151
RRLVG+L+YLT TRPDIS A
Sbjct: 362 RRLVGKLIYLTITRPDISFA 421
>BM307983
Length = 406
Score = 120 bits (302), Expect = 3e-27
Identities = 64/133 (48%), Positives = 87/133 (65%), Gaps = 2/133 (1%)
Frame = +2
Query: 885 IGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAK-LTTVRVILALAAS 943
+G +W+Y VK D TL RYKARLVAKGY Q G+DY +TF+ K + + A
Sbjct: 2 VGCRWIYTVKY*ADDTLDRYKARLVAKGYIQTYGIDYEETFAQWQK*IQSGSSSP*QQAQ 181
Query: 944 QNWHLHQLDVDNAFLHGNLDEDVYMTIPAGV-PSVKPNQVCKLLKSLYGLKQASRKWYEK 1002
W +HQ DV NAFLHG+L+E+VYM IP G S N+VC+L K+LYGLKQ+ R W+ +
Sbjct: 182 FGWEMHQFDVKNAFLHGSLEEEVYMEIPPGYGASNGGNKVCRLKKALYGLKQSPRAWFGR 361
Query: 1003 LSAHLETLGFKQT 1015
+ + +LG+KQ+
Sbjct: 362 FTQAMLSLGYKQS 400
>TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotein
(Fragment), partial (28%)
Length = 865
Score = 119 bits (297), Expect = 1e-26
Identities = 67/181 (37%), Positives = 103/181 (56%), Gaps = 3/181 (1%)
Frame = +2
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVH 477
L H RLGH L L + S+ K + C+ C L K R S+ F ++H
Sbjct: 311 LLHERLGH---PHLSKLKIMVPSLEKIKDLFCESCQLGKHVRSSXRHVESRVDSPFLVIH 481
Query: 478 MDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVK 537
DIWGP +S+ +++YF+T +D++S+ V L+ + E+ + + + +KTQFG+ +K
Sbjct: 482 XDIWGPNRVSSM-SYRYFVTFIDEFSQCTRVFLMKERSEILSFLTS-VNKIKTQFGKTIK 655
Query: 538 AIRSDNGPEF---LLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKL 594
+RSDN E+ ++ F SAQGI+HQ SC TPQQN ERK++H++ AR LL +
Sbjct: 656 ILRSDNAKEYFSSVISPFXSAQGILHQFSCPHTPQQNDIAERKNRHLVETARTLLLHANE 835
Query: 595 P 595
P
Sbjct: 836 P 838
>BU548243
Length = 599
Score = 117 bits (293), Expect = 3e-26
Identities = 62/147 (42%), Positives = 89/147 (60%)
Frame = -1
Query: 1200 DADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLY 1259
DA WA D RS G ++G +L+SW ++KQ ++SS EAEYR++A + EL W+
Sbjct: 587 DAGWASDVDDHRSTLGSAIFLGPNLISWWSRKQQVTAQSSTEAEYRSIAQTSAELTWIQA 408
Query: 1260 LLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIP 1319
LL +L++ T PV+ CDN+SA+ IA N VFH RTKH++ID V EK+ + L++ IP
Sbjct: 407 LLMELQIPFT-PPVILCDNKSAVAIAHNLVFHSRTKHMEIDVFFVHEKVLSKQLQIFHIP 231
Query: 1320 TTLQVADVFTKALQPRVFQGFATKLAM 1346
Q A + TK L F +KL +
Sbjct: 230 ALDQWAGILTKPLSSARFTFLKSKLTV 150
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.144 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,867,658
Number of Sequences: 63676
Number of extensions: 789006
Number of successful extensions: 6061
Number of sequences better than 10.0: 149
Number of HSP's better than 10.0 without gapping: 5915
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6009
length of query: 1346
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1237
effective length of database: 5,698,948
effective search space: 7049598676
effective search space used: 7049598676
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0366.1


