

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0349b.2
(400 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
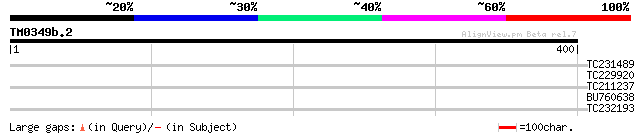
Score E
Sequences producing significant alignments: (bits) Value
TC231489 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F1... 30 2.5
TC229920 weakly similar to UP|O64855 (O64855) Expressed protein ... 28 5.6
TC211237 weakly similar to UP|Q8H3A4 (Q8H3A4) ABC transporter pe... 28 7.3
BU760638 weakly similar to GP|23237921|db contains ESTs D41667(S... 28 7.3
TC232193 weakly similar to UP|Q94AF3 (Q94AF3) AT4g28740/F16A16_1... 28 9.6
>TC231489 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;} , partial (22%)
Length = 634
Score = 29.6 bits (65), Expect = 2.5
Identities = 41/161 (25%), Positives = 64/161 (39%), Gaps = 8/161 (4%)
Frame = +3
Query: 6 MDPPEFDLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLI--- 62
MDP F ++ K + D+ R R R L+LEDS HL+ A QR +
Sbjct: 177 MDPNHFQHQS---KPIQFDSSPTKRAR-RDLLVLEDS------HLNPASAPQQQRRLWVK 326
Query: 63 ---DDYF--LNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPL 117
D++ N P + + FR +RM K F I L S+ + + +E I
Sbjct: 327 DRSKDWWQRCNSPDFPEEEFRLHFRMSKATFDMICQHLDSA----VTKKNTMLREAIPVR 494
Query: 118 AKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGI 158
+ + LA G V + +G +T + + C I
Sbjct: 495 QRVAVCIWRLATGDPLREVSKRFGLGISTCHKLVLEVCSAI 617
>TC229920 weakly similar to UP|O64855 (O64855) Expressed protein
(At2g44200/F6E13.34), partial (19%)
Length = 1372
Score = 28.5 bits (62), Expect = 5.6
Identities = 15/28 (53%), Positives = 18/28 (63%)
Frame = -1
Query: 231 *SLDLACLFWMSGNVELYKRSRPVTSV* 258
*SL CL W SG++ LY S PV S+*
Sbjct: 631 *SLLFLCLCWCSGDLSLYFLSSPVPSL* 548
>TC211237 weakly similar to UP|Q8H3A4 (Q8H3A4) ABC transporter permease
protein-like protein, partial (13%)
Length = 655
Score = 28.1 bits (61), Expect = 7.3
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKG 157
ANGVA ++Y+ + ET CLR + +G
Sbjct: 65 ANGVALSKDEDYLVVCETWKYRCLRHWLEG 154
>BU760638 weakly similar to GP|23237921|db contains ESTs D41667(S4320)
AU033165(S4320)~mucin-like protein, partial (27%)
Length = 430
Score = 28.1 bits (61), Expect = 7.3
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKG 157
ANGVA ++Y+ + ET CLR + +G
Sbjct: 224 ANGVALSKDEDYLVVCETWKYRCLRHWLEG 313
>TC232193 weakly similar to UP|Q94AF3 (Q94AF3) AT4g28740/F16A16_150, partial
(48%)
Length = 760
Score = 27.7 bits (60), Expect = 9.6
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 8/91 (8%)
Frame = -2
Query: 131 VAADAVDEYIKIGE--------TTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVN 182
VA+ +V K GE T+T R KG +RL + A + R + +N
Sbjct: 729 VASSSVSSNSKAGEFPSVTKGTTSTPLSKRLSVKGSLRLKDSVTKDAGPAQMTSRAIPLN 550
Query: 183 EMWGVPKHDREY*LRALRVEKLS*SMRRSIC 213
E G+ R+LR +KLS RR+IC
Sbjct: 549 EFTGMIFFSST---RSLRFDKLSSLDRRAIC 466
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,301,721
Number of Sequences: 63676
Number of extensions: 287464
Number of successful extensions: 1961
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1961
length of query: 400
length of database: 12,639,632
effective HSP length: 99
effective length of query: 301
effective length of database: 6,335,708
effective search space: 1907048108
effective search space used: 1907048108
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0349b.2


