

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0348.12
(180 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
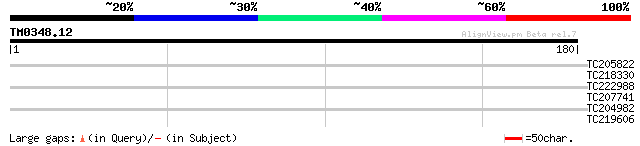
Score E
Sequences producing significant alignments: (bits) Value
TC205822 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {A... 32 0.12
TC218330 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {A... 28 2.3
TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%) 28 2.3
TC207741 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, ... 28 2.3
TC204982 PIR|T06394|T06394 isoprenylated protein - soybean (frag... 27 6.8
TC219606 similar to UP|O48722 (O48722) Heme oxygenase 2 precurso... 26 8.9
>TC205822 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {Arabidopsis
thaliana;} , partial (79%)
Length = 1373
Score = 32.3 bits (72), Expect = 0.12
Identities = 36/120 (30%), Positives = 47/120 (39%)
Frame = +1
Query: 59 PRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAYQNAFGSGL 118
PRCS+R + Y+D YL L A +A G L L+ G +
Sbjct: 652 PRCSLRSRKVYSDLSVD--YLKMLLLNVPATVIALGLFFFLDDLT-----------GFEI 792
Query: 119 SHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGACPNCGEE 178
++ P FS I + L I L +A VK D V L+G CPNCG E
Sbjct: 793 TYLLELPE-PFSFIFTWFAAVPLIVWIALSLTNAIVK--------DFVILKGPCPNCGTE 945
>TC218330 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {Arabidopsis
thaliana;} , partial (61%)
Length = 1066
Score = 28.1 bits (61), Expect = 2.3
Identities = 29/122 (23%), Positives = 52/122 (41%), Gaps = 2/122 (1%)
Frame = +3
Query: 59 PRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAY--QNAFGS 116
PRCS+R + Y+D + ++ L+ A + A+G+ + + G
Sbjct: 294 PRCSLRSRKVYSDLSVDYLKMFLLNVPATVV---------------ALGLFFFLDDVTGF 428
Query: 117 GLSHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGACPNCG 176
+S+ +P I+ F + F++ LA + + I D + L+G CPNCG
Sbjct: 429 EISYMIKSPEPVSFILT---WFAAIPFILW--LAQSITRAIV----QDFLILKGPCPNCG 581
Query: 177 EE 178
E
Sbjct: 582 TE 587
>TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%)
Length = 474
Score = 28.1 bits (61), Expect = 2.3
Identities = 17/59 (28%), Positives = 27/59 (44%)
Frame = +2
Query: 87 WALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVI 145
W + +AFG +A A + N G G S + FSI+V + V+GF++
Sbjct: 239 WEMMIAFGFMA-------AASVRVANELGRGSSKAAK-----FSIVVTVLTSFVIGFIL 379
>TC207741 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, partial
(24%)
Length = 947
Score = 28.1 bits (61), Expect = 2.3
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = +3
Query: 135 NIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGA 171
NI+ +++ FV SAP+ +Q LW N ++ GA
Sbjct: 111 NIVALIINFVSACITGSAPLTAVQLLWVNLIMDTLGA 221
>TC204982 PIR|T06394|T06394 isoprenylated protein - soybean (fragment)
{Glycine max;} , complete
Length = 1230
Score = 26.6 bits (57), Expect = 6.8
Identities = 16/34 (47%), Positives = 17/34 (49%), Gaps = 4/34 (11%)
Frame = -3
Query: 101 PLSYAVGIAY----QNAFGSGLSHGSHNPGLGFS 130
P A+G A N G GLS G PGLGFS
Sbjct: 745 PTGTAIGAAICGCTGNGAGVGLSLGFSGPGLGFS 644
>TC219606 similar to UP|O48722 (O48722) Heme oxygenase 2 precursor (HO2),
partial (66%)
Length = 948
Score = 26.2 bits (56), Expect = 8.9
Identities = 12/39 (30%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Frame = -1
Query: 111 QNAFGSGLSHGSHNPGLGFSI--IVNNIIFIVLGFVIGY 147
QN FG+ + G H+ +GF++ I + +F+ + GY
Sbjct: 138 QNLFGTEIGSGKHSCRMGFAVNHIAPSFLFLPC*LIDGY 22
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,605,224
Number of Sequences: 63676
Number of extensions: 144483
Number of successful extensions: 887
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 880
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 886
length of query: 180
length of database: 12,639,632
effective HSP length: 91
effective length of query: 89
effective length of database: 6,845,116
effective search space: 609215324
effective search space used: 609215324
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0348.12


