

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
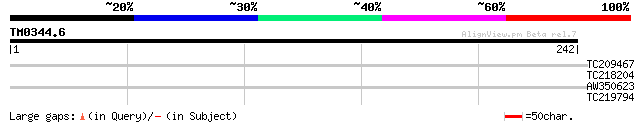
Query= TM0344.6
(242 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC209467 similar to UP|XT32_ARATH (Q9SJL9) Probable xyloglucan e... 35 0.040
TC218204 similar to UP|Q9FJ27 (Q9FJ27) Polygalacturonase-like pr... 28 4.9
AW350623 27 6.4
TC219794 similar to UP|Q9FED2 (Q9FED2) AtHVA22e (Abscisic acid-i... 27 6.4
>TC209467 similar to UP|XT32_ARATH (Q9SJL9) Probable xyloglucan
endotransglucosylase/hydrolase protein 32 precursor
(At-XTH32) (XTH-32) , partial (93%)
Length = 1223
Score = 34.7 bits (78), Expect = 0.040
Identities = 20/73 (27%), Positives = 32/73 (43%), Gaps = 6/73 (8%)
Frame = -2
Query: 3 NLTLNQHANNSEEND------SNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEK 56
N + N S+EN N + NW PK Q+ TSI +FNP+ ++ +
Sbjct: 1108 NFLKEKRQNKSQENSVLVLGTKNQHHNIPNW--PKCSNFQMLNTSILQFNPNTQVLGCDL 935
Query: 57 SFAVQGPYQSISI 69
S+ + Y+ I
Sbjct: 934 SWGLDNSYKPCGI 896
>TC218204 similar to UP|Q9FJ27 (Q9FJ27) Polygalacturonase-like protein,
partial (51%)
Length = 1014
Score = 27.7 bits (60), Expect = 4.9
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +3
Query: 83 GYLHIGLVQVAVKPLTRLRYC 103
G+LH+ ++Q+AVKPL R C
Sbjct: 324 GHLHLPVLQLAVKPLEGWRMC 386
>AW350623
Length = 713
Score = 27.3 bits (59), Expect = 6.4
Identities = 12/32 (37%), Positives = 15/32 (46%), Gaps = 3/32 (9%)
Frame = +1
Query: 135 PAYFNCFSSYPVCLRDPHVTT---CVDLDIKI 163
P+Y+ C YP C P V + C DL I
Sbjct: 436 PSYYPCCGFYPFCFHQPLVKSAVRCYDLSFSI 531
>TC219794 similar to UP|Q9FED2 (Q9FED2) AtHVA22e (Abscisic acid-induced-like
protein), partial (90%)
Length = 709
Score = 27.3 bits (59), Expect = 6.4
Identities = 25/89 (28%), Positives = 41/89 (45%), Gaps = 8/89 (8%)
Frame = +1
Query: 15 ENDSNL--EQKLQNWSIPKIRT-NQVYKTSIFEFNPD-YEIKTVEKSFAVQGPYQSISIL 70
E+ S L EQ L W I T ++ I E+ P Y++K + ++ V + + L
Sbjct: 277 ESQSKLDDEQWLAYWIIYSFLTLTEMVLQPILEWIPIWYDVKLLTVAWLVLPQFAGAAYL 456
Query: 71 DSRIANQHVRKY----GYLHIGLVQVAVK 95
R +H+RKY YL++ Q + K
Sbjct: 457 YERFVREHIRKYITEKEYLYVNHQQQSKK 543
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,567,257
Number of Sequences: 63676
Number of extensions: 160836
Number of successful extensions: 682
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 681
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 682
length of query: 242
length of database: 12,639,632
effective HSP length: 95
effective length of query: 147
effective length of database: 6,590,412
effective search space: 968790564
effective search space used: 968790564
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0344.6


