

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0342b.6
(427 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
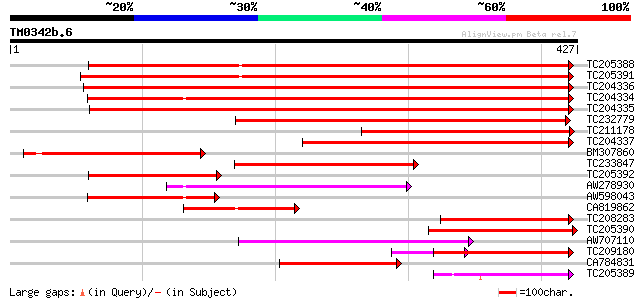
Score E
Sequences producing significant alignments: (bits) Value
TC205388 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partia... 435 e-122
TC205391 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partia... 435 e-122
TC204336 similar to UP|Q41695 (Q41695) Pectinacetylesterase prec... 409 e-114
TC204334 weakly similar to UP|Q41695 (Q41695) Pectinacetylestera... 403 e-113
TC204335 homologue to UP|Q41695 (Q41695) Pectinacetylesterase pr... 394 e-110
TC232779 weakly similar to UP|Q9FF93 (Q9FF93) Pectinacetylestera... 280 9e-76
TC211178 267 8e-72
TC204337 weakly similar to UP|Q41695 (Q41695) Pectinacetylestera... 212 2e-55
BM307860 172 3e-43
TC233847 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2... 161 5e-40
TC205392 143 2e-34
AW278930 weakly similar to PIR|S68805|S68 pectin acetylesterase ... 141 6e-34
AW598043 similar to PIR|S68805|S688 pectin acetylesterase (EC 3.... 136 2e-32
CA819862 129 3e-30
TC208283 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2... 120 2e-27
TC205390 116 2e-26
AW707110 weakly similar to PIR|S68805|S68 pectin acetylesterase ... 114 6e-26
TC209180 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2... 104 9e-23
CA784831 similar to GP|16226325|gb| AT3g05910/F2O10_3 {Arabidops... 96 3e-20
TC205389 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2... 94 2e-19
>TC205388 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partial (86%)
Length = 1433
Score = 435 bits (1119), Expect = e-122
Identities = 192/365 (52%), Positives = 260/365 (70%)
Frame = +1
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++P T + A +GA+CLDG+ PG+HF GFGSG+ SWL+ LEGGGWCN+ISSC RK T
Sbjct: 172 MVPLTLIQGAASKGAVCLDGTLPGYHFHPGFGSGANSWLIQLEGGGWCNTISSCVFRKTT 351
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSSKYME + F+G+LS+ +NPDFFNWN+V +RYCDGASF+G ++E L FRG
Sbjct: 352 RRGSSKYMEKQLAFTGLLSNKAEENPDFFNWNRVKVRYCDGASFSGDSQNEAAQ-LQFRG 528
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
Q IW+A M ELL GM KA QALLSGCSAGGLA+++HCD+FR + P T VKCL+DAGFF
Sbjct: 529 QKIWQAAMQELLFKGMQKANQALLSGCSAGGLASIIHCDEFRSLFPTSTKVKCLSDAGFF 708
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
LD D+ G +T+R+ + VV+LQ V K+L CL +++P+ C FP ++ ++TP+FL++
Sbjct: 709 LDAVDVSGGHTLRNLFGGVVKLQEVQKNLPNSCLNQLDPTSCFFPQNLINYVETPLFLLN 888
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQ++ LVP +DP G W CK N +CN++ I L FR+ +L + F + Q
Sbjct: 889 AAYDAWQVQESLVPHSADPHGSWNDCKANHAHCNSSQIQFLQDFRNQMLNDVKGFSETSQ 1068
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
G+F+NSCF HCQ+ R +TW++ +SP I N +A +V DWF DR+ + IDC YPC+ TC
Sbjct: 1069TGLFINSCFAHCQSERQDTWFADDSPLINNVPVAIAVGDWFLDRKTVKAIDCAYPCDNTC 1248
Query: 420 RNLDF 424
NL F
Sbjct: 1249HNLVF 1263
>TC205391 similar to UP|Q8L7Z5 (Q8L7Z5) At1g57590/T8L23_6, partial (82%)
Length = 1469
Score = 435 bits (1118), Expect = e-122
Identities = 194/371 (52%), Positives = 265/371 (71%)
Frame = +1
Query: 54 SHSLSNLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSC 113
S S ++ T + A +GA+CLDG+ PG+H +GFGSG+ SWL+HLEGGGWCN+I +C
Sbjct: 178 SSSQPLMVDLTLIQGADSKGAVCLDGTVPGYHLDRGFGSGADSWLIHLEGGGWCNTIRNC 357
Query: 114 SRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGS 173
RK T GSSKYME +PF+GILS+ +NPDFFNWN+V +RYCDGASF+G E E
Sbjct: 358 VYRKNTRRGSSKYMENQIPFTGILSNKPEENPDFFNWNRVKLRYCDGASFSGDSEDESAQ 537
Query: 174 GLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCL 233
L FRGQ IW A M+EL+S GM KA QALLSGCSAGGLA+++HCD+FR + K + VKCL
Sbjct: 538 -LQFRGQKIWLAAMEELMSKGMQKADQALLSGCSAGGLASIIHCDEFRSLFGKSSKVKCL 714
Query: 234 ADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKT 293
+D GFFLD D+ G T+R+ + VVQLQ V K+L K CL +++P+ C FP ++++++T
Sbjct: 715 SDGGFFLDAMDVSGGRTLRTLFGGVVQLQEVQKNLPKSCLDQLDPTSCFFPQNMIEHVET 894
Query: 294 PVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNE 353
P+FL+++AYD WQ++ L P +D G W CK N NC+++ + L FR+ +L + +
Sbjct: 895 PLFLLNAAYDVWQVQASLAPPSADRLGSWNECKSNHANCSSSQMQFLQDFRNQMLGDIKD 1074
Query: 354 FQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPY 413
F Q G+F+NSCF HCQ+ R ETW++ +SP I++K IA +V DW+FDREV+ IDC Y
Sbjct: 1075FSSSSQTGLFINSCFAHCQSERQETWFADDSPLIEDKPIAVAVGDWYFDREVVKAIDCAY 1254
Query: 414 PCNPTCRNLDF 424
PC+ +C NL F
Sbjct: 1255PCDNSCHNLVF 1287
>TC204336 similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor ,
partial (98%)
Length = 1592
Score = 409 bits (1050), Expect = e-114
Identities = 176/370 (47%), Positives = 259/370 (69%), Gaps = 1/370 (0%)
Frame = +1
Query: 56 SLSNLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSR 115
S ++ +P T + NA+ +GA+CLDGS P +HF +GFGSG +WL+ EGGGWCN++++C
Sbjct: 112 SEASYVPITIVQNAVAKGAVCLDGSPPAYHFDRGFGSGINNWLVAFEGGGWCNNVTTCLA 291
Query: 116 RKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKG-SG 174
RK LGSSK M ++ FSGIL++ NPDF+NWN++ +RYCDG+SF G E+ +
Sbjct: 292 RKTNRLGSSKQMAKLIAFSGILNNREMFNPDFYNWNRIKVRYCDGSSFTGDVEAVNPVTK 471
Query: 175 LFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLA 234
L FRG I+ A+M++LL+ GM A+ A++SGCSAGGL +++HCD FR +LP+ VKCL+
Sbjct: 472 LHFRGGRIFNAVMEDLLAKGMKNARNAIISGCSAGGLTSVLHCDRFRALLPRGARVKCLS 651
Query: 235 DAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTP 294
DAG+F++ KD+ G + +++ VV G A++L + C +++ P C FP ++ I TP
Sbjct: 652 DAGYFINGKDVLGEQHIEQYFSQVVATHGSARNLPQSCTSRLSPRLCFFPQYLVSRITTP 831
Query: 295 VFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEF 354
+F V++AYD WQI+NIL P +DP GHW SCKL+I+NC+ + +D++ FR+ L+A+
Sbjct: 832 IFFVNAAYDSWQIKNILAPGVADPEGHWHSCKLDINNCSPDQLDLMQGFRTEFLRAITVL 1011
Query: 355 QQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYP 414
GMF++SC+ HCQT ETW +SP+++ TIA++VADWF++R + IDCPYP
Sbjct: 1012GNSSSKGMFIDSCYAHCQTEMQETWLRSDSPELKKTTIAKAVADWFYERRPFHQIDCPYP 1191
Query: 415 CNPTCRNLDF 424
CNPTC N F
Sbjct: 1192CNPTCHNRVF 1221
>TC204334 weakly similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor
, partial (95%)
Length = 1508
Score = 403 bits (1036), Expect = e-113
Identities = 179/367 (48%), Positives = 254/367 (68%), Gaps = 1/367 (0%)
Frame = +1
Query: 59 NLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKF 118
+L+P T + NA +GA+CLDGS P +HF GF G ++W++H+EGGGWCN++ SC RK
Sbjct: 154 SLVPLTLVKNAESKGAVCLDGSPPAYHFDNGFEEGIKNWIVHIEGGGWCNNVESCLYRKD 333
Query: 119 TALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFF 177
+ LGSSK ME + FS ILS+++ NPDF+NWN+V +RYCDG+SF G E ++ + L F
Sbjct: 334 SRLGSSKQMEDLY-FSAILSNEQEYNPDFYNWNRVKVRYCDGSSFTGDVEEVDQTTNLHF 510
Query: 178 RGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAG 237
RG I+ A+M+ELL+ G+ KA+ A+LSGCSAGGL T++HCD F+ +LP E VKC+ DAG
Sbjct: 511 RGARIFSAVMEELLAKGLEKAENAILSGCSAGGLTTILHCDRFKNLLPSEANVKCVPDAG 690
Query: 238 FFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFL 297
+F++ +DI G + + FY++VV G AK+L C +K P C FP + +I TP+F+
Sbjct: 691 YFVNVEDISGTHFIEKFYSEVVSTHGSAKNLPSSCTSKFSPELCFFPQYVASHISTPIFV 870
Query: 298 VHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQR 357
V++AYD WQI+NI VP +DPS W SCKL+I NC+ + + + F+S KA++
Sbjct: 871 VNAAYDSWQIQNIFVPGSADPSDSWHSCKLDISNCSPDQLSKMQDFKSEFEKAVSVVGDS 1050
Query: 358 KQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNP 417
GMF++SC+ HCQT ETWY +SP++ N TIA++V DWF+ R +DC YPCNP
Sbjct: 1051SSKGMFIDSCYAHCQTESQETWYKSDSPQLANTTIAKAVGDWFYGRSSFRHVDCNYPCNP 1230
Query: 418 TCRNLDF 424
+C+N F
Sbjct: 1231SCQNRVF 1251
>TC204335 homologue to UP|Q41695 (Q41695) Pectinacetylesterase precursor ,
complete
Length = 1724
Score = 394 bits (1013), Expect = e-110
Identities = 169/365 (46%), Positives = 247/365 (67%), Gaps = 1/365 (0%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+ T + NA+ +GA+CLDGS P +HF KG G+G +W++H EGGGWCN++++C R+ T
Sbjct: 315 VGITFVENAVAKGAVCLDGSPPAYHFHKGSGAGINNWIVHFEGGGWCNNVTTCLSRRDTR 494
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
LGSSK M+T + FSG S+ + NPDF++WN++ +RYCDG+SF G E+ + + L FRG
Sbjct: 495 LGSSKKMDTSLSFSGFFSNSKKFNPDFYDWNRIKVRYCDGSSFTGDVEAVDPKTNLHFRG 674
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
++ ++++LL+ GM A+ A++SGCSAGGLA++++CD F+ +LP T VKCLADAGFF
Sbjct: 675 ARVFAVVVEDLLAKGMKNAQNAIISGCSAGGLASILNCDRFKSLLPATTKVKCLADAGFF 854
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
++ KD+ G + FY+ VVQ G AK+L C +++ P C FP ++ I TP+F V+
Sbjct: 855 INVKDVSGAQRIEEFYSQVVQTHGSAKNLPTSCTSRLRPGLCFFPQNVVSQISTPIFFVN 1034
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+NIL P +DP G W+ CKL+I NC+ N + ++ FR+ L+A +
Sbjct: 1035AAYDSWQIKNILAPGAADPRGQWRECKLDIKNCSPNQLSVMQGFRTDFLRAFSVVGNAAS 1214
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
G F++ C+ HCQT ETW +SP + +IA++V DWF+DR IDC YPCNPTC
Sbjct: 1215KGHFIDGCYAHCQTGIQETWLRNDSPVVAKTSIAKAVGDWFYDRRPFREIDCAYPCNPTC 1394
Query: 420 RNLDF 424
N F
Sbjct: 1395HNRIF 1409
>TC232779 weakly similar to UP|Q9FF93 (Q9FF93) Pectinacetylesterase, partial
(55%)
Length = 1178
Score = 280 bits (716), Expect = 9e-76
Identities = 134/254 (52%), Positives = 175/254 (68%), Gaps = 2/254 (0%)
Frame = +3
Query: 171 KGSGLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTV 230
K + L F+GQ IWEAI+ +LL +G+ KA++ALLSGCSAGGLAT HCD+F + LP +V
Sbjct: 69 KTTTLHFKGQKIWEAIIRDLLPLGLGKARKALLSGCSAGGLATFHHCDNFAKYLPTNASV 248
Query: 231 KCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECL-AKMEPSKCLFPSEILK 289
KCL+DAGFFLDE+DI N+TMR + +VQLQG+ K+L+ C A P C FP L+
Sbjct: 249 KCLSDAGFFLDERDISLNHTMRYNFKSLVQLQGIEKNLNGNCTRALYFPDLCFFPQYALR 428
Query: 290 NIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLK 349
I TP F+++SAYD +Q +ILVP +D GHWK CK N+ C A+ ID L FR +L
Sbjct: 429 YISTPYFILNSAYDVFQFSHILVPPSADMHGHWKQCKANLAECTADQIDTLQGFRLDMLG 608
Query: 350 ALNEF-QQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINL 408
AL F ++ GMF+NSCF HCQ+ ETW+ +SP+I NKTIAE++ DW+F R +
Sbjct: 609 ALRPFYMNSRRGGMFINSCFAHCQSELQETWFGDDSPRINNKTIAEAIGDWYFSRNISKE 788
Query: 409 IDCPYPCNPTCRNL 422
IDC YPC+ TC NL
Sbjct: 789 IDCAYPCDATCHNL 830
>TC211178
Length = 852
Score = 267 bits (682), Expect = 8e-72
Identities = 118/161 (73%), Positives = 140/161 (86%), Gaps = 1/161 (0%)
Frame = +1
Query: 266 KSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSC 325
KSLHK+C+AKMEPSKCLFPSEI KNIKTP+FLVH AYDFWQIRNILVP+GSDP GHW+ C
Sbjct: 1 KSLHKDCIAKMEPSKCLFPSEIAKNIKTPLFLVHPAYDFWQIRNILVPQGSDPDGHWQRC 180
Query: 326 KLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSP 385
+L+I +CNAN+ID L+ +R SLLKA+NEFQQRK+IGMF++SCF+HCQT TW+SPNSP
Sbjct: 181 RLDIRSCNANMIDKLDSYRGSLLKAVNEFQQRKEIGMFIDSCFVHCQTEMEVTWHSPNSP 360
Query: 386 KIQNKTIAESVADWFFDREVINLIDC-PYPCNPTCRNLDFT 425
KI +KTIAESV DW+FDRE + IDC + CNPTC N+DFT
Sbjct: 361 KINDKTIAESVGDWYFDREAVKRIDCSSFSCNPTCHNMDFT 483
>TC204337 weakly similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor
, partial (49%)
Length = 730
Score = 212 bits (540), Expect = 2e-55
Identities = 92/204 (45%), Positives = 134/204 (65%)
Frame = +1
Query: 221 RQILPKETTVKCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSK 280
+ LP VKC+ DAG+F++ +DI G + ++ +Y++VV G AK+L C +K+ P+
Sbjct: 1 KTFLPSRANVKCVPDAGYFVNVEDISGAHFIQQYYSEVVSTHGSAKNLPTSCTSKLSPTL 180
Query: 281 CLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDIL 340
C FP + +I TP+F+V+SAYD WQIR I VP +DPS W SCK+N+ NC+ + + L
Sbjct: 181 CFFPQYVASHISTPIFVVNSAYDSWQIRYIFVPGSADPSDSWNSCKVNMSNCSPDQLSKL 360
Query: 341 NRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWF 400
F+S +AL+E GMF++SC+ HCQT ETW+ +SPK+ N TIA++VADWF
Sbjct: 361 QGFKSEFERALSEVGDSPSKGMFIDSCYAHCQTEPQETWFKTDSPKLANTTIAKAVADWF 540
Query: 401 FDREVINLIDCPYPCNPTCRNLDF 424
+ R +DC YPCNP+C+N F
Sbjct: 541 YGRSSFRHVDCNYPCNPSCQNRVF 612
>BM307860
Length = 418
Score = 172 bits (436), Expect = 3e-43
Identities = 87/138 (63%), Positives = 104/138 (75%), Gaps = 1/138 (0%)
Frame = +2
Query: 11 NLRLRTLVVFFRKLTTKDYAIAAFAFFLLLSLAFFSQLHHHHHSHSL-SNLIPFTPLPNA 69
NLRLRTL ++ ++KDYAIAAF F +L SL FS L + S S L+P T L NA
Sbjct: 17 NLRLRTLRIW----SSKDYAIAAFIFLILFSLLLFSHLDSRYDSQPHHSKLVPLTLLRNA 184
Query: 70 ILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYMET 129
A+CLDGSAPG+HFQ GFGSGSR+WL+H+EGGGWCNSI SC +RKFT LGSS +ME
Sbjct: 185 NQTRALCLDGSAPGYHFQSGFGSGSRNWLIHIEGGGWCNSIPSCYQRKFTHLGSSDHMEK 364
Query: 130 VVPFSGILSSDRSQNPDF 147
++PFSGILSSD +QNPDF
Sbjct: 365 LIPFSGILSSDPAQNPDF 418
>TC233847 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2O10_3
{Arabidopsis thaliana;} , partial (33%)
Length = 421
Score = 161 bits (408), Expect = 5e-40
Identities = 71/139 (51%), Positives = 105/139 (75%)
Frame = +3
Query: 170 EKGSGLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETT 229
+K + L FRGQ IW A +++L+S GM A+QALLSGCSAGGLAT++HCD+FR P+ T
Sbjct: 3 DKAAQLQFRGQRIWSAAIEDLMSKGMRFARQALLSGCSAGGLATIIHCDEFRGFFPQTTK 182
Query: 230 VKCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILK 289
VKCL+DAG FLD D+ +T+R+F++ VV+LQGV K+L C + ++P+ C FP ++
Sbjct: 183 VKCLSDAGLFLDAIDVSRGHTIRNFFSGVVRLQGVQKNLPHICTSHLDPTSCFFPQNLIA 362
Query: 290 NIKTPVFLVHSAYDFWQIR 308
I+TP+F++++AYD WQ++
Sbjct: 363 GIRTPLFILNTAYDSWQVQ 419
>TC205392
Length = 517
Score = 143 bits (360), Expect = 2e-34
Identities = 60/100 (60%), Positives = 77/100 (77%)
Frame = +3
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++P T + A +GA+CLDG+ PG+HF GFGSG+ SWL+ LEGGGWCN+I SC RK T
Sbjct: 135 MVPLTLIQGAASKGAVCLDGTLPGYHFHPGFGSGANSWLIQLEGGGWCNTIRSCVFRKTT 314
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCD 159
GSSKYME + F+GILS+ +NPDFFNWN+V++RYCD
Sbjct: 315 RRGSSKYMEKQLAFTGILSNKAEENPDFFNWNRVIVRYCD 434
>AW278930 weakly similar to PIR|S68805|S68 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (35%)
Length = 622
Score = 141 bits (355), Expect = 6e-34
Identities = 75/185 (40%), Positives = 111/185 (59%), Gaps = 1/185 (0%)
Frame = +2
Query: 119 TALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFF 177
T LGS K ME + FS ILS+++ NPDF+N N+V RYC G+S G E ++ + L F
Sbjct: 2 THLGSFKQMEDLY-FSAILSNEQEYNPDFYNXNRVKARYCXGSSVTGDVEEVDQTTNLHF 178
Query: 178 RGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAG 237
RG I A+M+ELL+ + KA+ A+LSGC AGGL T++HCD + +LP E VKC++DAG
Sbjct: 179 RGARILPAVMEELLANRLEKAENAILSGCPAGGLTTILHCDRCKNLLPSEAYVKCVSDAG 358
Query: 238 FFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFL 297
++ +DI + + FY VV G+A++ + P C FP + +I T +
Sbjct: 359 HXVNVEDIIRTHIIEKFYTQVVSTHGLAQNSPSCRTY*VSPQLCYFPQYVASHISTSILF 538
Query: 298 VHSAY 302
++ Y
Sbjct: 539 GNAHY 553
>AW598043 similar to PIR|S68805|S688 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (23%)
Length = 414
Score = 136 bits (342), Expect = 2e-32
Identities = 59/100 (59%), Positives = 76/100 (76%)
Frame = +3
Query: 59 NLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKF 118
+L+P + NA +GA+CLDGS P +HF KGFG G SW++H+EGGGWCN+I SC RK
Sbjct: 117 SLVPLILVENAESKGAVCLDGSPPAYHFDKGFGEGINSWIVHIEGGGWCNNIESCLDRKD 296
Query: 119 TALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYC 158
T LGSSK ME + FSGILS+++ NPDF+NWN+V +RYC
Sbjct: 297 TRLGSSKQMEDIY-FSGILSNEQQFNPDFYNWNRVKVRYC 413
>CA819862
Length = 409
Score = 129 bits (323), Expect = 3e-30
Identities = 58/87 (66%), Positives = 73/87 (83%)
Frame = +1
Query: 132 PFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRGQIIWEAIMDELL 191
PF GILS+ +NPDFFNWN++ IRYCDGASF+G ++ G+GL+FRGQ IW+A M++L+
Sbjct: 1 PFIGILSNKAEENPDFFNWNRIKIRYCDGASFSGDSQNA-GAGLYFRGQRIWQAAMEDLM 177
Query: 192 SIGMSKAKQALLSGCSAGGLATLVHCD 218
S GM AKQALLSGCSAGGLAT++HCD
Sbjct: 178 SKGMRYAKQALLSGCSAGGLATIIHCD 258
>TC208283 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2O10_3
{Arabidopsis thaliana;} , partial (25%)
Length = 871
Score = 120 bits (300), Expect = 2e-27
Identities = 51/100 (51%), Positives = 69/100 (69%)
Frame = +2
Query: 325 CKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNS 384
C+LN C ++ I L FR+ +L A+ F + +Q G+F+NSCF HCQ+ R +TW++ NS
Sbjct: 23 CRLNHAKCTSSQIQYLQGFRNQMLNAIKGFSRSRQNGLFINSCFAHCQSERQDTWFADNS 202
Query: 385 PKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
P I NK IA SV DW+FDR V+ IDCPYPC+ TC +L F
Sbjct: 203 PVIGNKAIALSVGDWYFDRAVVKAIDCPYPCDNTCHHLVF 322
>TC205390
Length = 804
Score = 116 bits (290), Expect = 2e-26
Identities = 51/112 (45%), Positives = 68/112 (60%)
Frame = +2
Query: 316 SDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWR 375
+DP G CK N CN++ I L FR+ +L + F Q G+F+NSC HCQ+ R
Sbjct: 17 ADPHGSXXDCKSNHARCNSSQIQFLQDFRNQMLNDVKGFSGTSQTGLFINSCXAHCQSER 196
Query: 376 GETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDFTRV 427
+TW++ +SP I N IA +V DWFFDR+ + IDC YPC+ TC NL F V
Sbjct: 197 QDTWFADDSPLINNMPIAIAVGDWFFDRKTVKAIDCAYPCDNTCHNLVFNAV 352
>AW707110 weakly similar to PIR|S68805|S68 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (42%)
Length = 547
Score = 114 bits (286), Expect = 6e-26
Identities = 58/177 (32%), Positives = 100/177 (55%)
Frame = +2
Query: 173 SGLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKC 232
+ L FR ++ ++ +LLS G+ A+ A++S SA L ++++ D F+ +LP T VK
Sbjct: 17 TNLHFRRARVFAVVIKDLLSKGIKNAQNAIISRYSAKKLTSILNYDQFKSLLPTTTKVKY 196
Query: 233 LADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIK 292
L D FF++ K+I ++ FY+ V+Q AK+L +K+ P FP I+ I
Sbjct: 197 LTDTNFFINSKNISKAQHIKEFYSQVIQTHNSAKNLPTSYTSKLRPGLYFFPQNIISQIS 376
Query: 293 TPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLK 349
TP+F +++AY+ QI+NIL P ++P + +L+I N + N + I+ + LK
Sbjct: 377 TPIFFINTAYNS*QIKNILTPNTTNPHNQ*REYELDIKNYSPNQLSIVTDIHNRFLK 547
>TC209180 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2O10_3
{Arabidopsis thaliana;} , partial (25%)
Length = 678
Score = 104 bits (259), Expect = 9e-23
Identities = 45/105 (42%), Positives = 67/105 (62%)
Frame = +3
Query: 320 GHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETW 379
G + + N C+A I L FR+ +L++ F + + G+F+NSCF HCQ+ R +TW
Sbjct: 99 GTGMNAEKNYARCSAPQIQYLQGFRNQMLRSTRAFSRSYKNGLFINSCFAHCQSERQDTW 278
Query: 380 YSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
++ +SP+I N+ IAESV +W+F R + I CPYPC+ TC NL F
Sbjct: 279 FAHDSPRIGNRGIAESVGNWYFGRVSVQAIGCPYPCDKTCHNLVF 413
Score = 44.3 bits (103), Expect = 1e-04
Identities = 18/59 (30%), Positives = 32/59 (53%)
Frame = +2
Query: 288 LKNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSS 346
+ ++TP+FL+++AYD WQI+ L P +D +W C+ + + +RF S
Sbjct: 2 IAGVRTPLFLLNAAYDTWQIQASLAPPSADYHWNWYECRKKLCTLQCTTDTVSSRFSKS 178
>CA784831 similar to GP|16226325|gb| AT3g05910/F2O10_3 {Arabidopsis
thaliana}, partial (20%)
Length = 395
Score = 95.9 bits (237), Expect = 3e-20
Identities = 43/92 (46%), Positives = 64/92 (68%)
Frame = +3
Query: 204 SGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQG 263
SGCSAGGLAT++HCD+FR P+ T VKCL+DAG FLD D+ +T+R+F++ VV+LQG
Sbjct: 3 SGCSAGGLATIIHCDEFRGFFPQTTKVKCLSDAGLFLDAIDVSRGHTIRNFFSGVVRLQG 182
Query: 264 VAKSLHKECLAKMEPSKCLFPSEILKNIKTPV 295
V K+L C + ++P+ + +N++ V
Sbjct: 183 VQKNLPHICTSHLDPTSVMINIFYFENLEKSV 278
>TC205389 similar to GB|AAL16135.1|16226325|AF428303 AT3g05910/F2O10_3
{Arabidopsis thaliana;} , partial (16%)
Length = 538
Score = 93.6 bits (231), Expect = 2e-19
Identities = 45/107 (42%), Positives = 62/107 (57%), Gaps = 2/107 (1%)
Frame = +2
Query: 320 GHWKSCKLNIHNCNANLIDILNRFRSSLLKALNE--FQQRKQIGMFVNSCFIHCQTWRGE 377
G W C +H +L + R + ++ E Q Q G+F+NSCF HCQ+ R +
Sbjct: 5 GSWNDCNQIMHAVT-HLNSVPPRLQEIKVRK*CERLSQGHSQTGLFINSCFAHCQSERQD 181
Query: 378 TWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
TW++ +SP I N IA +V DWFFDR+ + IDC YPC+ TC NL F
Sbjct: 182 TWFADDSPLINNMPIAIAVGDWFFDRKTVKAIDCAYPCDNTCHNLVF 322
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.138 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,017,032
Number of Sequences: 63676
Number of extensions: 458206
Number of successful extensions: 6587
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 4905
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5841
length of query: 427
length of database: 12,639,632
effective HSP length: 100
effective length of query: 327
effective length of database: 6,272,032
effective search space: 2050954464
effective search space used: 2050954464
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0342b.6


