

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
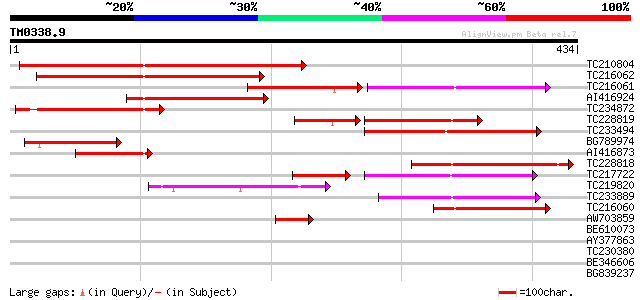
Query= TM0338.9
(434 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC210804 homologue to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein ... 332 2e-91
TC216062 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 293 8e-80
TC216061 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 137 5e-54
AI416924 181 7e-46
TC234872 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 171 5e-43
TC228819 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 83 3e-32
TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 133 2e-31
BG789974 similar to GP|21553837|gb| phytochelatin synthetase-lik... 129 3e-30
AI416873 107 8e-24
TC228818 similar to UP|CBL1_ARATH (Q9SRT7) COBRA-like protein 1 ... 107 1e-23
TC217722 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 ... 63 3e-22
TC219820 similar to GB|AAO64838.1|29028918|BT005903 At4g16120 {A... 74 1e-13
TC233889 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 ... 67 2e-11
TC216060 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 66 3e-11
AW703859 homologue to GP|26452135|dbj GPI-anchored protein {Arab... 58 9e-09
BE610073 29 3.6
AY377863 29 4.7
TC230380 similar to GB|AAO64866.1|29028974|BT005931 At4g29000 {A... 28 6.2
BE346606 similar to GP|13605910|gb| AT3g55400/T22E16_60 {Arabido... 28 6.2
BG839237 similar to GP|3688193|emb MAP3K alpha 1 protein kinase ... 28 6.2
>TC210804 homologue to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor,
partial (47%)
Length = 759
Score = 332 bits (851), Expect = 2e-91
Identities = 147/220 (66%), Positives = 178/220 (80%)
Frame = +1
Query: 8 RFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEP 67
R V+ ++L S + AYDPLDPNGNITI+WD++SWT DGYVA VT++NFQ +RHI P
Sbjct: 103 RVVISAVCAIVLFSYAAAYDPLDPNGNITIKWDVVSWTPDGYVAVVTMSNFQMFRHIMNP 282
Query: 68 GWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVA 127
GW+LGWTWAK E+IW M+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++Q +
Sbjct: 283 GWTLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQQFS 459
Query: 128 NCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVK 187
NCCKGGV+ + QDP++AV+SF+V +G AGT+ K+V+LPKNFT PGPGYTCGPA++V
Sbjct: 460 NCCKGGVVAAWGQDPSSAVSSFQVSIGLAGTSNKTVKLPKNFTLLGPGPGYTCGPAKVVP 639
Query: 188 PTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSL 227
T FL DKRR TQA+M+WNVTCTYSQFLA K P CCVSL
Sbjct: 640 STVFLTPDKRRKTQALMTWNVTCTYSQFLARKNPGCCVSL 759
>TC216062 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (40%)
Length = 550
Score = 293 bits (751), Expect = 8e-80
Identities = 127/175 (72%), Positives = 150/175 (85%)
Frame = +1
Query: 21 SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEI 80
+S EAYDPLDP GNITI+WD++SWT DGYVA VT+ NFQQYRHIQ PGWSLGWTWAK E+
Sbjct: 28 TSIEAYDPLDPTGNITIKWDVISWTPDGYVAVVTMYNFQQYRHIQAPGWSLGWTWAKKEV 207
Query: 81 IWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQ 140
IW M+G QTTEQGDCSKFK + PHCCKKDP VDL+PG PY++Q+ANCCKGGVL S Q
Sbjct: 208 IWNMMGAQTTEQGDCSKFKAGI-PHCCKKDPTVVDLLPGTPYNQQIANCCKGGVLNSWGQ 384
Query: 141 DPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYD 195
DP+NAV+SF++ VG AGTT K+V++PKNFT K+PGPGYTCGPA++VKPT F+ D
Sbjct: 385 DPSNAVSSFQISVGSAGTTNKTVKMPKNFTLKAPGPGYTCGPAKVVKPTVFITND 549
>TC216061 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (52%)
Length = 1100
Score = 137 bits (345), Expect(2) = 5e-54
Identities = 60/92 (65%), Positives = 75/92 (81%), Gaps = 4/92 (4%)
Frame = +2
Query: 183 ARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCA 242
A++ KPT F+ DKRR TQAMM+WN+TCTYSQFLA KAP+CCVSLS+FYNDT+V+CPTC
Sbjct: 2 AKVEKPTVFITNDKRRTTQAMMTWNITCTYSQFLAQKAPSCCVSLSSFYNDTVVNCPTCT 181
Query: 243 CGCQS----GNCVDPNSPNLTSVISNPGNNGH 270
CGC++ G+CVDPNSP+L SV+S G +
Sbjct: 182 CGCRNKTEPGSCVDPNSPHLASVVSASGKTAN 277
Score = 92.4 bits (228), Expect(2) = 5e-54
Identities = 60/140 (42%), Positives = 85/140 (59%)
Frame = +1
Query: 275 PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQLRVI 334
P++ S+LACEA+LQ VL + ++ FQLQ+EL +E R PAL F L+V
Sbjct: 301 PYVSYSSALACEAQLQRVLEGQDHNHKFQLQNELLSMEPCCAAS*FR*YYPALQF*LQVT 480
Query: 335 NSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSEDLFQ 394
SL G+ E*Y A+G+ VLQ*F+ SW SW C+VR K +N + * +G ++LFQ
Sbjct: 481 KSL*GL-E*YKYAVGSQVLQ*FSFFSWISW*CSVRNLT*KRQINLHI**GMGLP*KNLFQ 657
Query: 395 W*QLCDATT*CLSKVTECRF 414
W LC A++ CL ++C+F
Sbjct: 658 WG*LCYASSRCLPLASKCKF 717
>AI416924
Length = 324
Score = 181 bits (458), Expect = 7e-46
Identities = 80/109 (73%), Positives = 95/109 (86%)
Frame = +1
Query: 90 TEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQDPANAVASF 149
TEQGDCSKFKG + PHCCKKDP VDL+PG PY++Q+ANCCKGGVL+S QDP NAV+SF
Sbjct: 1 TEQGDCSKFKGGI-PHCCKKDPTVVDLLPGTPYNQQIANCCKGGVLSSWAQDPTNAVSSF 177
Query: 150 RVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRR 198
+V VGRAGTT K+V++PKNFT K+PGPGYTCGPA+IV PT+F+ DKRR
Sbjct: 178 QVSVGRAGTTNKTVKVPKNFTLKAPGPGYTCGPAKIVAPTKFITSDKRR 324
>TC234872 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (21%)
Length = 427
Score = 171 bits (434), Expect = 5e-43
Identities = 74/114 (64%), Positives = 90/114 (78%)
Frame = +1
Query: 5 LLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHI 64
+LF +L CT +S++AYDPLDPNGNITI+WD++SWT DGYVA VT+NNF +RHI
Sbjct: 100 ILFLVLLSCTCF----TSTDAYDPLDPNGNITIKWDVISWTPDGYVAVVTMNNFLAFRHI 267
Query: 65 QEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIP 118
PGWS+ WTWAK E+IW M+GGQ TEQGDCSKFKG + PH CKK+P VDL+P
Sbjct: 268 PSPGWSMRWTWAKKEVIWNMVGGQATEQGDCSKFKGNI-PHSCKKNPTVVDLLP 426
>TC228819 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (31%)
Length = 460
Score = 82.8 bits (203), Expect(2) = 3e-32
Identities = 52/91 (57%), Positives = 64/91 (70%)
Frame = +2
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
M+* ++P+ LAC A LQ VLA EG Y* LQDELF*+E T KL QSD A+ FQL
Sbjct: 188 MY*TYVPSQYPLAC*A*LQGVLACEGYYY*L*LQDELF*MEHGCSTSKL*QSDSAVQFQL 367
Query: 332 RVINSLWGVNE*YSNALGN*VLQ*FAHASWP 362
+VI+SLW N+*Y NA+G+*VL * + SWP
Sbjct: 368 QVIDSLW-FNK*YCNAMGS*VL**LSKPSWP 457
Score = 74.3 bits (181), Expect(2) = 3e-32
Identities = 33/55 (60%), Positives = 41/55 (74%), Gaps = 5/55 (9%)
Frame = +3
Query: 219 KAPTCCVSLSTFYNDTIVSCPTCACGC-----QSGNCVDPNSPNLTSVISNPGNN 268
K P+C VSLS+FYNDT+V C TCACGC QSG CVDP++P+L SV++ G N
Sbjct: 3 KTPSCGVSLSSFYNDTLVPCLTCACGCQSNSSQSGTCVDPDTPHLASVVAGSGKN 167
>TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (29%)
Length = 617
Score = 133 bits (334), Expect = 2e-31
Identities = 82/137 (59%), Positives = 98/137 (70%), Gaps = 1/137 (0%)
Frame = +2
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
M+* ++PNPSSLA AKLQ +LA EG Y*F LQDELF*LEL+ TP L+QSD ++ FQL
Sbjct: 2 MY*AYVPNPSSLARYAKLQEILACEGYCY*F*LQDELF*LELACSTP*LQQSDSSVWFQL 181
Query: 332 RVINSLWGVNE*YSNALGN*VLQ-*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSE 390
+V + +*YSNA+GN*V * + SWP C+ R KG* NFYF * LGFSS+
Sbjct: 182 QV-TGFGCIYK*YSNAVGN*VPS**HSQPSWP*RQCSSRATLQKG*SNFYF**GLGFSSK 358
Query: 391 DLFQW*QLCDATT*CLS 407
DL QW* +CDATT*CLS
Sbjct: 359 DLLQW*HMCDATT*CLS 409
>BG789974 similar to GP|21553837|gb| phytochelatin synthetase-like protein
{Arabidopsis thaliana}, partial (14%)
Length = 438
Score = 129 bits (323), Expect = 3e-30
Identities = 56/79 (70%), Positives = 64/79 (80%), Gaps = 5/79 (6%)
Frame = +3
Query: 12 PCTLLLLLPS-----SSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
PC L L L S S++AYDPLDPNGNITI+WDI+SWT DGYVA VT+NNFQQYRHI
Sbjct: 201 PCILFLALLSCTCFTSTDAYDPLDPNGNITIKWDIISWTPDGYVAVVTMNNFQQYRHIAS 380
Query: 67 PGWSLGWTWAKNEIIWQMI 85
PGWS+GWTWAK E+IW M+
Sbjct: 381 PGWSMGWTWAKKEVIWSMM 437
>AI416873
Length = 379
Score = 107 bits (268), Expect = 8e-24
Identities = 44/59 (74%), Positives = 50/59 (84%)
Frame = +2
Query: 51 AAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKK 109
A VT++NFQ +RHI PGW+LGWTWAK E+IW MIG QTTEQGDCSKFKG + PHCCKK
Sbjct: 203 AVVTMHNFQMFRHIMNPGWTLGWTWAKKEVIWSMIGAQTTEQGDCSKFKGNI-PHCCKK 376
>TC228818 similar to UP|CBL1_ARATH (Q9SRT7) COBRA-like protein 1 precursor,
partial (26%)
Length = 736
Score = 107 bits (266), Expect = 1e-23
Identities = 69/124 (55%), Positives = 84/124 (67%)
Frame = +2
Query: 308 LF*LELSNRTPKLRQSDPAL*FQLRVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCT 367
LF*+E KL QSD + FQL+VI+SLW N+*Y NA+G+*VL *F+ SWP W C+
Sbjct: 41 LF*MEHGCSXCKL*QSDSTVQFQLQVIDSLW-FNK*YCNAMGS*VL**FSKPSWP*WQCS 217
Query: 368 VRPAFWKG*VNFYFS*RLGFSSEDLFQW*QLCDATT*CLSKVTECRFAAKGFPLTCFDVN 427
+R KG* NFYF * LGFSS+ L Q *QLC+ATT* LS VT C A+GF L F +
Sbjct: 218 IRATLPKG*SNFYF**GLGFSSKGLLQR*QLCNATT*FLSVVT*CWC*ARGF-LVFFSDS 394
Query: 428 ILGN 431
I G+
Sbjct: 395 IFGS 406
>TC217722 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor,
partial (47%)
Length = 905
Score = 62.8 bits (151), Expect(2) = 3e-22
Identities = 28/46 (60%), Positives = 33/46 (70%), Gaps = 1/46 (2%)
Frame = +1
Query: 217 AAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-GNCVDPNSPNLTSV 261
A K P CCVSLS+FYN+TI CPTCACGCQ+ NCV +S + V
Sbjct: 16 ARKNPGCCVSLSSFYNETITPCPTCACGCQNRRNCVKSDSKRINMV 153
Score = 60.5 bits (145), Expect(2) = 3e-22
Identities = 47/133 (35%), Positives = 73/133 (54%)
Frame = +3
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
+H ++ N +LA EA+L +LAS+G FQLQDEL L+ K Q P+ * L
Sbjct: 195 VHSSYVSNKGALAREAQL*GLLASQGCCDEFQLQDELLPLDSCCSASKS*QCHPSF*L*L 374
Query: 332 RVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSED 391
+ SL V++*+ L + +L * + SW +W C++R + + N + R+G SS+
Sbjct: 375 QAFTSL-*VHK*HWYVLWHEIL**SLNGSWTNWECSIRNSSSEEPGNIHIQARMGISSQG 551
Query: 392 LFQW*QLCDATT* 404
L W* + AT+*
Sbjct: 552 LL*W*GMHAATS* 590
>TC219820 similar to GB|AAO64838.1|29028918|BT005903 At4g16120 {Arabidopsis
thaliana;} , partial (37%)
Length = 616
Score = 73.9 bits (180), Expect = 1e-13
Identities = 43/147 (29%), Positives = 70/147 (47%), Gaps = 8/147 (5%)
Frame = +2
Query: 107 CKKDPVAVDLIPGAPYSE----QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKS 162
C++ P +DL P +++ ++ CC+ G + DP+ + + F++ V +
Sbjct: 47 CERRPTIIDL-PPTKFNDSDLGKIPFCCRNGTILPPSMDPSMSASRFQMQVFKMPPALNR 223
Query: 163 VRLPKNFTFKSPG---PGYTCGPARIVKPTRFLRYDKRRVTQAMM-SWNVTCTYSQFLAA 218
+L +K G P Y CGP V PT + +M SW V C +
Sbjct: 224 SQLSPPQNWKISGTLNPDYECGPPVRVSPTENPDPSGLPSNKTVMASWQVVCNITTAKRT 403
Query: 219 KAPTCCVSLSTFYNDTIVSCPTCACGC 245
+ CCVS S++YND+++ C TCACGC
Sbjct: 404 SSK-CCVSFSSYYNDSVIPCKTCACGC 481
>TC233889 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor,
partial (31%)
Length = 531
Score = 66.6 bits (161), Expect = 2e-11
Identities = 49/124 (39%), Positives = 70/124 (55%)
Frame = +1
Query: 283 LACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQLRVINSLWGVNE 342
LACE L +LAS+G QLQ+E L K +QS + FQL+ +SLW +++
Sbjct: 7 LACED*L*GLLASQGCHNKLQLQNESLPLVSCCSASKSQQSHTSFQFQLQATSSLW-IHK 183
Query: 343 *YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSEDLFQW*QLCDAT 402
*Y + L + VLQ* ++ SW +W C++R A +G + LG SS+ L W* + AT
Sbjct: 184 *YWHVLWHEVLQ*SSNGSWTNWECSIRVASSEGQGCIHIQAGLGISSQGLL*W**MHAAT 363
Query: 403 T*CL 406
T* L
Sbjct: 364 T*HL 375
>TC216060 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (22%)
Length = 661
Score = 65.9 bits (159), Expect = 3e-11
Identities = 41/90 (45%), Positives = 57/90 (62%)
Frame = +3
Query: 325 PAL*FQLRVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*R 384
PAL F L+V +L G+ E*Y A+G+ VLQ*F+ SW SW C+VR K +N + *
Sbjct: 24 PALQF*LQVTYTL*GL-E*YKYAVGSQVLQ*FSFFSWISW*CSVRNLT*KRQINLHI**G 200
Query: 385 LGFSSEDLFQW*QLCDATT*CLSKVTECRF 414
+G +DLFQW LC A++ CL ++C+F
Sbjct: 201 MGLP*KDLFQWG*LCYASSRCLPLASKCKF 290
>AW703859 homologue to GP|26452135|dbj GPI-anchored protein {Arabidopsis
thaliana}, partial (7%)
Length = 366
Score = 57.8 bits (138), Expect = 9e-09
Identities = 23/29 (79%), Positives = 27/29 (92%)
Frame = +2
Query: 204 MSWNVTCTYSQFLAAKAPTCCVSLSTFYN 232
++WNVTCTYSQFLA K P+CCVSLS+FYN
Sbjct: 278 VTWNVTCTYSQFLAQKTPSCCVSLSSFYN 364
>BE610073
Length = 423
Score = 29.3 bits (64), Expect = 3.6
Identities = 19/79 (24%), Positives = 33/79 (41%), Gaps = 1/79 (1%)
Frame = -2
Query: 54 TLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKD-PV 112
T+ N+ QY ++ W W +NE++ + + T+ C K CCK P+
Sbjct: 374 TILNYPQYLQRKQELWQT--CWCQNEVLTSQLKQELTDHPACGK---HFYILCCKHSRPL 210
Query: 113 AVDLIPGAPYSEQVANCCK 131
+ P ++ NC K
Sbjct: 209 VRKVFPCIDSQIKLKNCIK 153
>AY377863
Length = 449
Score = 28.9 bits (63), Expect = 4.7
Identities = 33/107 (30%), Positives = 52/107 (47%)
Frame = -2
Query: 325 PAL*FQLRVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*R 384
P+L* + +SL +++*+ + L +LQ* S C V A +G + Y
Sbjct: 424 PSL*LWIHPASSL*-IHK*HRHVLWFKILQ*SVDGSRA*RECAV*GAHEEGQEHIYSQAG 248
Query: 385 LGFSSEDLFQW*QLCDATT*CLSKVTECRFAAKGFPLTCFDVNILGN 431
+G S E L QW* + +T+* +S + AK +P T + I GN
Sbjct: 247 MGISKEGLLQW**MYASTS*FIS------YVAKLWP*TTNNYYIDGN 125
>TC230380 similar to GB|AAO64866.1|29028974|BT005931 At4g29000 {Arabidopsis
thaliana;} , partial (52%)
Length = 1463
Score = 28.5 bits (62), Expect = 6.2
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +1
Query: 153 VGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARI 185
VGR G +VR+P + SP PG GPA +
Sbjct: 607 VGRQGQQANNVRVPAPSSSLSPIPGARVGPATL 705
>BE346606 similar to GP|13605910|gb| AT3g55400/T22E16_60 {Arabidopsis
thaliana}, partial (4%)
Length = 418
Score = 28.5 bits (62), Expect = 6.2
Identities = 15/48 (31%), Positives = 23/48 (47%), Gaps = 6/48 (12%)
Frame = -1
Query: 223 CCVSLSTFYNDTIVSCPTCA------CGCQSGNCVDPNSPNLTSVISN 264
CC+ L + CP+ A C C + +CV NS L+S +S+
Sbjct: 268 CCIKLVVGIXKLCIYCPSQAWCNRKECNCNNPHCV*YNSKVLSSCLSS 125
>BG839237 similar to GP|3688193|emb MAP3K alpha 1 protein kinase {Brassica
napus}, partial (17%)
Length = 777
Score = 28.5 bits (62), Expect = 6.2
Identities = 17/53 (32%), Positives = 24/53 (45%)
Frame = +3
Query: 136 TSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKP 188
+S+ P + + FR+ T +RL N KSPGPG + GP P
Sbjct: 69 SSVTPPPPSTIIQFRLPTPTPTPTEDMMRLXVNVRSKSPGPG-SRGPTSPTSP 224
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,841,397
Number of Sequences: 63676
Number of extensions: 425588
Number of successful extensions: 3383
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 3324
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3361
length of query: 434
length of database: 12,639,632
effective HSP length: 100
effective length of query: 334
effective length of database: 6,272,032
effective search space: 2094858688
effective search space used: 2094858688
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0338.9


