

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
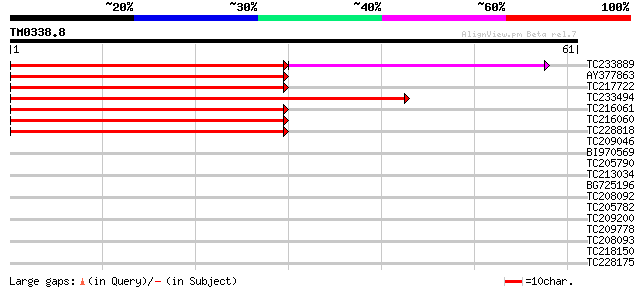
Score E
Sequences producing significant alignments: (bits) Value
TC233889 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 ... 67 5e-13
AY377863 67 2e-12
TC217722 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 ... 63 3e-11
TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 62 6e-11
TC216061 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 61 1e-10
TC216060 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 61 1e-10
TC228818 similar to UP|CBL1_ARATH (Q9SRT7) COBRA-like protein 1 ... 60 2e-10
TC209046 similar to UP|CBL8_ARATH (Q9LIB6) COBRA-like protein 8 ... 32 0.050
BI970569 32 0.066
TC205790 weakly similar to UP|Q8W162 (Q8W162) Methylthioadenosin... 31 0.11
TC213034 28 0.95
BG725196 27 2.1
TC208092 27 2.8
TC205782 similar to UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, pa... 26 4.7
TC209200 26 4.7
TC209778 similar to UP|Q88GI3 (Q88GI3) Major facilitator family ... 25 6.2
TC208093 weakly similar to UP|Q9LQS6 (Q9LQS6) T4O12.12, partial ... 25 6.2
TC218150 similar to UP|Q70I27 (Q70I27) SAR DNA-binding protein-l... 25 6.2
TC228175 similar to UP|Q9LTG1 (Q9LTG1) Gb|AAD38661.1, partial (44%) 25 8.0
>TC233889 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor,
partial (31%)
Length = 531
Score = 66.6 bits (161), Expect(2) = 5e-13
Identities = 26/30 (86%), Positives = 29/30 (96%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+ F+FKQGWAFPRKVYFNGDECM+PPP
Sbjct: 278 KDKDAFTFKQGWAFPRKVYFNGDECMLPPP 367
Score = 22.3 bits (46), Expect(2) = 5e-13
Identities = 14/28 (50%), Positives = 15/28 (53%)
Frame = +3
Query: 31 TPTHFCQILLLQA*FPSRHSSSHYSSFL 58
TPT F LLLQA S S + SS L
Sbjct: 369 TPTLFSLTLLLQAYLTSLRSCCYCSSCL 452
>AY377863
Length = 449
Score = 67.0 bits (162), Expect = 2e-12
Identities = 26/30 (86%), Positives = 29/30 (96%)
Frame = -3
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDKNTF+ KQGWAFPR+VYFNGDECM+PPP
Sbjct: 279 KDKNTFTLKQGWAFPRRVYFNGDECMLPPP 190
>TC217722 similar to UP|CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor,
partial (47%)
Length = 905
Score = 63.2 bits (152), Expect = 3e-11
Identities = 24/30 (80%), Positives = 29/30 (96%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K++ TF+FKQGWAFPRKVYFNG+ECM+PPP
Sbjct: 499 KNQETFTFKQGWAFPRKVYFNGEECMLPPP 588
>TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (29%)
Length = 617
Score = 62.0 bits (149), Expect = 6e-11
Identities = 26/43 (60%), Positives = 33/43 (76%)
Frame = +3
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA 43
KDK TF+F +GWAFPR++YFNGD C+MPPP F I+ LQ+
Sbjct: 309 KDKATFTFDKGWAFPRRIYFNGDICVMPPPDAYPF*TIIRLQS 437
>TC216061 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (52%)
Length = 1100
Score = 60.8 bits (146), Expect = 1e-10
Identities = 22/30 (73%), Positives = 28/30 (93%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 596 KDKSTFTFDKGWAFPRRIYFNGDNCVMPPP 685
>TC216060 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (22%)
Length = 661
Score = 60.8 bits (146), Expect = 1e-10
Identities = 22/30 (73%), Positives = 28/30 (93%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 169 KDKSTFTFDKGWAFPRRIYFNGDNCVMPPP 258
>TC228818 similar to UP|CBL1_ARATH (Q9SRT7) COBRA-like protein 1 precursor,
partial (26%)
Length = 736
Score = 60.1 bits (144), Expect = 2e-10
Identities = 23/30 (76%), Positives = 27/30 (89%)
Frame = +3
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK TF+F +GWAFPR+VYFNGD C+MPPP
Sbjct: 237 KDKATFTFDKGWAFPRRVYFNGDNCVMPPP 326
>TC209046 similar to UP|CBL8_ARATH (Q9LIB6) COBRA-like protein 8 precursor,
partial (21%)
Length = 756
Score = 32.3 bits (72), Expect = 0.050
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +1
Query: 10 QGWAFPRKVYFNGDECMMPPPTPT 33
+G FP KV+FNG+EC +P P+
Sbjct: 349 RGDGFPTKVFFNGEECSLPSVYPS 420
>BI970569
Length = 788
Score = 32.0 bits (71), Expect = 0.066
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = -1
Query: 11 GWAFPRKVYFNGDECMMPPPTPT 33
G FP KV+FNG+EC +P P+
Sbjct: 455 GDGFPTKVFFNGEECSLPSVYPS 387
>TC205790 weakly similar to UP|Q8W162 (Q8W162) Methylthioadenosine/S-adenosyl
homocysteine nucleosidase , partial (71%)
Length = 960
Score = 31.2 bits (69), Expect = 0.11
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +1
Query: 14 FPRKVYFNGDECMMPPP 30
+PR++YFNG+ C MP P
Sbjct: 757 YPRRIYFNGESCEMP*P 807
>TC213034
Length = 442
Score = 28.1 bits (61), Expect = 0.95
Identities = 15/46 (32%), Positives = 21/46 (45%)
Frame = +1
Query: 15 PRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSHYSSFLFY 60
P K +FN PPP+ +H C + PS + S SF F+
Sbjct: 202 PSKTFFNNSSFHFPPPSSSHTCLLHFAPKASPSGNFSGD-DSFGFF 336
>BG725196
Length = 388
Score = 26.9 bits (58), Expect = 2.1
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 8/37 (21%)
Frame = +3
Query: 32 PTHF--------CQILLLQA*FPSRHSSSHYSSFLFY 60
P+HF C IL FP+ HS++ Y F FY
Sbjct: 54 PSHFTWLFFFSSCFILFSSRRFPTSHSANFYLIFFFY 164
>TC208092
Length = 513
Score = 26.6 bits (57), Expect = 2.8
Identities = 11/35 (31%), Positives = 17/35 (48%)
Frame = -2
Query: 23 DECMMPPPTPTHFCQILLLQA*FPSRHSSSHYSSF 57
D C FC++LLL +P RH H++ +
Sbjct: 245 DLCTFQEHNLRAFCKLLLLDDNYPKRHIPKHWNKW 141
>TC205782 similar to UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, partial (91%)
Length = 1377
Score = 25.8 bits (55), Expect = 4.7
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -3
Query: 28 PPPTPTHFCQILLLQA*FPSRHSSSH 53
P P CQ+LLL+ FP + +S H
Sbjct: 592 PQNLPRQICQLLLLRDVFPLQRTSVH 515
>TC209200
Length = 322
Score = 25.8 bits (55), Expect = 4.7
Identities = 7/11 (63%), Positives = 10/11 (90%)
Frame = -2
Query: 26 MMPPPTPTHFC 36
++PPP+P HFC
Sbjct: 216 LLPPPSPLHFC 184
>TC209778 similar to UP|Q88GI3 (Q88GI3) Major facilitator family transporter,
partial (4%)
Length = 837
Score = 25.4 bits (54), Expect = 6.2
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 30 PTPTHFCQILLLQA*FPSRHSSSHYSSFLFYL 61
P PTH LL+ P+ H+ S ++FL+ L
Sbjct: 463 PIPTHQHATLLMLLQIPTHHAVSTSTTFLYLL 368
>TC208093 weakly similar to UP|Q9LQS6 (Q9LQS6) T4O12.12, partial (17%)
Length = 622
Score = 25.4 bits (54), Expect = 6.2
Identities = 11/33 (33%), Positives = 15/33 (45%)
Frame = -3
Query: 23 DECMMPPPTPTHFCQILLLQA*FPSRHSSSHYS 55
D C FC +LLL +P RH H++
Sbjct: 413 DLCTFQEHDLRSFCTLLLLDDNYPKRHIPKHWN 315
>TC218150 similar to UP|Q70I27 (Q70I27) SAR DNA-binding protein-like protein,
partial (11%)
Length = 463
Score = 25.4 bits (54), Expect = 6.2
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 2/33 (6%)
Frame = -1
Query: 30 PTPTHFC--QILLLQA*FPSRHSSSHYSSFLFY 60
P + FC L Q PSRH+S +SSFL +
Sbjct: 259 PHFSFFCLSSCLAFQEILPSRHASFSFSSFLLH 161
>TC228175 similar to UP|Q9LTG1 (Q9LTG1) Gb|AAD38661.1, partial (44%)
Length = 1122
Score = 25.0 bits (53), Expect = 8.0
Identities = 12/36 (33%), Positives = 18/36 (49%)
Frame = -2
Query: 12 WAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPS 47
W F +K+++ D P P H+ + QA FPS
Sbjct: 380 WPFLQKLHYGMDFLSAPWQHPRHYSSQVHKQAMFPS 273
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,842,040
Number of Sequences: 63676
Number of extensions: 86432
Number of successful extensions: 658
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 656
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 658
length of query: 61
length of database: 12,639,632
effective HSP length: 37
effective length of query: 24
effective length of database: 10,283,620
effective search space: 246806880
effective search space used: 246806880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0338.8


