

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.4
(197 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
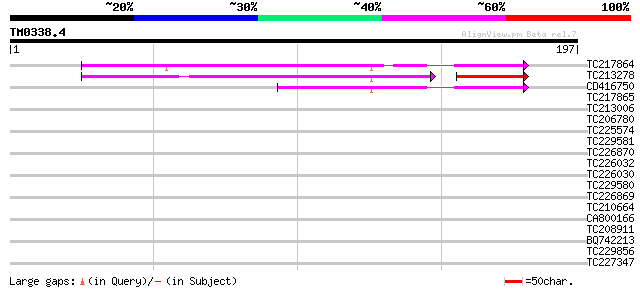
Score E
Sequences producing significant alignments: (bits) Value
TC217864 similar to GB|AAP12861.1|30017255|BT006212 At2g40095 {A... 119 1e-27
TC213278 weakly similar to GB|AAP12861.1|30017255|BT006212 At2g4... 100 5e-26
CD416750 63 1e-10
TC217865 35 0.017
TC213006 similar to UP|Q9KJW7 (Q9KJW7) YajC homolog, partial (11%) 29 1.6
TC206780 similar to UP|2AAA_PEA (P36875) Protein phosphatase PP2... 29 1.6
TC225574 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein... 28 2.1
TC229581 similar to UP|Q94F37 (Q94F37) At1g67850/F12A21_2, parti... 28 2.1
TC226870 28 2.7
TC226032 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, ... 28 2.7
TC226030 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, ... 28 2.7
TC229580 similar to UP|Q94F37 (Q94F37) At1g67850/F12A21_2, parti... 28 3.6
TC226869 similar to UP|Q8H6S5 (Q8H6S5) CTV.2, partial (40%) 28 3.6
TC210664 28 3.6
CA800166 27 4.7
TC208911 similar to UP|Q9SGD8 (Q9SGD8) T23G18.8, partial (39%) 27 4.7
BQ742213 27 8.0
TC229856 similar to UP|Q7Z615 (Q7Z615) Specificity protein 8 sho... 27 8.0
TC227347 similar to UP|O04097 (O04097) BRCA1-associated RING dom... 27 8.0
>TC217864 similar to GB|AAP12861.1|30017255|BT006212 At2g40095 {Arabidopsis
thaliana;} , partial (59%)
Length = 1220
Score = 119 bits (297), Expect = 1e-27
Identities = 66/158 (41%), Positives = 94/158 (58%), Gaps = 3/158 (1%)
Frame = +1
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFT--CIRCYGITKHFGRYAFKSSLTDIPLVSLMR 83
+ L + S PV++V+ D LVP L+S + + HF Y F+ SL DIPLVS++R
Sbjct: 406 ISLLLFSAPVLLVIADALVPSALLSTLSPSSFSLETLASHFHNYDFRYSLIDIPLVSIIR 585
Query: 84 SLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTC-VFTVNSRIEAEASASLKR 142
S II CVYS+CD P LS GPYLG T+CS+LS++ +S K +F+V+ + S ++
Sbjct: 586 SFIIFCVYSLCDGPRLSRGPYLGITTMCSVLSLMFVSFKAVYIFSVSG---IDRSGYVRA 756
Query: 143 QRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
+ LF+ S ++GH+VVA RTSCR RR
Sbjct: 757 TEI---------ALFVCSCALAVGHVVVAYRTSCRERR 843
>TC213278 weakly similar to GB|AAP12861.1|30017255|BT006212 At2g40095
{Arabidopsis thaliana;} , partial (75%)
Length = 788
Score = 100 bits (248), Expect(2) = 5e-26
Identities = 52/124 (41%), Positives = 75/124 (59%), Gaps = 1/124 (0%)
Frame = +3
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSL 85
+ L + S PV++V+ D LVP L+S F+ + HF Y F SL DIPLVS+ RSL
Sbjct: 201 ISLLLFSAPVLLVIADALVPSALLSAFSPSFLFS---HFENYDFGYSLIDIPLVSIARSL 371
Query: 86 IILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTC-VFTVNSRIEAEASASLKRQR 144
+I CVYS CD P LSHGPYLG LCS++S+ + +K V ++N+ + S + +
Sbjct: 372 VIFCVYSFCDRPRLSHGPYLGVTMLCSVMSLTFVCLKAVYVLSLNTNVSGRESGYDRSSQ 551
Query: 145 LHLK 148
+ L+
Sbjct: 552 IALR 563
Score = 34.3 bits (77), Expect(2) = 5e-26
Identities = 15/25 (60%), Positives = 19/25 (76%)
Frame = +1
Query: 156 LFLSSVVFSLGHIVVACRTSCRARR 180
LF+ S ++GH+VVA RTSCR RR
Sbjct: 556 LFVWSCALAVGHVVVAYRTSCRERR 630
>CD416750
Length = 723
Score = 62.8 bits (151), Expect = 1e-10
Identities = 33/88 (37%), Positives = 49/88 (55%), Gaps = 1/88 (1%)
Frame = -1
Query: 94 CDAPTLSHGPYLGTVTLCSILSIVLLSVKTC-VFTVNSRIEAEASASLKRQRLHLKKSWG 152
CD P LSHGP LG LCS++S+ + +K V ++N+ + S + ++
Sbjct: 720 CDRPRLSHGPNLGVTMLCSVMSLTFVCLKAVYVLSLNTNVSGRESGYDRSSQI------- 562
Query: 153 MPVLFLSSVVFSLGHIVVACRTSCRARR 180
LF+ S ++GH+VVA RTSCR RR
Sbjct: 561 --ALFVWSCALAVGHVVVAYRTSCRERR 484
>TC217865
Length = 557
Score = 35.4 bits (80), Expect = 0.017
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Frame = +3
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFT--CIRCYGITKHFGRYAFKSSLTDIPLVSL 81
+ L + S PV++V+ D LVP L S+ + + HF Y F++ DIPL +L
Sbjct: 285 ISLLLFSAPVLLVIADALVPSALRSSLSQPSFSMETLASHFHNYDFRNKNIDIPLGAL 458
>TC213006 similar to UP|Q9KJW7 (Q9KJW7) YajC homolog, partial (11%)
Length = 494
Score = 28.9 bits (63), Expect = 1.6
Identities = 21/50 (42%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Frame = -3
Query: 109 TLCSI-----LSIVLLSVKT--CVFTVNSRIEAEASASLKRQRLHLKKSW 151
T CSI I LL V T C+FTV+ EAEA A+ R H + +W
Sbjct: 264 TKCSISRQELFIIHLLQVATGYCIFTVHV*HEAEAEAANLLARFHRQTNW 115
>TC206780 similar to UP|2AAA_PEA (P36875) Protein phosphatase PP2A regulatory
subunit A (PR65) (Fragment), partial (31%)
Length = 667
Score = 28.9 bits (63), Expect = 1.6
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 1/50 (2%)
Frame = +2
Query: 64 FGRYAFKSSLTDIPLVSLMRSLIILCV-YSICDAPTLSHGPYLGTVTLCS 112
F + F T ++ + + ILC+ Y+ CD LS PYL LCS
Sbjct: 392 FSYFFFLIRFTFSSVIDMTPTCPILCITYNCCDPALLSFYPYLCYRVLCS 541
>TC225574 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein, partial
(83%)
Length = 868
Score = 28.5 bits (62), Expect = 2.1
Identities = 15/30 (50%), Positives = 18/30 (60%)
Frame = +1
Query: 20 CTTRSKLDLTVSSLPVVVVVVDLLVPCVLI 49
CTTR +SLP+VV VV LL VL+
Sbjct: 79 CTTRMSQGRGSASLPIVVTVVSLLCLLVLL 168
>TC229581 similar to UP|Q94F37 (Q94F37) At1g67850/F12A21_2, partial (56%)
Length = 958
Score = 28.5 bits (62), Expect = 2.1
Identities = 12/22 (54%), Positives = 16/22 (72%)
Frame = -3
Query: 86 IILCVYSICDAPTLSHGPYLGT 107
+++ V+SIC PTL HGP L T
Sbjct: 698 VVIRVFSICQ-PTLEHGPLLST 636
>TC226870
Length = 882
Score = 28.1 bits (61), Expect = 2.7
Identities = 19/50 (38%), Positives = 25/50 (50%), Gaps = 5/50 (10%)
Frame = -2
Query: 145 LHLKKSWG---MPVLFLSSVVFS--LGHIVVACRTSCRARRSSCFIESTL 189
LHL S G MP+L S + + L H VAC +R +CFI S +
Sbjct: 179 LHLNLSAGTVRMPILPSSKLTYID*LSHEYVACVIGLASRGENCFIHSNV 30
>TC226032 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, partial
(41%)
Length = 612
Score = 28.1 bits (61), Expect = 2.7
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = +2
Query: 80 SLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVL 118
S MR +++ + + PT+ + PYL +L S LS+ L
Sbjct: 251 SAMRHMLVEVMQMMLSHPTIPNSPYLDEESLSSALSLTL 367
>TC226030 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, complete
Length = 651
Score = 28.1 bits (61), Expect = 2.7
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = +3
Query: 80 SLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVL 118
S MR +++ + + PT+ + PYL +L S LS+ L
Sbjct: 411 SAMRHMLVEVMQMMLSHPTIPNSPYLDEESLSSALSLTL 527
>TC229580 similar to UP|Q94F37 (Q94F37) At1g67850/F12A21_2, partial (19%)
Length = 479
Score = 27.7 bits (60), Expect = 3.6
Identities = 17/44 (38%), Positives = 27/44 (60%), Gaps = 5/44 (11%)
Frame = -2
Query: 69 FKSSLTDIPLVSL-MRSL----IILCVYSICDAPTLSHGPYLGT 107
+KS+L + L+S +R + +++ V+ IC PTL HGP L T
Sbjct: 295 YKSTLALVTLMSCGIRCISSHGLVIRVFGICQ-PTLEHGPLLST 167
>TC226869 similar to UP|Q8H6S5 (Q8H6S5) CTV.2, partial (40%)
Length = 1800
Score = 27.7 bits (60), Expect = 3.6
Identities = 19/48 (39%), Positives = 24/48 (49%), Gaps = 5/48 (10%)
Frame = -1
Query: 145 LHLKKSWG---MPVLFLSSVVFS--LGHIVVACRTSCRARRSSCFIES 187
LHL S G MP+L S + + L H VAC +R +CFI S
Sbjct: 1287 LHLNLSAGTVRMPILPSSKLTYID*LSHEYVACVIGLASRGENCFIHS 1144
>TC210664
Length = 891
Score = 27.7 bits (60), Expect = 3.6
Identities = 16/50 (32%), Positives = 26/50 (52%)
Frame = -3
Query: 85 LIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEA 134
LII+C+ IC A ++ G G S +++ S V TV S++E+
Sbjct: 217 LIIICLTKICSAASIIAGWQTGLQHSLSGSTVLQASSNLLVLTVFSKVES 68
>CA800166
Length = 437
Score = 27.3 bits (59), Expect = 4.7
Identities = 25/87 (28%), Positives = 40/87 (45%), Gaps = 6/87 (6%)
Frame = +1
Query: 89 CVYSICDAPTLSH--GPYLGTVTLCSILSIVLLSVKTCV---FTVNSRIEAEASASLKRQ 143
CV S+ LS G VT+C + +VLL + C+ F SRI ++ L
Sbjct: 118 CVESVHRVMPLSRFAAEAFGVVTICLVAILVLLGL-MCIAYSFYFRSRIHSQGLVQL--- 285
Query: 144 RLHLKKSWGMPVLF-LSSVVFSLGHIV 169
L+ W + + F L S+ + +G I+
Sbjct: 286 -LYFSGPWIIRITFILFSIWWGIGEII 363
>TC208911 similar to UP|Q9SGD8 (Q9SGD8) T23G18.8, partial (39%)
Length = 971
Score = 27.3 bits (59), Expect = 4.7
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -2
Query: 141 KRQRLHLKKSWGMPVLFLSSVVFSLG 166
KR+ + KKSWG + + SS+ F LG
Sbjct: 970 KRKDKYTKKSWGETIFYESSLGFFLG 893
>BQ742213
Length = 427
Score = 26.6 bits (57), Expect = 8.0
Identities = 21/82 (25%), Positives = 36/82 (43%)
Frame = -2
Query: 19 SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPL 78
S + ++L L SS + V L P + + +FT R T H GR + + + L
Sbjct: 348 STFSETRLLLPFSSSGAIQGRVPLTPPDIKVWHFTLDRPKSPTLHIGRCGSRRFTSKLSL 169
Query: 79 VSLMRSLIILCVYSICDAPTLS 100
++ C YSI A +++
Sbjct: 168 FKSK*TIFFPCRYSIPKATSIA 103
>TC229856 similar to UP|Q7Z615 (Q7Z615) Specificity protein 8 short isoform,
partial (5%)
Length = 821
Score = 26.6 bits (57), Expect = 8.0
Identities = 15/45 (33%), Positives = 24/45 (53%), Gaps = 1/45 (2%)
Frame = -1
Query: 76 IPLVSLMRSLIILCVYSICDAP-TLSHGPYLGTVTLCSILSIVLL 119
+ V L L+I C++S C+A L+H P++ IL I +L
Sbjct: 761 VEFVQLFTLLMITCIHSRCEASLCLAHPPFIAHFQNLFILVIQVL 627
>TC227347 similar to UP|O04097 (O04097) BRCA1-associated RING domain protein
isolog; 106935-111081, partial (17%)
Length = 982
Score = 26.6 bits (57), Expect = 8.0
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 3/30 (10%)
Frame = -2
Query: 69 FKSSLTDIPLVSLMRSLIILCVYSI---CD 95
F++ L + S++R +IILCVY I CD
Sbjct: 465 FRTMLMPFGISSMVRLIIILCVYQI*C*CD 376
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.343 0.149 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,342,740
Number of Sequences: 63676
Number of extensions: 167426
Number of successful extensions: 2064
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 2052
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2062
length of query: 197
length of database: 12,639,632
effective HSP length: 92
effective length of query: 105
effective length of database: 6,781,440
effective search space: 712051200
effective search space used: 712051200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0338.4


