

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.2
(282 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
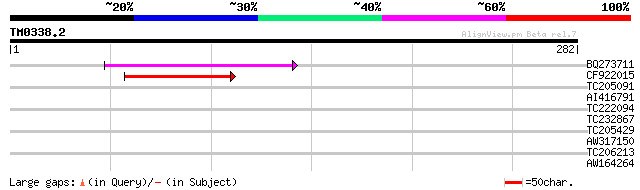
Score E
Sequences producing significant alignments: (bits) Value
BQ273711 84 7e-17
CF922015 60 1e-09
TC205091 weakly similar to UP|Q6TXI7 (Q6TXI7) LRRGT00012, partia... 35 0.050
AI416791 33 0.15
TC222094 32 0.25
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 32 0.32
TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich prote... 30 1.6
AW317150 28 4.7
TC206213 similar to UP|PMXA_MOUSE (Q62066) Paired mesoderm homeo... 28 4.7
AW164264 homologue to GP|24210303|em SI:dZ12F11.3 (collagen type... 28 6.1
>BQ273711
Length = 409
Score = 84.0 bits (206), Expect = 7e-17
Identities = 41/96 (42%), Positives = 57/96 (58%)
Frame = -2
Query: 48 QREAALDQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYL 107
+R+ + RGL FR+ PPKF+G DP+ A LW+ E EKIFE + E KV AT++
Sbjct: 288 ERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFM 109
Query: 108 LLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKY 143
L G+AE WW+ + A + WN+F+ FLE Y
Sbjct: 108 LQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 59.7 bits (143), Expect = 1e-09
Identities = 26/55 (47%), Positives = 39/55 (70%)
Frame = -3
Query: 58 GLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDA 112
GL+ F+R +PP F GG DP+ A+ W++EIEKIF V++ + KV AT++L +A
Sbjct: 167 GLDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>TC205091 weakly similar to UP|Q6TXI7 (Q6TXI7) LRRGT00012, partial (5%)
Length = 1360
Score = 34.7 bits (78), Expect = 0.050
Identities = 20/45 (44%), Positives = 24/45 (52%), Gaps = 1/45 (2%)
Frame = +1
Query: 73 GTDPDEADL-WIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWW 116
G D EA L W ++E++F SE KV LAT G A YWW
Sbjct: 1153 GKDNVEAYLDWEMKVEQLFACHHISEERKVPLATLSFQGYALYWW 1287
>AI416791
Length = 420
Score = 33.1 bits (74), Expect = 0.15
Identities = 33/125 (26%), Positives = 54/125 (42%), Gaps = 7/125 (5%)
Frame = -3
Query: 129 EVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVD 188
++ W S ++A LE+Y D E ++GS + +Y + E L ++
Sbjct: 382 DLTWESLKIALLERYGGHGDGDVYEQLTELKQEGS--VEDYITEFEYLIAQIPRLPEKQF 209
Query: 189 EPYMCKRFVRGLRADIEDSVRPLGIM----RFQAL-VEKATEVELM--KNRRMDRAGTGG 241
+ Y F+ GL+++I VR L M R + L V +A E E+ R G
Sbjct: 208 QGY----FLHGLQSEIRGRVRTLATMGEMSRTKLLQVTRAVEKEVKGGNGSGFGRGPKNG 41
Query: 242 PMRSS 246
P RS+
Sbjct: 40 PYRSN 26
>TC222094
Length = 984
Score = 32.3 bits (72), Expect = 0.25
Identities = 20/65 (30%), Positives = 32/65 (48%)
Frame = -2
Query: 103 LATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQG 162
L + L G+A W R +G N++ WN FL+KYFP ES+ + + G
Sbjct: 614 LFCFSLAGEARTWLRSFKG----NNLRT-WNEXXEKFLKKYFP-------ESKTIXXKGG 471
Query: 163 SMTIP 167
++ +P
Sbjct: 470 NLILP 456
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 32.0 bits (71), Expect = 0.32
Identities = 29/121 (23%), Positives = 54/121 (43%), Gaps = 8/121 (6%)
Frame = -3
Query: 96 SEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQ 155
++ ++ L ++ L G+A+ W +G N ++ +W+ FL+KYFP+S E ++
Sbjct: 695 ADAIRLSLLSFSLSGEAKRWLHSFKG----NSLK-SWDEVVEKFLKKYFPESKTAEGKAA 531
Query: 156 FLTLRQGSMTIPEYAAKLESLAKHFRFFRDQV--------DEPYMCKRFVRGLRADIEDS 207
+ Q P+ ESL++ FR + EP F+ LR + +
Sbjct: 530 ISSFHQ----FPD-----ESLSEALERFRGLLRKTPTHGFSEPIQLNIFIDELRPESKQL 378
Query: 208 V 208
V
Sbjct: 377 V 375
>TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich protein, partial
(49%)
Length = 611
Score = 29.6 bits (65), Expect = 1.6
Identities = 16/54 (29%), Positives = 24/54 (43%)
Frame = +3
Query: 224 TEVELMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMKKPYQRPTGEGFTPGPYRP 277
T + K + + + G P + S+QGKG+ + P PT TP P P
Sbjct: 213 TIISRTKKKDVKKLGATHPCMHACSSFQGKGRTPLPPPTPVPTPPPPTPPPVPP 374
>AW317150
Length = 315
Score = 28.1 bits (61), Expect = 4.7
Identities = 17/43 (39%), Positives = 20/43 (45%), Gaps = 2/43 (4%)
Frame = -1
Query: 234 MDRAGTGGPMRSSPRSYQGKGKLQMK--KPYQRPTGEGFTPGP 274
+ R GGP +PR +GKG K KP PT G GP
Sbjct: 183 LGRGNGGGPFLKTPRGKKGKGAKGKKWGKPRGFPTPGG*KRGP 55
>TC206213 similar to UP|PMXA_MOUSE (Q62066) Paired mesoderm homeobox protein
2A (Paired-like homeobox 2A) (PHOX2A homeodomain
protein) (Aristaless homeobox protein homolog), partial
(8%)
Length = 930
Score = 28.1 bits (61), Expect = 4.7
Identities = 17/34 (50%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Frame = +1
Query: 35 RRAAEDARE-LHRLQREAALDQNRGLNDFRRQDP 67
RR A D R LHR+ R AAL Q GL R++ P
Sbjct: 205 RRGAGDRRPGLHRVVRRAALRQPAGLQALRQELP 306
>AW164264 homologue to GP|24210303|em SI:dZ12F11.3 (collagen type XI alpha-2)
{Danio rerio}, partial (1%)
Length = 294
Score = 27.7 bits (60), Expect = 6.1
Identities = 16/39 (41%), Positives = 20/39 (51%), Gaps = 4/39 (10%)
Frame = -1
Query: 240 GGPMRSSPRSYQGKGKLQMK----KPYQRPTGEGFTPGP 274
GGP +P +GKG K + + RPTGE PGP
Sbjct: 198 GGPFAKNPTGEKGKGPQGKKGGEPQGFPRPTGE--KPGP 88
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,230,778
Number of Sequences: 63676
Number of extensions: 100807
Number of successful extensions: 506
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 504
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 506
length of query: 282
length of database: 12,639,632
effective HSP length: 96
effective length of query: 186
effective length of database: 6,526,736
effective search space: 1213972896
effective search space used: 1213972896
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0338.2


