

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
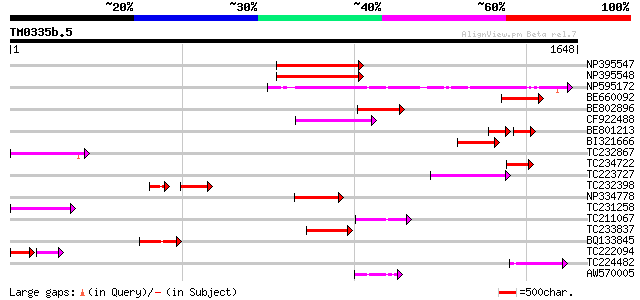
Query= TM0335b.5
(1648 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP395547 reverse transcriptase [Glycine max] 414 e-115
NP395548 reverse transcriptase [Glycine max] 411 e-114
NP595172 polyprotein [Glycine max] 384 e-106
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 193 5e-49
BE802896 184 3e-46
CF922488 173 5e-43
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 101 2e-41
BI321666 161 3e-39
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 138 2e-32
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 136 9e-32
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 134 3e-31
TC232398 90 1e-28
NP334778 reverse transcriptase [Glycine max] 119 1e-26
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 118 2e-26
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 114 5e-25
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 112 2e-24
BQ133845 107 3e-23
TC222094 70 5e-20
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 92 1e-18
AW570005 77 5e-14
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 414 bits (1064), Expect = e-115
Identities = 187/254 (73%), Positives = 222/254 (86%)
Frame = +1
Query: 775 VRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDY 834
VRKE+ KLL+AG+IYPISDS WVSPVQVVPKKGG+TVV N+ NELIPTR+VT+WR+CIDY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 835 RRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGV 894
R+LN TRKDH+PLPF+DQML RLA +Y FLDGYSGYNQI V P+DQEKTAFTCP+ V
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 895 FAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKR 954
FAY+RMPFGLCNA TFQRCM AIF D++E CIE+FMDDFS FG +F CL NL VL+R
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 955 CQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGH 1014
C+++NLVLNWEKCHFMV++GIVLGHK+S++GIEV + K++VI+KL PP N+KGI SFLGH
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 1015 AGFYRRFIKDFSKL 1028
GFYRRFIKDF+K+
Sbjct: 721 VGFYRRFIKDFTKV 762
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 411 bits (1056), Expect = e-114
Identities = 187/254 (73%), Positives = 221/254 (86%)
Frame = +1
Query: 775 VRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDY 834
VRKE++KLL+ G+IYPISDS WVSPV VV KK G+TV+ NE N+LIPTR VT W++CIDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 835 RRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGV 894
R+LN TRKDHFPLPF+DQML+RLAGH YYCFLD Y GYNQI V P+DQEK AFTCP+GV
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 895 FAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKR 954
FAY+R+PFGLCNAP TFQ CM AIF+D++E IE+FMDDFSVF P+ ++CL L +VL+R
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 955 CQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGH 1014
C ETNLVLNWEKCHFMVR+GIVLGHK+S +GIEVD+ KI+VIEKL PP+N+KGIRSFLG
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 1015 AGFYRRFIKDFSKL 1028
A FYRRFIKDF+K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 384 bits (986), Expect = e-106
Identities = 280/898 (31%), Positives = 434/898 (48%), Gaps = 12/898 (1%)
Frame = +1
Query: 749 HKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGG 808
H I L++ P+ R + KD + K I ++L G+I P S+S + P+ +V KK G
Sbjct: 1759 HAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGIIQP-SNSPFSLPILLVKKKDG 1935
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
WR C DYR LN++T KD FP+P +D++LD L G QY+ LD
Sbjct: 1936 ------------------SWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLD 2061
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
SGY+QI V PED+EKTAF +G + + MPFGL NAPATFQ M IF + +
Sbjct: 2062 LRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVL 2241
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+F DD ++ ++ L +L VL+ ++ L KC F + LGHKVS G+ +
Sbjct: 2242 VFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSM 2421
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
+ K++ + P N+K +R FLG G+YRRFIK ++ +A P+T+LL+K++ F ++
Sbjct: 2422 ENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDS-FLWNNEA 2598
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
AF +K+++ APV+ PD+S PF + DAS + +GAVL Q + I Y S+ L
Sbjct: 2599 EAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNG----HPIAYFSKKLA 2766
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
+ + +ELL + A KFR Y+LG K I+ TD +L+ L + P W+
Sbjct: 2767 PRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHK 2946
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFA 1228
+D +I + GKDN AD LSR + ++ EP ++
Sbjct: 2947 FLGYDFKIEYKPGKDNQAADALSR-------------------MFMLAWSEPHSIFLEEL 3069
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCH 1288
R+ PH + K DA Y + L+ D V+ E+ KIL H
Sbjct: 3070 RARLIS-DPHLKQLMETYKQGADASHYTVREGLLY--WKDRVVIPAEEEI-VNKILQEYH 3237
Query: 1289 GSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILE 1348
S GGH RT A+ L++ FYWP + + +A+++ C CQ+ N +P +
Sbjct: 3238 SSPIGGHAGITRTLAR-LKAQFYWPKMQEDVKAYIQKCLICQQA---KSNNTLPAGLLQP 3405
Query: 1349 IEL-FDVW---GIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTN-DSKVVVAFLKKN 1403
+ + VW +DF+ P SFG I+V +D ++K+ L + +SKVV +
Sbjct: 3406 LPIPQQVWEDVAMDFITGLPNSFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSH 3585
Query: 1404 IFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRIL 1463
I G+PR+I+SD F + ++ L + G +S+ YHPQ+ GQ E+ N+ L+ L
Sbjct: 3586 IVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYL 3765
Query: 1464 EKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKL 1523
K W + L A + Y TA+ +G +PF ++G+ P L +A
Sbjct: 3766 RCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGRE---PPTLTRQA------C 3918
Query: 1524 NFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNS 1583
+ D + QL + D + + + ++ K+ D+K L+ F G VL+
Sbjct: 3919 SIDDPAEVRE---QLTDRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQ 4089
Query: 1584 RLRLFPG------KLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTV-NGQRLKVYQG 1634
R KL R+ GPF V A ++E P I V + +LK + G
Sbjct: 4090 PYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHPVFHVSQLKPFNG 4263
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 193 bits (491), Expect = 5e-49
Identities = 88/123 (71%), Positives = 105/123 (84%)
Frame = -3
Query: 1429 SLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTA 1488
+LL+KYGV H+VSTPYHPQT+GQ EISNRE+KRILEK+V SRKDWS +LDDALWA+RTA
Sbjct: 370 ALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTA 191
Query: 1489 FKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRA 1548
+K PIG SP+ +VFGKACHLPVE+EHKAYWA++ NF A E+R LQL+ELDE RL A
Sbjct: 190 YKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRLEA 11
Query: 1549 YES 1551
YE+
Sbjct: 10 YEN 2
>BE802896
Length = 416
Score = 184 bits (467), Expect = 3e-46
Identities = 85/138 (61%), Positives = 110/138 (79%)
Frame = -2
Query: 1010 SFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPD 1069
SFLGHAGFYRRFI+DF K+A P++NLL+KE F F++ C AF+ +K +L+T P+I APD
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPD 236
Query: 1070 WSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACE 1129
W+ PFE+MCDAS+ ALG VL QK +++ VIYY+SR L+ AQ NYTTTEKELL +VFA E
Sbjct: 235 WTAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALE 56
Query: 1130 KFRPYILGFKVIVHTDHA 1147
KF Y+LG ++IV+ DHA
Sbjct: 55 KFHSYLLGTRIIVYIDHA 2
>CF922488
Length = 741
Score = 173 bits (439), Expect = 5e-43
Identities = 97/237 (40%), Positives = 136/237 (56%)
Frame = +3
Query: 830 VCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFT 889
+C+DYR LN + KD FPLP I+ ++D + F+DG+SGYNQI +APED EKT F
Sbjct: 30 MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAPEDMEKTTFI 209
Query: 890 CPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLA 949
+G F YK M FGL N AT+QR M A+F D++ IE++MDD V + L NL
Sbjct: 210 TLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRTEEEHLVNLR 389
Query: 950 LVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIR 1009
+ +R ++ L LN KC F V+ +L S +GIEVD K++VI ++ P K ++
Sbjct: 390 KLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMAKPHTEKQVQ 569
Query: 1010 SFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIV 1066
FLG + RFI +P+ LL K +D +C AFE IK+ L+ V+V
Sbjct: 570 GFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLINPHVLV 740
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 101 bits (252), Expect(2) = 2e-41
Identities = 46/63 (73%), Positives = 54/63 (85%)
Frame = +2
Query: 1464 EKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKL 1523
EK V SSRKDWS KL+DALWA +TA KTPIG +PF +V+ KACHLPVEL+HKAYWA++ L
Sbjct: 224 EKNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELKHKAYWAMKCL 403
Query: 1524 NFD 1526
NFD
Sbjct: 404 NFD 412
Score = 88.2 bits (217), Expect(2) = 2e-41
Identities = 38/66 (57%), Positives = 51/66 (76%)
Frame = +3
Query: 1391 NDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSG 1450
ND+K+V+ FLKKNIF+RFG+PR +ISDGG+HF +L+ V+HKV T YHPQT+G
Sbjct: 3 NDAKIVIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNG 182
Query: 1451 QVEISN 1456
Q ++SN
Sbjct: 183 QAKVSN 200
>BI321666
Length = 430
Score = 161 bits (407), Expect = 3e-39
Identities = 71/123 (57%), Positives = 95/123 (76%)
Frame = +2
Query: 1302 AAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMG 1361
+ VLQS FY P++ +++ +SC++CQRT ++S+RNE+PL ILE+E+FD WGIDF+G
Sbjct: 5 STNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVG 184
Query: 1362 PFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTH 1421
PFPPSF YILV VDYVSKWVEA A +D+K+V+ FLKK IF+R GVP +I +GG+H
Sbjct: 185 PFPPSFSNEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGSH 364
Query: 1422 FCN 1424
CN
Sbjct: 365 LCN 373
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 138 bits (348), Expect = 2e-32
Identities = 86/243 (35%), Positives = 130/243 (53%), Gaps = 13/243 (5%)
Frame = -3
Query: 1 SEDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLA 60
+EDP+AH+ +I CNT R+ V ++AIRLSL FSL A+ WL+S SL +W+++
Sbjct: 767 NEDPYAHLATYIEICNTIRLAGVPADAIRLSLLSFSLSGEAKRWLHSFKGNSLKSWDEVV 588
Query: 61 EKFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFY 120
EKF ++FP S + K I +F Q DE+L EA E F+ LLRK P H ++ + F
Sbjct: 587 EKFLKKYFPESKTAEGKAAISSFHQFPDESLSEALERFRGLLRKTPTHGFSEPIQLNIFI 408
Query: 121 DGLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMRAMNSENER---QIRRSSFEVETY 177
D L S+ +DA+ G+ P DLIE MA + +R ++S E+ +
Sbjct: 407 DELRPESKQLVDASVGGKIKMKTPDEAMDLIESMAASDIAILRDRAHIPTKKSLLELTSQ 228
Query: 178 DKLIASNKKLSEEVAEMQKHI----------QDTKSIGAKVTSLECKFCGESHDSNQCTV 227
D L+A NK LS+++ + K + Q + S +VT C GE+H+S C
Sbjct: 227 DTLLAQNKLLSKQLETLTKTLSKLPTQLHSAQTSHSSILQVTG--CTIFGEAHESGCCIP 54
Query: 228 NDD 230
N++
Sbjct: 53 NEE 45
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 136 bits (342), Expect = 9e-32
Identities = 59/78 (75%), Positives = 69/78 (87%)
Frame = -3
Query: 1444 YHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFG 1503
YHPQT+GQ E+SN+E+KR+LE +V SSRKDW+ KLDDA WAYR AFKTPIG SPF LV+G
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 1504 KACHLPVELEHKAYWAIR 1521
KACHL VELEHKAYWA++
Sbjct: 54 KACHLSVELEHKAYWALK 1
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 134 bits (338), Expect = 3e-31
Identities = 73/236 (30%), Positives = 119/236 (49%), Gaps = 2/236 (0%)
Frame = +1
Query: 1222 PWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFE 1281
PWY + ++ ++ K+ A + L+K D RC+ +
Sbjct: 136 PWYFDIKRYVISKEYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREAN 315
Query: 1282 KILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEM 1341
++ H S+G H +G A K+L++G+YW T+ + V C +CQ +
Sbjct: 316 HMIEEVHEGSFGTHANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPH 495
Query: 1342 PLKNILEIELFDVWGIDFMGPFPP--SFGC*YILVAVDYVSKWVEASALSTNDSKVVVAF 1399
PL + F +WGID +G P S G +ILVA+DY +KWVEA++ + VVV F
Sbjct: 496 PLNVMSSPWPFSMWGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRF 675
Query: 1400 LKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEIS 1455
+KK I R+G+PR II+D GT+ N+ + E++ ++H TPY P+ + VE++
Sbjct: 676 IKKEIICRYGLPRKIITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>TC232398
Length = 1054
Score = 90.1 bits (222), Expect(2) = 1e-28
Identities = 38/93 (40%), Positives = 66/93 (70%)
Frame = +1
Query: 498 KRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLIL 557
+R+ ++ PT ++LQ+AD S+ +G++ED++VKV + +F VDFVI+D+EED+++ LIL
Sbjct: 172 RRIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLIL 351
Query: 558 GRPFLATGEAEIKVAKGTLTLKVGEDEVLFNIF 590
G PF+ T + + + KG L + V + + FN+F
Sbjct: 352 GWPFMVTAKCVVDMGKGNLEMSVEDQKATFNLF 450
Score = 56.6 bits (135), Expect(2) = 1e-28
Identities = 30/60 (50%), Positives = 41/60 (68%)
Frame = +2
Query: 406 LHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTEECSAILQRKMPKKRRDPG 465
L I IPF E ++QMP+Y KF+KDIL K+ K ETI++ E C A++Q K+P K +D G
Sbjct: 2 LEITIPFGERIQQMPLYKKFLKDILIKKGKYIN-SETIVVGEYCRALIQ-KLPPKFKDLG 175
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 119 bits (297), Expect = 1e-26
Identities = 61/141 (43%), Positives = 85/141 (60%)
Frame = +3
Query: 829 RVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAF 888
R+C+DYR LN + KD+FPLP ID ++ +A + F+DG+SGYNQI +APED EKT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 889 TCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNL 948
+G F YK M FGL N AT+ R M A+F D++ IE ++D+ + L NL
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 949 ALVLKRCQETNLVLNWEKCHF 969
+ + ++ L LN KC F
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVF 425
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 118 bits (296), Expect = 2e-26
Identities = 66/192 (34%), Positives = 98/192 (50%), Gaps = 2/192 (1%)
Frame = +2
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
EDPH H+++F C+T + +V + I L FP SL+ A++WL S+ W+DL
Sbjct: 572 EDPHKHLKEFHIVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRSIFNWDDLKR 751
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYD 121
F +FFP S ++ DI Q + E+LYE WE FKK CP H +++ + FY+
Sbjct: 752 VFLEKFFPASRTTTIRKDISGIRQLSRESLYEYWERFKKSCASCPHHQISKQLLL*YFYE 931
Query: 122 GLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMRA--MNSENERQIRRSSFEVETYDK 179
L R +DAAS G + P +LI+KMA + ++ N+ + R EV T
Sbjct: 932 ELSNMKRSMIDAASGGALGDMTPTEARNLIKKMASNSQQFSARNDAIVLRGVHEVATDSS 1111
Query: 180 LIASNKKLSEEV 191
NK L E +
Sbjct: 1112SSTENKSLRENL 1147
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 114 bits (284), Expect = 5e-25
Identities = 61/163 (37%), Positives = 96/163 (58%)
Frame = +1
Query: 1004 NIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAP 1063
++ IRSF G A FYRRF+ +FS +A P+ L++K FT+ E +AF +KE L AP
Sbjct: 109 SVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKAP 288
Query: 1064 VIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLG 1123
V+ PD+S FE+ CDAS + + AVL Q + I Y S L+ A NY T +KEL
Sbjct: 289 VLALPDFSKTFELECDASGVGVRAVLLQGG----HPIAYFSEKLHSATLNYPTYDKELYA 456
Query: 1124 VVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWV 1166
++ A + + +++ + ++H+DH +L+++ K R +WV
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWV 585
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 112 bits (279), Expect = 2e-24
Identities = 59/133 (44%), Positives = 82/133 (61%)
Frame = +2
Query: 864 YCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLI 923
+ F+DG+SGYNQI +A ED EKT F +G F+Y+ M FGL N AT+QR M A+F D++
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 924 ETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSE 983
IE+++DD L NL + R Q+ L LN KC F V+ G +LG VS+
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 984 KGIEVDRAKIEVI 996
KGIE+D K++ +
Sbjct: 362 KGIEIDPEKVKAL 400
>BQ133845
Length = 389
Score = 107 bits (268), Expect = 3e-23
Identities = 54/123 (43%), Positives = 83/123 (66%)
Frame = +2
Query: 377 ELRKIPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKL 436
E +++P+P K+ ++ +KFL++FKKL I +PF EAL+QMP+YA F+KD+L+K+
Sbjct: 23 ECKEVPYPLVPS*KDKEQHLAKFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNWY 202
Query: 437 SEVDETILMTEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRM 496
D+ I++ CSA++QR +P DPG T+P I + + +AL DLGASINLM L M
Sbjct: 203 IHSDK-IVVEGNCSAVIQRILPP*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSM 379
Query: 497 FKR 499
++
Sbjct: 380 CRQ 388
>TC222094
Length = 984
Score = 70.5 bits (171), Expect(2) = 5e-20
Identities = 32/71 (45%), Positives = 43/71 (60%)
Frame = -2
Query: 1 SEDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLA 60
+EDP+AH+ +I CNT ++V V +AI L LF FSL A WL S +L TW +
Sbjct: 707 TEDPYAHLATYIDICNTVKIVGVPEDAIHLDLFCFSLAGEARTWLRSFKGNNLRTWNEXX 528
Query: 61 EKFTTRFFPRS 71
EKF ++FP S
Sbjct: 527 EKFLKKYFPES 495
Score = 47.4 bits (111), Expect(2) = 5e-20
Identities = 28/79 (35%), Positives = 42/79 (52%)
Frame = -3
Query: 77 KNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYDGLLYSSRFGLDAASS 136
K +I +F Q E+L EA + F LL K P H ++ + F DG+ S+ LDA++
Sbjct: 478 KVEISSFHQHPHESLSEALDRFHGLLWKTPTHGFSEPVQLNIFIDGMQPHSKQLLDASAG 299
Query: 137 GEFDALPPQAGYDLIEKMA 155
G+ P+ +LIE MA
Sbjct: 298 GKIKLKTPEEAIELIENMA 242
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 92.4 bits (228), Expect = 1e-18
Identities = 56/170 (32%), Positives = 91/170 (52%), Gaps = 1/170 (0%)
Frame = +1
Query: 1453 EISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVEL 1512
E +N+ +K+I++K+ S KDW L AL YRT+ +T G +PF LV+G LP E+
Sbjct: 1 EAANKNIKKIIQKMT-VSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 1513 EHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREF 1572
E + + + ++ R QLN ++ RL A +Y+++ K D+K+ R+F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 1573 VSGQLVL-LFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNI 1621
G LVL + ++ GK + GPFVVKR F GA+ + N + + +
Sbjct: 358 HEGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEEL 507
>AW570005
Length = 413
Score = 77.4 bits (189), Expect = 5e-14
Identities = 44/141 (31%), Positives = 78/141 (55%)
Frame = -2
Query: 1001 PPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLV 1060
PP + +R FL GFYRRFIK ++ +A P+++LL K++ F + AF+++K +
Sbjct: 409 PPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDS-FVWSPEADVAFQALKNVVT 233
Query: 1061 TAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKE 1120
V+ PD++ PF + DAS +GAVL Q+ + +++ + R+ +T E
Sbjct: 232 NTLVLALPDFTKPFTVETDASGSDMGAVLSQEGHP---IAFFSKEFCPKLVRS-STYVHE 65
Query: 1121 LLGVVFACEKFRPYILGFKVI 1141
L + +K+R Y+LG ++
Sbjct: 64 LAAITNVVKKWRQYLLGHHLV 2
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 72,415,091
Number of Sequences: 63676
Number of extensions: 1036259
Number of successful extensions: 4615
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 4541
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4599
length of query: 1648
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1538
effective length of database: 5,635,272
effective search space: 8667048336
effective search space used: 8667048336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0335b.5


