

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0334a.1
(1167 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
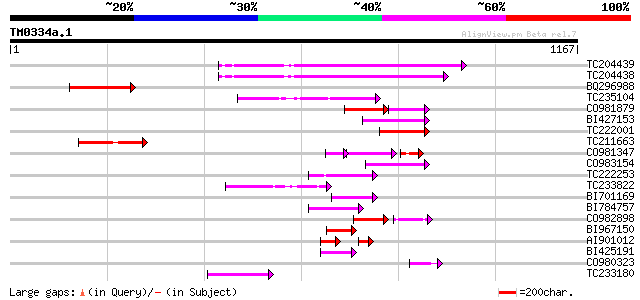
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 164 2e-40
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 160 3e-39
BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativ... 154 2e-37
TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotei... 144 2e-34
CO981879 94 7e-30
BI427153 113 4e-25
TC222001 similar to UP|C716_NEPRA (O04164) Cytochrome P450 71A6 ... 108 2e-23
TC211663 107 3e-23
CO981347 65 5e-22
CO983154 100 5e-21
TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement p... 87 3e-17
TC233822 75 2e-13
BI701169 72 2e-12
BI784757 70 4e-12
CO982898 50 9e-12
BI967150 60 6e-09
AI901012 39 2e-07
BI425191 weakly similar to GP|14586969|gb| pol polyprotein {Citr... 54 3e-07
CO980323 53 9e-07
TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, parti... 53 9e-07
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 164 bits (415), Expect = 2e-40
Identities = 146/526 (27%), Positives = 235/526 (43%), Gaps = 15/526 (2%)
Frame = +1
Query: 430 HSTLSYSSQYDSRDWIIDSGASDHICFNKLCFDTLNRIKPVS---VRLPNGISIQTCYAG 486
H++L S++ DW +DSG S H+ K + L I+P S V +G + G
Sbjct: 1651 HTSLRASAK---EDWYLDSGCSRHMTGVK---EFLLNIEPCSTSYVTFGDGSKGKIIGMG 1812
Query: 487 VVSITKQILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSL 546
+ L VL V T NLIS+ +L V F + C V + ++ + GS
Sbjct: 1813 KLVHDGLPSLNKVLLVKGLTANLISISQLCDEG-FNVNFTKSECLVTNEKSEVLMK-GSR 1986
Query: 547 KEDDLYHLVV--TSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHL---SHDRILAL 601
+D+ Y TS SSTC + S D++ +WH R GHL +I+
Sbjct: 1987 SKDNCYLWTPQETSYSSTCLS--------SKEDEV----RIWHQRFGHLHLRGMKKIIDK 2130
Query: 602 NAL--YPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCN-FDLLHMDIWGPISVSSVHS 658
A+ P++ + + +C C + K + H +LLHMD+ GP+ V S+
Sbjct: 2131 GAVRGIPNLKIEEGRICGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGG 2310
Query: 659 HRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---M 715
RY V+DD SRF WV ++ K E + K ++ + +K IRSD+G EF
Sbjct: 2311 KRYAYVVVDDFSRFTWVNFIREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSR 2490
Query: 716 LHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVF 775
+F ++ GI H+ S TPQQNG VERK++ + AR +L LP +W A+ A +
Sbjct: 2491 FTEFCTSEGITHEFSAAITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACY 2670
Query: 776 LMSRIQSKVLENKSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLG 835
+ +R+ + + YEI + G+K ++ +FGS C+ R K+D ++ +FLG
Sbjct: 2671 IHNRVTLRRGTPTTLYEI-WKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLG 2847
Query: 836 FKQGVKGFVLLDLNSHEIFVSRDVTFHEQILPYKSATSTAWTCLDLEVKDQSSSSLNIPE 895
+ + + + + + + S +V + K V D + S N
Sbjct: 2848 YSTNSRAYRVFNSRTRTVMESINVVVDDLSPARKKDVEEDVRTSGDNVADAAKSGENAEN 3027
Query: 896 TTSAAQEL-ISDNENFSNT*LPCHTQSETIIEEENTQSETLIEEEE 940
+ SA E I+ + S+T + E II + N T E E
Sbjct: 3028 SDSATDESNINQPDKRSSTRIQKMHPKELIIGDPNRGVTTRSREVE 3165
Score = 31.6 bits (70), Expect = 2.1
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +1
Query: 317 CTHCKKPGHTVDVCYRLHGYP 337
C +C K GH CY LHG+P
Sbjct: 1510 CHYCGKYGHIKPFCYHLHGHP 1572
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 160 bits (405), Expect = 3e-39
Identities = 135/487 (27%), Positives = 220/487 (44%), Gaps = 14/487 (2%)
Frame = +1
Query: 430 HSTLSYSSQYDSRDWIIDSGASDHICFNKLCFDTLNRIKPVS---VRLPNGISIQTCYAG 486
H++L S++ DW +DSG S H+ K + L I+P S V +G + G
Sbjct: 1654 HTSLRASAK---EDWYLDSGCSRHMTGVK---EFLVNIEPCSTSYVTFGDGSKGKITGMG 1815
Query: 487 VVSITKQILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSL 546
+ L VL V T NLIS+ +L V F + C V + ++ + GS
Sbjct: 1816 KLVHDGLPSLNKVLLVKGLTANLISISQLCDEG-FNVNFTKSECLVTNEKSEVLMK-GSR 1989
Query: 547 KEDDLYHLVV--TSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHL---SHDRILAL 601
+D+ Y TS SSTC S D++ +WH R GHL +I+
Sbjct: 1990 SKDNCYLWTPQETSYSSTCL--------FSKEDEV----KIWHQRFGHLHLRGMKKIIDK 2133
Query: 602 NAL--YPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCN-FDLLHMDIWGPISVSSVHS 658
A+ P++ + + +C C + K + H +LLHMD+ GP+ V S+
Sbjct: 2134 GAVRGIPNLKIEEGRICGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGG 2313
Query: 659 HRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---M 715
RY V+DD SRF WV ++ K + + K ++ + +K IRSD+G EF
Sbjct: 2314 KRYAYVVVDDFSRFTWVNFIREKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSK 2493
Query: 716 LHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVF 775
+F ++ GI H+ S TPQQNG VERK++ + AR +L LP +W A+ A +
Sbjct: 2494 FTEFCTSEGITHEFSAAITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACY 2673
Query: 776 LMSRIQSKVLENKSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLG 835
+ +R+ + + YEI + G+K ++ +FGS C+ R K+D ++ +FLG
Sbjct: 2674 IHNRVTLRRGTPTTLYEI-WKGRKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLG 2850
Query: 836 FKQGVKGFVLLDLNSHEIFVSRDVTFHEQILPYKSATSTAWTCLDLEVKDQSSSSLNIPE 895
+ + + + + + + S +V + K V D + S+ N
Sbjct: 2851 YSTNSRAYRVFNSRTRTVMESINVVVDDLTPARKKDVEEDVRTSGDNVADTAKSAENAEN 3030
Query: 896 TTSAAQE 902
+ SA E
Sbjct: 3031 SDSATDE 3051
Score = 31.6 bits (70), Expect = 2.1
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +1
Query: 317 CTHCKKPGHTVDVCYRLHGYP 337
C +C K GH CY LHG+P
Sbjct: 1513 CHYCGKYGHIKPFCYHLHGHP 1575
>BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 408
Score = 154 bits (389), Expect = 2e-37
Identities = 75/135 (55%), Positives = 94/135 (69%)
Frame = -1
Query: 124 RCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQN 183
RCN LIHSWILNSV SI+ SIVF++NA D W DL++RFSQG LVR++E+Q EI L Q
Sbjct: 408 RCNMLIHSWILNSVEPSISRSIVFMDNASDVWLDLKERFSQGDLVRVSEIQQEIYALTQG 229
Query: 184 TLSVNDYYTEIKTLWEELEQYRPIPQCRCVVPCRCEAIEHAKMFREQDNAIRFLLGLNET 243
T SV +Y++ K LWEELE Y PIP C C C C+A+ A+ + +RFL GLN+
Sbjct: 228 TRSVTTFYSDKKALWEELEIYMPIPNCTCHHRCSCDAMRLARRHHHTLHVMRFLTGLNDE 49
Query: 244 FSVVNSQILMSNPLP 258
F+ V SQIL+ PLP
Sbjct: 48 FNAVKSQILLIEPLP 4
>TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotein
(Fragment), partial (28%)
Length = 865
Score = 144 bits (364), Expect = 2e-34
Identities = 98/297 (32%), Positives = 151/297 (49%), Gaps = 3/297 (1%)
Frame = +2
Query: 469 PVSVRLPNGISIQTCYAGVVSITKQILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEF 528
P + L +G + G VS T + L V+++ +N+ S+ +L + C V F
Sbjct: 11 PYFITLADGSRVVATGIGHVSPTSSLSLNSVVFILGCPFNITSLSQLTRFRNCSVTFDAN 190
Query: 529 TCFVQEKHTKKRIGLGSLKEDDLYHLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHF 588
+ +QE T IG+G ++ LY+L + S C A+ + L H
Sbjct: 191 SFVIQECGTGWTIGVG-IESHGLYYLK-PNLSWVCSAVTSP--------------KLLHE 322
Query: 589 RLGHLSHDRILALNALYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNFDLLHMDIW 648
RLGH + L + PS++ K C+ C L K R F ++H DIW
Sbjct: 323 RLGH---PHLSKLKIMVPSLEKIKDLFCESCQLGKHVRSSXRHVESRVDSPFLVIHXDIW 493
Query: 649 GPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRS 708
GP VSS+ S+RYF+T +D+ S+ V L+K + E+ + + V +KTQF +K++RS
Sbjct: 494 GPNRVSSM-SYRYFVTFIDEFSQCTRVFLMKERSEILSFLTS-VNKIKTQFGKTIKILRS 667
Query: 709 DNGPEF---MLHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLP 762
DN E+ ++ F SA GILHQ SC +TPQQN ERK++H++ AR LL ++ P
Sbjct: 668 DNAKEYFSSVISPFXSAQGILHQFSCPHTPQQNDIAERKNRHLVETARTLLLHANEP 838
>CO981879
Length = 576
Score = 94.0 bits (232), Expect(2) = 7e-30
Identities = 46/94 (48%), Positives = 62/94 (65%), Gaps = 3/94 (3%)
Frame = -1
Query: 689 KNFVALVKTQFSHHVKVIRSDNGPEFM---LHDFYSAHGILHQRSCVNTPQQNGRVERKH 745
K F +++TQF +KV RSDNG E+ L +GI+HQ SCV+TPQQNG ERK+
Sbjct: 564 KTFFQMIQTQFQVKIKVFRSDNGREYFNKHLSKXXLENGIIHQSSCVDTPQQNGVAERKN 385
Query: 746 QHILNIARALLFQSHLPKKMWGYAILHAVFLMSR 779
+H+ +ARALLFQ+ PK WG AIL +L ++
Sbjct: 384 RHLXEVARALLFQNKAPKYXWGEAILTGTYLKNK 283
Score = 56.6 bits (135), Expect(2) = 7e-30
Identities = 30/90 (33%), Positives = 50/90 (55%), Gaps = 4/90 (4%)
Frame = -2
Query: 779 RIQSKVLENKSPYEIL---FGGKKIDLQ-ELRVFGSLCFASTSSSSRSKLDSRARKCVFL 834
R+ SK+L ++P ++ F ++ L++FG F ++ KL+ RA+KCVF+
Sbjct: 284 RMPSKILNFRTPLDVFTSAFPNNRLSCTLPLKIFGCTVFVHIHEPNQGKLEPRAKKCVFV 105
Query: 835 GFKQGVKGFVLLDLNSHEIFVSRDVTFHEQ 864
G+ KG+ D S + FV+ DVTF E+
Sbjct: 104 GYAPNQKGYKCFDPTSKKTFVTIDVTFFEK 15
>BI427153
Length = 422
Score = 113 bits (283), Expect = 4e-25
Identities = 57/137 (41%), Positives = 82/137 (59%)
Frame = +1
Query: 727 HQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLE 786
HQ +C +TPQQNG ERK+ H+L AR+L+ S++P WG A+L A FL++R+ S LE
Sbjct: 1 HQSTCPHTPQQNGIAERKNHHLLETARSLMLNSNVPTHHWGDAVLTACFLINRMPSSSLE 180
Query: 787 NKSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLL 846
N+ P+ I+F + +VFG CF S KL +R+ KCVFLG+ + KG+
Sbjct: 181 NQIPHSIVFPNDLLFYVSPKVFGCTCFVHDLSPGLDKLSARSVKCVFLGYSRLQKGYTCY 360
Query: 847 DLNSHEIFVSRDVTFHE 863
N ++S +VTF E
Sbjct: 361 FPNMRRYYMSANVTFFE 411
>TC222001 similar to UP|C716_NEPRA (O04164) Cytochrome P450 71A6 (Fragment)
, partial (21%)
Length = 912
Score = 108 bits (269), Expect = 2e-23
Identities = 50/103 (48%), Positives = 70/103 (67%)
Frame = -2
Query: 761 LPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGGKKIDLQELRVFGSLCFASTSSSS 820
+P W YA+LHA +L++ I + L+N SPYE L G D+ LR+FG LC+AST ++
Sbjct: 911 MPPNFWNYALLHAAYLINCIPTPFLQNTSPYERLHGHIP-DISHLRIFGCLCYASTIKAN 735
Query: 821 RSKLDSRARKCVFLGFKQGVKGFVLLDLNSHEIFVSRDVTFHE 863
R KL+ RA C+F+GFK KG++L DL+SH I SR+V F+E
Sbjct: 734 RKKLEPRAHPCIFIGFKPNTKGYMLYDLHSHNIITSRNVVFYE 606
>TC211663
Length = 426
Score = 107 bits (267), Expect = 3e-23
Identities = 54/144 (37%), Positives = 90/144 (62%)
Frame = -3
Query: 141 IANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQNTLSVNDYYTEIKTLWEE 200
IA +++ ++A + W DL++ FSQG L++IAELQ EI LKQ + +V D++T++K +WEE
Sbjct: 424 IAQIVIYFDHATNIWNDLKEGFSQGDLLQIAELQEEIYRLKQGSHTVLDFFTKLKFVWEE 245
Query: 201 LEQYRPIPQCRCVVPCRCEAIEHAKMFREQDNAIRFLLGLNETFSVVNSQILMSNPLPPI 260
L+ Y + C C ++ + +QD I FL GL+E FSVV S++L+ + LP
Sbjct: 244 LDNYGLMNLCTC----------PSRTYHQQDFVIHFLKGLDERFSVVCSEVLLMDHLPST 95
Query: 261 AKVVSLAMQHERQSETGENEESKS 284
++ S+ +QHE Q + + E ++
Sbjct: 94 KRIFSMVIQHETQHASHTSAEDQN 23
>CO981347
Length = 624
Score = 65.1 bits (157), Expect(3) = 5e-22
Identities = 39/106 (36%), Positives = 55/106 (51%), Gaps = 3/106 (2%)
Frame = +2
Query: 694 LVKTQFSHHVKVIRSDNGPEFML---HDFYSAHGILHQRSCVNTPQQNGRVERKHQHILN 750
L+ Q +KV+R+DNG EF+L ++F GI + +TP QNG ER + IL
Sbjct: 134 LIGNQLGTKLKVLRTDNGLEFVLEQFNEFCRKIGIKRHKIVPHTP*QNGLAERMNMTILE 313
Query: 751 IARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFG 796
R +L + LPK WG A +L++R S L K+P E G
Sbjct: 314 RVRCMLLSARLPKTFWGEAANTTSYLINRCPSSTLGFKTPMEAWSG 451
Score = 42.4 bits (98), Expect(3) = 5e-22
Identities = 23/48 (47%), Positives = 30/48 (61%)
Frame = +3
Query: 805 LRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLDLNSHE 852
L+VFGSL F + KLD+RA KCVF+G+ +GVK + L L E
Sbjct: 474 LKVFGSLAFDHVK---QGKLDARAVKCVFIGYPKGVKRYKLWKLEPGE 608
Score = 36.6 bits (83), Expect(3) = 5e-22
Identities = 18/47 (38%), Positives = 28/47 (59%)
Frame = +3
Query: 650 PISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVK 696
P V + YFLT++DD SR VW+ +LK K E + +N + L++
Sbjct: 3 PSRVKTHGGSSYFLTIIDDFSRRVWLYVLKNKSESFKNSENDILLLE 143
>CO983154
Length = 568
Score = 100 bits (248), Expect = 5e-21
Identities = 51/131 (38%), Positives = 77/131 (57%)
Frame = +3
Query: 733 NTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYE 792
+TPQQNG ERK++H+L AR+L+ ++P WG A+L + FL++R+ S LEN+ P+
Sbjct: 3 HTPQQNGIAERKNRHLLETARSLMLNLNVPIHHWGDAVLTSCFLINRMPSSSLENQIPHS 182
Query: 793 ILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLDLNSHE 852
++F + +VFG CF S KL +R+ KCVFLG+ + KG+
Sbjct: 183 LVFPHDPLFHVSPKVFGCTCFVHDLSPGLDKLSARSVKCVFLGYSRLQKGYKCYSPTMRR 362
Query: 853 IFVSRDVTFHE 863
++S DVTF E
Sbjct: 363 YYMSADVTFFE 395
>TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement pol
polyprotein, partial (4%)
Length = 919
Score = 87.4 bits (215), Expect = 3e-17
Identities = 49/144 (34%), Positives = 77/144 (53%), Gaps = 2/144 (1%)
Frame = +1
Query: 615 VCDICHLAKLKRKMFPDSLH-NAKCNFDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFV 673
VCD C + K R+ FP K ++H+D+ + + + + YF+T +DD S+ +
Sbjct: 328 VCDTCEIGKKHRESFPTGKSWRMKKLLKIVHLDLC-TVEIPTHGDNNYFITFIDDFSKKM 504
Query: 674 WVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEFMLHD-FYSAHGILHQRSCV 732
WV LK+K E K F A + Q VK + D G E++ + F+ HGI HQ +
Sbjct: 505 WVYFLKQKSEACNAFKMFKAFAEKQNGCKVKALIIDKGQEYLSYTIFFEKHGIQHQLTTK 684
Query: 733 NTPQQNGRVERKHQHILNIARALL 756
TPQ NG ERK++ I+++ R +L
Sbjct: 685 YTPQHNGVTERKNKTIMDMVRCML 756
>TC233822
Length = 632
Score = 75.1 bits (183), Expect = 2e-13
Identities = 68/221 (30%), Positives = 103/221 (45%), Gaps = 3/221 (1%)
Frame = +2
Query: 445 IIDSGASDHICFNKLCFDTLNRIK-PVSVRLPNGISIQTCYAGVVSITKQILLTHVLYVP 503
+IDS ASDHI N F + + K P + L NG + + G SI + L + +VP
Sbjct: 38 VIDSTASDHIFGNSSLFTSQSPPKIPHLITLANGTKVTSKGFGKFSIFPSLNLDPIHFVP 217
Query: 504 EFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTSPSSTC 563
+NL+S+ +L K + F + +QE+ T IG+ + LY+L ++S S +C
Sbjct: 218 NCPFNLVSLSQLTKALNFSITFDADSFVIQERDTSWLIGV-EHESRGLYYLEISS-SMSC 391
Query: 564 FAIPNVVERISDSDQLIPPGALWHFRLGHLSHDRILALNALYPSIDVSKHFVCDICHLAK 623
FA P S +L+ HL H +L L + PS++ + C+ C L K
Sbjct: 392 FATP--------SPKLLH---------NHLGHPHLLKLK-MVPSLNKLQVLECESCQLGK 517
Query: 624 LKRKMFPDSLHNAKCN--FDLLHMDIWGPISVSSVHSHRYF 662
R FP +C+ F + DIW P V+S RYF
Sbjct: 518 HVR-FFPKRT-ETRCSSVFSTIPSDIWDPSCVTS-FGFRYF 631
>BI701169
Length = 407
Score = 71.6 bits (174), Expect = 2e-12
Identities = 40/98 (40%), Positives = 56/98 (56%), Gaps = 3/98 (3%)
Frame = +2
Query: 662 FLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEFMLHDFY- 720
F +DD SR WV K K EV + K F A+V+ + +K +RSD G EF ++F
Sbjct: 113 FFLFIDDFSRKTWVYFFKHKLEVFENFKKFKAIVEKESGFKIKAMRSDRGGEF*SNEFQK 292
Query: 721 --SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALL 756
HGI + +PQQNG ERK++ ILN+AR++L
Sbjct: 293 YCDDHGIRRPLMVLRSPQQNGVAERKNRTILNMARSML 406
>BI784757
Length = 430
Score = 70.5 bits (171), Expect = 4e-12
Identities = 35/118 (29%), Positives = 66/118 (55%), Gaps = 4/118 (3%)
Frame = +1
Query: 615 VCDICHLAKLKRKMFPDSLH-NAKCNFDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFV 673
VCD C K R F ++ AK ++++ D+ GP+ S+ +RYF++ +D+ +R V
Sbjct: 64 VCDGCLQCKQSRSTFKQNVPIRAKEKLEVIYSDVCGPMQTESLGGNRYFISFIDELTRKV 243
Query: 674 WVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEFM---LHDFYSAHGILHQ 728
WV L++RK + + + F + K Q +K++R++ G E++ +F GI+H+
Sbjct: 244 WVYLIRRKSDFFEVFEKFKNMAKKQSGSLIKILRTNGGGEYVSTEFQEFCDQQGIIHE 417
>CO982898
Length = 816
Score = 49.7 bits (117), Expect(2) = 9e-12
Identities = 30/73 (41%), Positives = 44/73 (60%), Gaps = 1/73 (1%)
Frame = -3
Query: 709 DNGPEF-MLHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWG 767
D G EF L +F + GI+H+ +T QN VERK +HI+ +LL + LP K WG
Sbjct: 814 DWGGEF*PLTEFLND*GIIHKLIFPHTQYQNV-VERKQRHIVKFGLSLLSHASLPLKFWG 638
Query: 768 YAILHAVFLMSRI 780
+A + VFL++R+
Sbjct: 637 HAFITPVFLINRL 599
Score = 39.7 bits (91), Expect(2) = 9e-12
Identities = 27/80 (33%), Positives = 38/80 (46%)
Frame = -1
Query: 790 PYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLDLN 849
PY LF + +D RVFG CF + K + ++ +C+FLG K + L
Sbjct: 570 PYTQLFN-QMLDYSFPRVFGCACFPLIRPYNIQKFNFKSHECIFLGNYTSHKNYKCLAPA 394
Query: 850 SHEIFVSRDVTFHEQILPYK 869
VS+DV F+E PYK
Sbjct: 393 GK--LVSKDVIFNEI*FPYK 340
>BI967150
Length = 574
Score = 60.1 bits (144), Expect = 6e-09
Identities = 29/63 (46%), Positives = 44/63 (69%), Gaps = 1/63 (1%)
Frame = +1
Query: 652 SVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHH-VKVIRSDN 710
++SS+H H YFL V+DD++R W+ L+K K E ++ + NFVAL +T F HH + R++N
Sbjct: 277 ALSSLHGHGYFLIVVDDYTRRTWLFLMKLKSEARKLMYNFVALTQT*F*HHCFETARTNN 456
Query: 711 GPE 713
G E
Sbjct: 457 GNE 465
>AI901012
Length = 410
Score = 39.3 bits (90), Expect(2) = 2e-07
Identities = 16/31 (51%), Positives = 22/31 (70%)
Frame = +2
Query: 719 FYSAHGILHQRSCVNTPQQNGRVERKHQHIL 749
F ++HG++HQ SC +T QQNG K QHI+
Sbjct: 317 FMTSHGMIHQSSCPHTLQQNGVANHKLQHII 409
Score = 35.4 bits (80), Expect(2) = 2e-07
Identities = 14/40 (35%), Positives = 25/40 (62%)
Frame = +1
Query: 641 DLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKR 680
+++H +IWG + S+ RY++T +DD SR +L +R
Sbjct: 76 NIVHSNIWGHSCILSILGFRYYVTFIDDFSRHTXTLLNER 195
>BI425191 weakly similar to GP|14586969|gb| pol polyprotein {Citrus x
paradisi}, partial (7%)
Length = 423
Score = 54.3 bits (129), Expect = 3e-07
Identities = 27/73 (36%), Positives = 42/73 (56%)
Frame = +3
Query: 641 DLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFS 700
+++H DI P V+S RYF+T +DD+S + +V LL K + ++ + V+ Q
Sbjct: 105 EIMHTDICRPFDVNSFGKERYFITFIDDYSHYGYVYLLHEKFQAVDVLEIHLNEVERQLD 284
Query: 701 HHVKVIRSDNGPE 713
VKV+RSD G E
Sbjct: 285 RKVKVVRSDRGGE 323
>CO980323
Length = 769
Score = 52.8 bits (125), Expect = 9e-07
Identities = 27/69 (39%), Positives = 37/69 (53%)
Frame = -2
Query: 823 KLDSRARKCVFLGFKQGVKGFVLLDLNSHEIFVSRDVTFHEQILPYKSATSTAWTCLDLE 882
K D RA KC F G+K GVKG++ ++S EI +SR+ F+E I P CL L
Sbjct: 216 KFDGRA*KCAFWGYKAGVKGYLAYGIHSKEILISRNAQFYEHIFP----------CLSLS 67
Query: 883 VKDQSSSSL 891
+ S S+
Sbjct: 66 ISQSSLLSM 40
>TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partial (11%)
Length = 916
Score = 52.8 bits (125), Expect = 9e-07
Identities = 40/138 (28%), Positives = 65/138 (46%), Gaps = 2/138 (1%)
Frame = +2
Query: 408 KAFANLVTK-SAVCGANTAGNGKHSTLSYSSQYDSRDWIIDSGASDHICFNKLCFDTLN- 465
+AF + +K S+ C AG S LS+++ IIDSG +DH+ + F +
Sbjct: 344 RAFMDPFSKTSSSCSLTMAGKSS-SFLSFNASGTENI*IIDSGVTDHMTPHSSYFSSYTF 520
Query: 466 RIKPVSVRLPNGISIQTCYAGVVSITKQILLTHVLYVPEFTYNLISVHKLAKRNRCEVVF 525
I + + NG I G + + + L +VLYVP+ + NL+S+HK+ C V F
Sbjct: 521 LIGNQHIIVANGSHIPIIGCGNIQLQSSLHLNNVLYVPKLSNNLLSIHKIT*DLNCVVTF 700
Query: 526 GEFTCFVQEKHTKKRIGL 543
C + + IG+
Sbjct: 701 FHSHCVF*DLAMGRMIGI 754
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,484,894
Number of Sequences: 63676
Number of extensions: 841697
Number of successful extensions: 5446
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 5256
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5412
length of query: 1167
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1059
effective length of database: 5,762,624
effective search space: 6102618816
effective search space used: 6102618816
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0334a.1


