

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0333b.7
(293 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
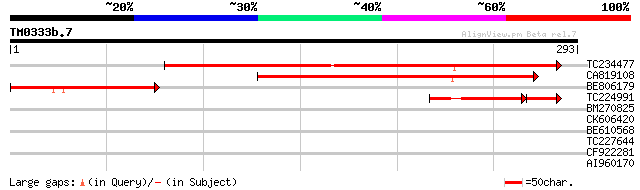
Score E
Sequences producing significant alignments: (bits) Value
TC234477 weakly similar to UP|DRPE_CRAPL (P22242) Desiccation-re... 241 3e-64
CA819108 192 2e-49
BE806179 75 5e-14
TC224991 homologue to UP|S25K_SOYBN (P10742) Stem 31 kDa glycopr... 47 8e-10
BM270825 30 1.7
CK606420 29 2.9
BE610568 28 3.7
TC227644 homologue to UP|Q9M4Z8 (Q9M4Z8) Beta-ketoacyl-ACP synth... 27 8.3
CF922281 27 8.3
AI960170 similar to SP|P78352|DLG4_ Presynaptic density protein ... 27 8.3
>TC234477 weakly similar to UP|DRPE_CRAPL (P22242) Desiccation-related
protein PCC13-62 precursor, partial (57%)
Length = 766
Score = 241 bits (615), Expect = 3e-64
Identities = 121/218 (55%), Positives = 162/218 (73%), Gaps = 13/218 (5%)
Frame = +1
Query: 81 LDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDP 140
LD + D+ QFAFQ+ GHLRAI +V+GFPRPLL++S ++FA+++++AFGKPL PPFDP
Sbjct: 16 LDSLTNDVILQFAFQEVGHLRAIKSKVRGFPRPLLDLSSKSFAKLMDNAFGKPLVPPFDP 195
Query: 141 YANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERR 200
YANSLN+++A+Y++PYVGLTGYVG N +L +R LV+GLLG +SGQD ++R LLYER+
Sbjct: 196 YANSLNFIIASYVIPYVGLTGYVGAN-RLLSATSRELVAGLLGVESGQDVVLR*LLYERK 372
Query: 201 YLQVKPYQWTVYEVTHRLSLLRNDLGKK-------------QAEGKVVGQVFGLNMYSLT 247
V PY V E T+R+S+LR+ LG + AEG+V G + ++ SL
Sbjct: 373 EQLVPPYGVAVEEFTNRISILRSKLGIRGLKDEGIVVPTGLGAEGRVRGNILAGDVNSLA 552
Query: 248 YPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSYL 285
Y R+PEEILRIVYGSGDEHV GGF+PKG +G+IA+ YL
Sbjct: 553 YSRTPEEILRIVYGSGDEHVRGGFYPKGASGHIAQCYL 666
>CA819108
Length = 474
Score = 192 bits (487), Expect = 2e-49
Identities = 95/158 (60%), Positives = 117/158 (73%), Gaps = 13/158 (8%)
Frame = +1
Query: 129 AFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQ 188
AFG+PL PPFDPYANS+NYLLA+Y++PYVGLTGYVG NP LQ ++ LV+GLLG +SGQ
Sbjct: 1 AFGRPLVPPFDPYANSINYLLASYVIPYVGLTGYVGANPLLQNATSKRLVAGLLGVESGQ 180
Query: 189 DAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGK-------------KQAEGKVV 235
DA+IRTLLYER+ V+PY+ TV E T R+S+LRN LG + AEG V
Sbjct: 181 DAVIRTLLYERQASLVQPYKVTVAEFTDRISMLRNKLGNAGVKDEGLVVPRVQGAEGSVT 360
Query: 236 GQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFP 273
+ + SL+YPR+PEEILRI+YG GDEHVPGGF+P
Sbjct: 361 DNILAGDKDSLSYPRTPEEILRIIYGGGDEHVPGGFYP 474
>BE806179
Length = 286
Score = 74.7 bits (182), Expect = 5e-14
Identities = 42/85 (49%), Positives = 54/85 (63%), Gaps = 8/85 (9%)
Frame = +2
Query: 1 MSLHTSSIFFLLSSLVLSLFI------PIFCS--FEDVVDVDLVQFHLNLEFLEAEFYFY 52
+S +SIF LL+SLVL L + +F S F D DL++F LNLE+LEAEF+ +
Sbjct: 32 ISRDRASIFVLLASLVLPLILLDYFSSALFASAKFPKSKDADLLEFALNLEYLEAEFFLF 211
Query: 53 GTLGWGLDHADPKLAQGGPPPVGGQ 77
G LG GLD A P L GGPPP+G +
Sbjct: 212 GALGHGLDVAAPNLTGGGPPPIGAK 286
>TC224991 homologue to UP|S25K_SOYBN (P10742) Stem 31 kDa glycoprotein
precursor (Vegetative storage protein VSP25) (Fragment),
partial (16%)
Length = 904
Score = 47.0 bits (110), Expect(2) = 8e-10
Identities = 25/50 (50%), Positives = 32/50 (64%)
Frame = +1
Query: 218 LSLLRNDLGKKQAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHV 267
LSL R + G + V G + + SL+YPR+P EILRI+YG GDEHV
Sbjct: 613 LSLPRAEFGTR-----VTGYILVGDKDSLSYPRTPREILRIIYGGGDEHV 747
Score = 33.5 bits (75), Expect(2) = 8e-10
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = +2
Query: 268 PGGFFPKGGNGNIAKSYL 285
PGGF+PKG +G IAK YL
Sbjct: 749 PGGFYPKGASGRIAKYYL 802
>BM270825
Length = 364
Score = 29.6 bits (65), Expect = 1.7
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 1/43 (2%)
Frame = +3
Query: 9 FFLLSSLVLSLFIPIFCSFEDVVD-VDLVQFHLNLEFLEAEFY 50
FF++S V +F FC +VD V +VQ H N FLE + +
Sbjct: 174 FFIMSFKVQYVFFVFFCHCISLVDQVYIVQVHQNKTFLEKKVH 302
>CK606420
Length = 498
Score = 28.9 bits (63), Expect = 2.9
Identities = 17/40 (42%), Positives = 20/40 (49%), Gaps = 3/40 (7%)
Frame = -3
Query: 41 NLEFLEAEFYFYGTLGWGLDH---ADPKLAQGGPPPVGGQ 77
+L F E E F G G G DH GGPPP+GG+
Sbjct: 370 SLGFSELEVIFQGG*GGGSDHPWGGGSWGPGGGPPPLGGE 251
>BE610568
Length = 413
Score = 28.5 bits (62), Expect = 3.7
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 11/59 (18%)
Frame = +1
Query: 3 LHTSSIFFLLSSLVLSLFIPIFCSFEDVVDVDLVQ--FH---------LNLEFLEAEFY 50
L +SSI+ + SL+ +F+ F+D + ++L++ FH L+ +F AEFY
Sbjct: 160 LFSSSIYIISLSLLFVIFLGSIFKFDDFLYLELLERVFHIIGVYA*SFLSCQFQNAEFY 336
>TC227644 homologue to UP|Q9M4Z8 (Q9M4Z8) Beta-ketoacyl-ACP synthetase 2,
complete
Length = 2440
Score = 27.3 bits (59), Expect = 8.3
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +3
Query: 156 YVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAM 191
Y L G NP+L+ T++++ LLGA G +A+
Sbjct: 1404 YQALMHCFGQNPELRVNSTKSMIGHLLGAAGGVEAV 1511
>CF922281
Length = 520
Score = 27.3 bits (59), Expect = 8.3
Identities = 11/22 (50%), Positives = 12/22 (54%)
Frame = +2
Query: 55 LGWGLDHADPKLAQGGPPPVGG 76
LGW + PK GGPPP G
Sbjct: 356 LGWEIKKKPPKKPPGGPPPQFG 421
>AI960170 similar to SP|P78352|DLG4_ Presynaptic density protein 95 (PSD-95)
(Discs large homolog 4) (Postsynaptic density-95).
[Human], partial (1%)
Length = 397
Score = 27.3 bits (59), Expect = 8.3
Identities = 11/18 (61%), Positives = 11/18 (61%)
Frame = -1
Query: 56 GWGLDHADPKLAQGGPPP 73
GWG PK QGGPPP
Sbjct: 340 GWGNFSLFPKFWQGGPPP 287
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,431,848
Number of Sequences: 63676
Number of extensions: 192930
Number of successful extensions: 968
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 961
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 964
length of query: 293
length of database: 12,639,632
effective HSP length: 97
effective length of query: 196
effective length of database: 6,463,060
effective search space: 1266759760
effective search space used: 1266759760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0333b.7


