

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0328.16
(738 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
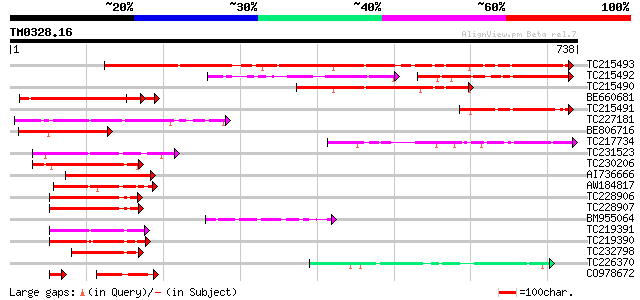
Score E
Sequences producing significant alignments: (bits) Value
TC215493 weakly similar to UP|DX_DROME (Q23985) Deltex protein, ... 830 0.0
TC215492 similar to UP|Q9VHB7 (Q9VHB7) CG8208-PA (LD22928p), par... 188 5e-97
TC215490 weakly similar to UP|Q8I7C3 (Q8I7C3) Lasp protein, part... 304 1e-82
BE660681 246 3e-65
TC215491 208 8e-54
TC227181 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, parti... 177 1e-44
BE806716 157 2e-38
TC217734 132 4e-31
TC231523 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {A... 130 3e-30
TC230206 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {A... 128 1e-29
AI736666 similar to GP|20197587|gb| expressed protein {Arabidops... 123 3e-28
AW184817 112 8e-25
TC228906 similar to UP|Q6ID88 (Q6ID88) At3g26510, partial (48%) 96 4e-20
TC228907 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {A... 96 6e-20
BM955064 96 6e-20
TC219391 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {A... 93 4e-19
TC219390 similar to GB|AAT41790.1|48310282|BT014807 At3g48240 {A... 92 6e-19
TC232798 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {A... 77 4e-14
TC226370 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pr... 68 1e-11
CO978672 54 2e-10
>TC215493 weakly similar to UP|DX_DROME (Q23985) Deltex protein, partial (6%)
Length = 2072
Score = 830 bits (2145), Expect = 0.0
Identities = 451/627 (71%), Positives = 499/627 (78%), Gaps = 16/627 (2%)
Frame = +3
Query: 124 SLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKS 183
SLISVTTDEDLENMIDEYDRT S++KPSRIRLFLFPTKPES HSIP QILD+SAKS
Sbjct: 3 SLISVTTDEDLENMIDEYDRTAAAATSAVKPSRIRLFLFPTKPESTHSIPAQILDTSAKS 182
Query: 184 DDWFLDALNGAGLLNRGFSDSASVNCLLGLDDVVAGNNNNIEQGGREGVE--AGGGGASL 241
DDWFL+ALNGAGLLNRGFSDSASVNCLLGLDD VAG NN+E G ++GV+ GGGGAS
Sbjct: 183 DDWFLNALNGAGLLNRGFSDSASVNCLLGLDDEVAG--NNLEPGSKDGVDGGVGGGGASQ 356
Query: 242 PGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALANLPPIRTHAEDGGGGGVRVQQ 301
GSF N KN KQDV S+PDSPMLET+SSFGSTSSSP+LANLPPIR H
Sbjct: 357 GGSFGNGKNLKQDVHSIPDSPMLETTSSFGSTSSSPSLANLPPIRVHV-----------V 503
Query: 302 QDQKVLGIEEQFAQLGVAAGVGVGQK---QDEGFVVITSPPPPPVPTTLSAPAMGVPINS 358
+DQKVLGIEEQFAQ+ GVGVGQK QDEGFV+++SPPPPPVP TL+ A+GVPI
Sbjct: 504 EDQKVLGIEEQFAQM----GVGVGQKQQQQDEGFVLLSSPPPPPVPATLA--AVGVPIGP 665
Query: 359 A-VVSGDYQNRVFSDDERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRS 417
A VV+G+Y NRV SDDERSDHGV VGYRKPPTP PQ Q+Q Q +Q Q+Q+ QFQQ+S
Sbjct: 666 ATVVAGEYHNRVVSDDERSDHGVSVGYRKPPTPPPQVQSQTQAQSQAQTQTLAPQFQQKS 845
Query: 418 SGG---TDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVS 474
+GG DLPSPDSVSSDSSLANA+SR KP +YQEQV IQSGTTRVP SNPVDPKLNVS
Sbjct: 846 TGGGAVVDLPSPDSVSSDSSLANAISRSKPAVYQEQVQIQSGTTRVP--SNPVDPKLNVS 1019
Query: 475 DPQSRIQVQQH-HEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQ 533
DP RIQ+QQH + GYLLQQQF+Q Q L Q Q QQQ +QQ HQ QQ
Sbjct: 1020DPHGRIQMQQHVQDPGYLLQQQFEQHQQL------QSQQPQQQFEQQHHQ----HQQTLP 1169
Query: 534 QQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQAN 593
QQQQQ+IHGT FIHHNP AIPAYYPVYPSQQ HPQH QVYYV ARQAQAYNL +QQAN
Sbjct: 1170QQQQQFIHGTHFIHHNP--AIPAYYPVYPSQQ--HPQHPQVYYVTARQAQAYNLPVQQAN 1337
Query: 594 MGE------SGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQ 647
MGE S +PQTPPNP+TLV Q A Y NP+RNAP+PKTEM A YR AT G+PQLVQ
Sbjct: 1338MGESAGNIASSRPQTPPNPSTLVQQPATY-NPIRNAPMPKTEMNA--YRAATAGNPQLVQ 1508
Query: 648 VPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYAYDYVDPAHAQIYYSQPLAPTMPSQ 707
VP + QHQQQYVTYSQIHHPSQS+APNSA PANYA+DY DPAHAQIYYSQP+APT+PSQ
Sbjct: 1509VPTS--QHQQQYVTYSQIHHPSQSMAPNSAAPANYAFDYADPAHAQIYYSQPMAPTIPSQ 1682
Query: 708 YQTMTAAAMVLPEGSSAQLPSDGMKQQ 734
YQTMTAAA+++ EG SAQ PSD +KQQ
Sbjct: 1683YQTMTAAAVMMQEG-SAQHPSDSVKQQ 1760
Score = 30.8 bits (68), Expect = 2.3
Identities = 18/41 (43%), Positives = 22/41 (52%), Gaps = 2/41 (4%)
Frame = +1
Query: 5 PQPPTPSAAP-PLSSVAPPPQPPPHLN-YADSVDSSPRSRN 43
P PPT ++ S APPP PPP N A + SSP + N
Sbjct: 13 PSPPTRTSRT*STSMTAPPPPPPPPSNPLASASSSSPPNPN 135
>TC215492 similar to UP|Q9VHB7 (Q9VHB7) CG8208-PA (LD22928p), partial (6%)
Length = 1424
Score = 188 bits (477), Expect(2) = 5e-97
Identities = 128/256 (50%), Positives = 150/256 (58%), Gaps = 6/256 (2%)
Frame = +1
Query: 258 VPDSPMLETSSSFGSTSSSPALANLPPIRTHAEDGGGGGVRVQQQDQKVLGIEEQFAQLG 317
V DSPMLE++SSFGS SS P NLPPI+ H E IEEQFAQLG
Sbjct: 31 VSDSPMLESNSSFGSGSSVPP--NLPPIKVHVE------------------IEEQFAQLG 150
Query: 318 VAAGVGVGQKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSD 377
V G Q++DEGFV PVP V NR+ SDDERSD
Sbjct: 151 V----GQKQQEDEGFVA-------PVP----------------VMEFSTNRLVSDDERSD 249
Query: 378 HGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGT-DLPSPDSVSSDSSLA 436
HGVP+G+RK PTP Q Q P QFQQ+S G DLPSPDS+SSDSSL+
Sbjct: 250 HGVPLGHRKSPTPPLQPQ--------------PHQFQQKSPHGVVDLPSPDSLSSDSSLS 387
Query: 437 NAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQH-HEQGYLLQQQ 495
NAM R KP IYQ+QV IQ G+++VP++++ VDPKLNV+D Q IQ+QQH E GY+LQ Q
Sbjct: 388 NAMLRPKPAIYQDQVQIQYGSSQVPSNAS-VDPKLNVTDQQGWIQMQQHVQEAGYVLQPQ 564
Query: 496 FDQ----QQHLLQQQF 507
FDQ QQH Q F
Sbjct: 565 FDQLPLPQQHQTQPPF 612
Score = 186 bits (471), Expect(2) = 5e-97
Identities = 115/213 (53%), Positives = 139/213 (64%), Gaps = 11/213 (5%)
Frame = +2
Query: 532 QQQQQQQYIHGTQFIHHNPANAIP---AYYPVYPSQQQTHPQHAQ-------VYYVPARQ 581
QQ Q QQYIH T IHH + ++P +YYPV QQQ H QH VYYV ARQ
Sbjct: 614 QQHQPQQYIHNTHLIHHTLSGSVPIPASYYPV--CQQQLHTQHLHHLDQQYPVYYVQARQ 787
Query: 582 AQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGG 641
AQ YNLS+Q AN ES P + + P +AAYN+ RN P EM A RTA
Sbjct: 788 AQPYNLSVQLANAAESATTM-PSSQSQNQPSSAAYNST-RN---PNFEMAAGACRTAPA- 949
Query: 642 SPQLVQVPANQLQHQQQYVTYSQIHHPS-QSIAPNSAPPANYAYDYVDPAHAQIYYSQPL 700
+PQLVQV +++ HQQQYV +SQI+H QS+ P SA PANYAY+Y DPA AQ++YSQPL
Sbjct: 950 TPQLVQVSSSK--HQQQYVAHSQIYHRHPQSMPPKSAVPANYAYNYADPALAQVFYSQPL 1123
Query: 701 APTMPSQYQTMTAAAMVLPEGSSAQLPSDGMKQ 733
AP MPS YQTMTAA+++ E SA+LPSD MK+
Sbjct: 1124APFMPSHYQTMTAASVMRTE-VSAKLPSDSMKK 1219
>TC215490 weakly similar to UP|Q8I7C3 (Q8I7C3) Lasp protein, partial (7%)
Length = 715
Score = 304 bits (778), Expect = 1e-82
Identities = 164/243 (67%), Positives = 180/243 (73%), Gaps = 13/243 (5%)
Frame = +3
Query: 374 ERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGG---TDLPSPDSVS 430
ERSDHGVPVGYRKPPTP PQ Q+Q Q +Q Q+Q+ QF Q+S+ G DLPSPDSVS
Sbjct: 3 ERSDHGVPVGYRKPPTPQPQVQSQTQAQSQAQTQTLAPQFHQKSTAGGAVVDLPSPDSVS 182
Query: 431 SDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQH-HEQG 489
SDSSLANA+SR K +YQEQ IQ GTTRVP SNPVDPKLNVSD RIQ+QQH H+ G
Sbjct: 183 SDSSLANAISRSKAAVYQEQAQIQPGTTRVP--SNPVDPKLNVSDLHGRIQMQQHVHDPG 356
Query: 490 YLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQ---QQQQQYIHGTQFI 546
YLLQQQF+Q Q L QQ Q + QQ Q QHQ +QHQ Q QQ QQQQQ+IHG FI
Sbjct: 357 YLLQQQFEQHQQLQSQQQQQFEQHQQLQSQPQHQFEQHQHQHQQTLPQQQQQFIHGAHFI 536
Query: 547 HHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGE------SGKP 600
HHNPA IPAYYPVYPSQQ HPQH QVYYV ARQAQAYNL +QQANMGE S +P
Sbjct: 537 HHNPA--IPAYYPVYPSQQ--HPQHPQVYYVTARQAQAYNLPLQQANMGESAGNIASSRP 704
Query: 601 QTP 603
QTP
Sbjct: 705 QTP 713
>BE660681
Length = 541
Score = 246 bits (628), Expect = 3e-65
Identities = 126/165 (76%), Positives = 134/165 (80%), Gaps = 1/165 (0%)
Frame = +3
Query: 14 PPLSSVAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFPPTS-KLRLMCSYGGHIVPRPH 72
PP S APPP PP HL+Y DSV+SSPRSRNTDSWD+PF P S KLRLMCSYGGHIVPRPH
Sbjct: 3 PPTPSTAPPPLPP-HLSYPDSVESSPRSRNTDSWDEPFAPASTKLRLMCSYGGHIVPRPH 179
Query: 73 DKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQLPNEDLDSLISVTTDE 132
DKSLCYVGGDTRI+V ERATSLAD+S RLSKTFL+GRPFTLKYQLPNEDLDSLISVTTDE
Sbjct: 180 DKSLCYVGGDTRIIVSERATSLADLSMRLSKTFLNGRPFTLKYQLPNEDLDSLISVTTDE 359
Query: 133 DLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQIL 177
DLENMIDEYDRT + P L LF TKPES P+ L
Sbjct: 360 DLENMIDEYDRTXASXPXX*NPXESAL-LFSTKPESLTLSLPKFL 491
Score = 57.4 bits (137), Expect = 2e-08
Identities = 29/43 (67%), Positives = 33/43 (76%)
Frame = +2
Query: 152 IKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWFLDALNGA 194
+KP RIR LF + HSIPPQILD+SAKSDDW L+ALNGA
Sbjct: 416 VKPXRIRSSLFH-QTRITHSIPPQILDTSAKSDDWVLNALNGA 541
>TC215491
Length = 647
Score = 208 bits (529), Expect = 8e-54
Identities = 111/155 (71%), Positives = 124/155 (79%), Gaps = 6/155 (3%)
Frame = +3
Query: 586 NLSMQQANMGESG------KPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTAT 639
NL +QQANMGES +PQTPPNP+TLV Q A YN P+RNAP+PKTEM A YR AT
Sbjct: 3 NLPVQQANMGESAGNIAXSRPQTPPNPSTLVQQPATYN-PIRNAPMPKTEMNA--YRAAT 173
Query: 640 GGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYAYDYVDPAHAQIYYSQP 699
G+PQLVQVP +Q HQQQYVTYSQIHHPSQS+APNSA PANYA+DY DPAHAQIYYSQP
Sbjct: 174 AGTPQLVQVPTSQ--HQQQYVTYSQIHHPSQSMAPNSAAPANYAFDYADPAHAQIYYSQP 347
Query: 700 LAPTMPSQYQTMTAAAMVLPEGSSAQLPSDGMKQQ 734
+APTMPSQYQT M++ EG SAQ PSD +KQQ
Sbjct: 348 MAPTMPSQYQT-----MMMQEG-SAQHPSDSVKQQ 434
Score = 33.5 bits (75), Expect = 0.35
Identities = 20/69 (28%), Positives = 31/69 (43%), Gaps = 4/69 (5%)
Frame = +3
Query: 529 QQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYY----VPARQAQA 584
Q Q QQQY+ +Q H + + A + P + P HAQ+YY P +Q
Sbjct: 195 QVPTSQHQQQYVTYSQIHHPSQSMAPNSAAPANYAFDYADPAHAQIYYSQPMAPTMPSQY 374
Query: 585 YNLSMQQAN 593
+ MQ+ +
Sbjct: 375 QTMMMQEGS 401
>TC227181 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, partial (38%)
Length = 1611
Score = 177 bits (450), Expect = 1e-44
Identities = 127/291 (43%), Positives = 164/291 (55%), Gaps = 10/291 (3%)
Frame = +3
Query: 7 PPTPSAAPPLSSVAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGH 66
PP P + + PP L+ ADSV SSPRS D + D P ++R MCS+GG
Sbjct: 33 PPNPHSKVIRRRMTSHSHYPPDLD-ADSVTSSPRS---DHFHDAPP---RIRFMCSFGGK 191
Query: 67 IVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRP-FTLKYQLPNEDLDSL 125
I+PRPHD L YVGGDTRIV V R+ + + + +LSK L G T KYQLPNEDLD+L
Sbjct: 192 ILPRPHDNQLRYVGGDTRIVAVNRSITFSALILKLSK--LSGMSNITAKYQLPNEDLDAL 365
Query: 126 ISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDD 185
ISVTTDED+ENM+DEYDR + + +R+RLFLFP +S S +L+ SAK ++
Sbjct: 366 ISVTTDEDVENMMDEYDRVAHN--QNPRSARLRLFLFPEGEDSRASSISSLLNGSAKREN 539
Query: 186 WFLDALN-GAGLLNRGFSDSASV-----NCLLGLDDVVAGNNNNIEQGGREGVEAGGGGA 239
WFLDALN G L RG S+++S+ + L GLD N+ I R+
Sbjct: 540 WFLDALNGGVSGLERGRSEASSMVSEVPDYLFGLD-----NSEEITNHQRD-------PR 683
Query: 240 SLPGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSS---PALANLPPIRT 287
PG S D P SP SS + STSS+ P+L NLP ++T
Sbjct: 684 PKPGPIIQDNVSVSD----PGSPAPVVSSPYCSTSSAPPVPSLPNLPYVKT 824
>BE806716
Length = 407
Score = 157 bits (396), Expect = 2e-38
Identities = 81/128 (63%), Positives = 99/128 (77%), Gaps = 5/128 (3%)
Frame = +1
Query: 12 AAPPLSSVAPPP-QPPPHLNYADSVDS-SPRSRNTDSWDD---PFPPTSKLRLMCSYGGH 66
+ PPL + PPP N+ +SV+S SP R ++W D P P +KLRLMCSYGGH
Sbjct: 22 STPPLLLMDPPPLSAATQKNHPESVESFSPCFRVAETWTDETLPAVPGAKLRLMCSYGGH 201
Query: 67 IVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQLPNEDLDSLI 126
I+PRPHDKSL Y+GGDTRIVVV+R +SL D+ +RLS+T L+GRPFTLKYQLPNEDL+SLI
Sbjct: 202 IMPRPHDKSLSYIGGDTRIVVVDRHSSLKDLCSRLSRTILNGRPFTLKYQLPNEDLESLI 381
Query: 127 SVTTDEDL 134
+VTTDEDL
Sbjct: 382 TVTTDEDL 405
>TC217734
Length = 1229
Score = 132 bits (333), Expect = 4e-31
Identities = 119/351 (33%), Positives = 166/351 (46%), Gaps = 26/351 (7%)
Frame = +2
Query: 414 QQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQV---LIQSGTTRVPTHSNPVDPK 470
Q R+SGG LPSPDSV+SDSS+A A S K V YQ+QV L+ + +P + K
Sbjct: 35 QPRTSGGVGLPSPDSVASDSSIAAANSFSKTVYYQDQVQAALLDNKIVAMP------NAK 196
Query: 471 LNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQ 530
+SD ++Q Q + GY L Q D +
Sbjct: 197 SEISDQMIQVQ-GQLQDSGYTLPPQLD------------------------------PNK 283
Query: 531 QQQQQQQQYIH-GTQFIHHNPANA---IPAYYPVY--PSQQQTHPQHAQVYYV------- 577
Q QQQ+ ++H T +IHH A + +YYPVY P QQ HP Q Y
Sbjct: 284 QHFQQQKPFVHASTHYIHHPAATGPVPVSSYYPVYAPPQPQQLHPPIGQQQYPVYVMPVG 463
Query: 578 PARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQ----TAAYNNPMRNAPLPKTEMTAA 633
P + Q YN+++ Q N+ + P + +L+PQ +AAY + P+ T+ ++
Sbjct: 464 PTQVTQPYNMAL-QPNIAD---PNVVASSRSLMPQSIVTSAAYKD--GTPPIYPTKSVSS 625
Query: 634 GYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSI--APNSAPP-ANYAYDYVDPA 690
+ +G +P +P NQ Q QYV Q HHP QSI AP+S NY Y+Y
Sbjct: 626 TMASNSGYAP----IPTNQF--QPQYVGLHQFHHPPQSIAVAPSSTTTNNNYGYEYGGHV 787
Query: 691 HAQIYYSQ--PLAPTMPSQYQTMT-AAAMVLPEGSSAQLPSDGMKQQTRPS 738
Q YY+Q AP +P QYQ+MT AAA +S P+D ++Q TR S
Sbjct: 788 QDQAYYTQQTTTAPLLP-QYQSMTPAAAAAALSDASK*FPADNIQQPTRAS 937
>TC231523 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;} , partial (32%)
Length = 878
Score = 130 bits (326), Expect = 3e-30
Identities = 84/205 (40%), Positives = 112/205 (53%), Gaps = 13/205 (6%)
Frame = +1
Query: 30 NYADSVDSSPRSRNTD-----SWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTR 84
+Y DS +SSPRSR D WD+ K + MCSYGG I PR HD L YVGGDT+
Sbjct: 37 SYPDSGESSPRSREIDFENPPPWDEQQNQNYKAKFMCSYGGKIQPRTHDNQLSYVGGDTK 216
Query: 85 IVVVERATSLADISTRLSKTFLDGRP--FTLKYQLPNEDLDSLISVTTDEDLENMIDEYD 142
I+ V+R+ ++L+ D P T KYQLP EDLD+LISVT D+DLE+M+ EYD
Sbjct: 217 ILAVDRSVKFPAFLSKLA-AVCDSAPQDLTFKYQLPGEDLDALISVTNDDDLEHMMHEYD 393
Query: 143 RTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWFLDALNGAGL------ 196
R ++KP R+RLFLF + +S S + D F++ALN +
Sbjct: 394 R---LYRPNLKPVRMRLFLFTLSNSNPNS-------SFSSERDRFVEALNSGPVPSQPDP 543
Query: 197 LNRGFSDSASVNCLLGLDDVVAGNN 221
+ ++V+ L GLD VA N
Sbjct: 544 IKTPPVTPSNVDYLFGLDKAVAPPN 618
>TC230206 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;} , partial (28%)
Length = 1305
Score = 128 bits (321), Expect = 1e-29
Identities = 74/154 (48%), Positives = 97/154 (62%), Gaps = 9/154 (5%)
Frame = +3
Query: 30 NYADSVDSSPRSRNTDSWDDPFP---PTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIV 86
+Y DS +SSPRSR D +D+P P K + MCSYGG I+PR HD L YV G+T+I+
Sbjct: 87 SYPDSGESSPRSREID-FDNPPPWDDQNYKAKFMCSYGGKILPRSHDNQLSYVAGETKIL 263
Query: 87 VVERATSLADISTRLSKTF--LDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRT 144
V+R+ + + +LS D T KYQLP EDLD+LISVT D+DL++M+ EYDR
Sbjct: 264 AVDRSIKFSAMLAKLSALCDAPDNNNLTFKYQLPGEDLDALISVTNDDDLDHMMHEYDRL 443
Query: 145 TTTTASSIKPSRIRLFLFPT----KPESAHSIPP 174
+A +PSR+RLFLFP+ P A S PP
Sbjct: 444 YRASA---RPSRMRLFLFPSDTLPSPPPAASAPP 536
Score = 29.3 bits (64), Expect = 6.6
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = +1
Query: 10 PSAAPPLSSVAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFPPTS 55
PS++PP S +PP P + S S P +T S P TS
Sbjct: 277 PSSSPPCSPNSPPSATPQTTTTSPSSTSFPAKTSTPSSPSPTTTTS 414
>AI736666 similar to GP|20197587|gb| expressed protein {Arabidopsis
thaliana}, partial (6%)
Length = 357
Score = 123 bits (308), Expect = 3e-28
Identities = 69/118 (58%), Positives = 80/118 (67%), Gaps = 1/118 (0%)
Frame = +3
Query: 73 DKSLCYVGGDTRIVVVERATSLADISTR-LSKTFLDGRPFTLKYQLPNEDLDSLISVTTD 131
+K + YVGGDTRI+ ERATSL D+ST ++ P EDL+ LI VTTD
Sbjct: 3 EKYV*YVGGDTRIIGCERATSLWDLSTAPINNIA*MEDP*RSSTSCATEDLECLIYVTTD 182
Query: 132 EDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWFLD 189
EDLENMI+EY+RT KP RIRLFLF TKPES HSIP QILD+ AKSDDWFL+
Sbjct: 183 EDLENMIEEYERTACVATCCDKPCRIRLFLFRTKPESTHSIPAQILDT*AKSDDWFLN 356
>AW184817
Length = 455
Score = 112 bits (279), Expect = 8e-25
Identities = 64/138 (46%), Positives = 85/138 (61%), Gaps = 3/138 (2%)
Frame = +3
Query: 58 RLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTL---K 114
+LMCS+GG I PRPHD L YV GDT+I+ V+R + +LS + + P L K
Sbjct: 3 KLMCSFGGSIQPRPHDNHLTYVSGDTKILAVDRHVKFPSLIAKLS-SLANNTPSNLSFFK 179
Query: 115 YQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPP 174
YQLP EDLD+LISVT D+DL M+ EYDR + +S +P+R+RLFLFP + + P
Sbjct: 180 YQLPGEDLDALISVTNDDDLHQMMIEYDR---LSRASPRPARLRLFLFPLH-NNCNFAPT 347
Query: 175 QILDSSAKSDDWFLDALN 192
+ S WF+DALN
Sbjct: 348 E----SKSERQWFVDALN 389
>TC228906 similar to UP|Q6ID88 (Q6ID88) At3g26510, partial (48%)
Length = 927
Score = 96.3 bits (238), Expect = 4e-20
Identities = 54/122 (44%), Positives = 77/122 (62%)
Frame = +2
Query: 52 PPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPF 111
PP S L+ +CSYGG I+PR D L Y+GG TRI+ V+R+ +++ +L +
Sbjct: 140 PPKSTLKFLCSYGGKILPRYPDGKLRYLGGHTRILAVDRSIPFSELLLKLEE-LCGASVR 316
Query: 112 TLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHS 171
L+ QLP+EDLD+L+S+T+DEDL N+I+EYDR SS+K IR FL P P + +
Sbjct: 317 HLRCQLPSEDLDALVSITSDEDLANLIEEYDR-----VSSLK---IRAFLSPPTPPTPPT 472
Query: 172 IP 173
P
Sbjct: 473 PP 478
>TC228907 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {Arabidopsis
thaliana;} , partial (53%)
Length = 537
Score = 95.9 bits (237), Expect = 6e-20
Identities = 55/124 (44%), Positives = 78/124 (62%), Gaps = 1/124 (0%)
Frame = +1
Query: 52 PPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPF 111
PP S L+ +CSYGG I+PR D L Y+GG TR++ V+R+ +++ +L +
Sbjct: 163 PPKSTLKFLCSYGGKILPRYPDGKLRYLGGHTRVLAVDRSIPFSELLLKLEE-LCGASVR 339
Query: 112 TLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTK-PESAH 170
L+ QLP+EDLD+L+S+T+DEDL N+I+EYDR SS+K IR FL P + P
Sbjct: 340 YLRCQLPSEDLDALVSITSDEDLANLIEEYDR-----VSSLK---IRAFLSPPRSPSLKV 495
Query: 171 SIPP 174
S PP
Sbjct: 496 STPP 507
>BM955064
Length = 425
Score = 95.9 bits (237), Expect = 6e-20
Identities = 69/173 (39%), Positives = 93/173 (52%), Gaps = 2/173 (1%)
Frame = +1
Query: 255 VSSVPDSPMLETSSSFGSTSSSPALANLPPIRTHAEDGGGGGVRVQQQDQKVLGIEEQFA 314
V S P SPM+E +SS S S SP+LANLPPIR +D G R+ Q+++ + +EEQFA
Sbjct: 10 VQSTPGSPMMENNSS-SSPSFSPSLANLPPIRVRVDDNGS---RLHQENKVGMVVEEQFA 177
Query: 315 QLGVAAGVGVGQKQDEGFVVITSP--PPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSD 372
Q+ +A+GV K D+G V + S P P +P ++ ++GV S + NRV +
Sbjct: 178 QMTIASGV----KPDDGLVNVVSSTVPVPVIPAAVTMASVGV----ITTSDNVMNRVVYE 333
Query: 373 DERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPS 425
DERSD G+RKPP P QL Q R+SGG LPS
Sbjct: 334 DERSD----PGFRKPPLP-------MQL------------VQPRTSGGVGLPS 423
>TC219391 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {Arabidopsis
thaliana;} , partial (44%)
Length = 718
Score = 93.2 bits (230), Expect = 4e-19
Identities = 52/129 (40%), Positives = 74/129 (57%)
Frame = +1
Query: 53 PTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFT 112
P ++L+CSYGG I+PR D L Y GG TR++ V R+ S +++ +LS+ G
Sbjct: 136 PGQTIKLLCSYGGKILPRATDGELRYAGGHTRVLTVARSISFSELMVKLSE--FCGSSVI 309
Query: 113 LKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSI 172
L+ QLP DL++LIS+T DEDL ++I+EYDR + A P +I+ L P K
Sbjct: 310 LRCQLPKGDLETLISITNDEDLASIIEEYDRASLKLA---HPLKIKAVLSPPKSSKKTPS 480
Query: 173 PPQILDSSA 181
P SSA
Sbjct: 481 APSSASSSA 507
>TC219390 similar to GB|AAT41790.1|48310282|BT014807 At3g48240 {Arabidopsis
thaliana;} , partial (53%)
Length = 845
Score = 92.4 bits (228), Expect = 6e-19
Identities = 54/131 (41%), Positives = 79/131 (60%)
Frame = +1
Query: 53 PTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFT 112
P ++L+CSYGG I+PR D L YVGG TR++ V+R+ S ++ +L + F G
Sbjct: 142 PAQTIKLLCSYGGKILPRATDGELRYVGGHTRVLTVDRSISFPELMVKL-RVFC-GSSVI 315
Query: 113 LKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSI 172
L+ QLP DL++LIS+T DEDL ++I+EYDR++ A P +I+ L P P+S
Sbjct: 316 LRCQLPKGDLETLISITNDEDLASIIEEYDRSSLKLA---HPLKIKAVLSP--PKSMKKA 480
Query: 173 PPQILDSSAKS 183
P +SA S
Sbjct: 481 SPVPSSASASS 513
>TC232798 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;} , partial (10%)
Length = 707
Score = 76.6 bits (187), Expect = 4e-14
Identities = 44/95 (46%), Positives = 61/95 (63%), Gaps = 1/95 (1%)
Frame = +1
Query: 81 GDTRIVVVERATSLADISTRLSKTF-LDGRPFTLKYQLPNEDLDSLISVTTDEDLENMID 139
GDT+I+ V+R+ + +LS T KYQLP EDLD+LISVT D+DL++M+
Sbjct: 1 GDTKILAVDRSIKFPAMLAKLSALCDAQDNNITFKYQLPGEDLDALISVTNDDDLDHMMH 180
Query: 140 EYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPP 174
EYDR +A +PSR+RLFLFP+ ++ S PP
Sbjct: 181 EYDRLYRASA---RPSRMRLFLFPS--DTPPSPPP 270
>TC226370 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (43%)
Length = 1810
Score = 68.2 bits (165), Expect = 1e-11
Identities = 89/337 (26%), Positives = 128/337 (37%), Gaps = 18/337 (5%)
Frame = +2
Query: 391 TPQQQAQQQL-HTQTQSQSHPAQFQQRSSGGTDLPSPDSVS--SDSSLANAMSRQ----- 442
T ++ A+ QL ++ S SH ++RSS TD D+ S ++ LA A+ Q
Sbjct: 455 TQKELAKLQLAQKESSSSSHSQSNEERSSPTTDPKKTDNASDANNQQLALALPHQIAPQP 634
Query: 443 KPVIYQEQVLIQS---GTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYL-LQQQFDQ 498
+P Q Q Q+ T+VP P + +P V QH + YL QQ+
Sbjct: 635 QPPAPQVQAQAQAPAPNVTQVPQQPPYYMPPTPLPNPA----VPQHPQNQYLPSDQQYRT 802
Query: 499 QQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYY 558
QH+ Q Q Q +QQQQ Q QQQQ QQQQ +P+
Sbjct: 803 PQHVAPQPTPSQVTPSPVQQFSHYQQQQQQPQQQQPQQQQ---------QWSQQVLPSQP 955
Query: 559 PVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNN 618
P P Q Q P V Y P + QA N P+P +P + A
Sbjct: 956 P--PMQSQVRPPSPNV-YPPYQPNQATN-----------------PSPAETLPNSMAMQM 1075
Query: 619 PMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAP 678
P P P + T A G + Q P Q + ++ PS ++ + P
Sbjct: 1076PYSGVPPPGSSRTDAIPYGYGGAGRTVPQQPPPQQMKSSFPAPPADMYGPSGNLP--ALP 1249
Query: 679 PANYAYDYVDPAHA------QIYYSQPLAPTMPSQYQ 709
P + AY D Q +++QP P + Q
Sbjct: 1250PPSGAYMMYDGEGGRTHHPPQPHFAQPGYPPTSASLQ 1360
>CO978672
Length = 725
Score = 53.5 bits (127), Expect(2) = 2e-10
Identities = 33/82 (40%), Positives = 50/82 (60%), Gaps = 1/82 (1%)
Frame = -3
Query: 113 LKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLF-PTKPESAHS 171
LKYQL EDLD+L+SV T+ED+++MI+E+DR T + +R FLF P+K +
Sbjct: 570 LKYQLIPEDLDALVSVRTEEDVKHMIEEHDRHHT-------GALLRAFLFPPSKQTGLVA 412
Query: 172 IPPQILDSSAKSDDWFLDALNG 193
P +L+ ++DA+NG
Sbjct: 411 CEPYLLEQR------YIDAING 364
Score = 30.4 bits (67), Expect(2) = 2e-10
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = -2
Query: 53 PTSKLRLMCSYGGHIVPRPHD 73
P +K++ +CSYGG ++PR D
Sbjct: 694 PRNKVKFLCSYGGKVLPRXXD 632
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.127 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,141,173
Number of Sequences: 63676
Number of extensions: 710089
Number of successful extensions: 27357
Number of sequences better than 10.0: 906
Number of HSP's better than 10.0 without gapping: 9177
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 19099
length of query: 738
length of database: 12,639,632
effective HSP length: 104
effective length of query: 634
effective length of database: 6,017,328
effective search space: 3814985952
effective search space used: 3814985952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0328.16


