

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0323a.10
(175 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
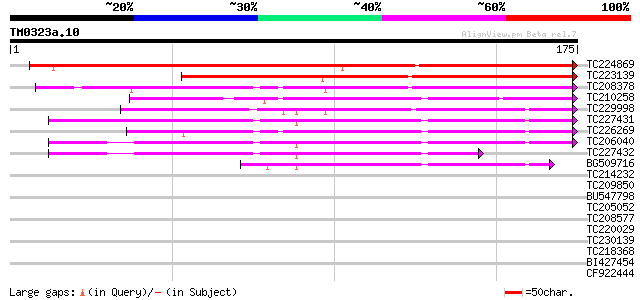
TC224869 weakly similar to UP|ST14_SOLTU (Q41495) STS14 protein ... 178 1e-45
TC223139 weakly similar to UP|Q8LDA3 (Q8LDA3) Sts14, partial (59%) 142 6e-35
TC208378 similar to UP|PR1_MEDTR (Q40374) Pathogenesis-related p... 114 2e-26
TC210258 weakly similar to UP|Q9LPM7 (Q9LPM7) F2J10.6 protein, p... 107 3e-24
TC229998 weakly similar to PIR|T04232|T04232 pathogenesis-relate... 103 4e-23
TC227431 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (... 100 3e-22
TC226269 homologue to UP|Q9XFB4 (Q9XFB4) PR1a precursor, complete 100 6e-22
TC206040 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (... 99 8e-22
TC227432 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (... 76 9e-15
BG509716 similar to GP|4928711|gb|A PR1a precursor {Glycine max}... 48 2e-06
TC214232 weakly similar to UP|ATF5_HUMAN (Q9Y2D1) Cyclic-AMP-dep... 33 0.090
TC209850 weakly similar to UP|Q947C2 (Q947C2) P450 monooxygenase... 30 0.45
BU547798 30 0.45
TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 30 0.76
TC208577 similar to UP|Q9SZ43 (Q9SZ43) SNF8 like protein, complete 27 3.8
TC220029 27 6.4
TC230139 UP|Q95RL3 (Q95RL3) LD21089p, partial (8%) 27 6.4
TC218368 similar to UP|GPDA_CUPLA (P52425) Glycerol-3-phosphate ... 26 8.4
BI427454 26 8.4
CF922444 26 8.4
>TC224869 weakly similar to UP|ST14_SOLTU (Q41495) STS14 protein precursor,
partial (65%)
Length = 1276
Score = 178 bits (452), Expect = 1e-45
Identities = 92/171 (53%), Positives = 114/171 (65%), Gaps = 2/171 (1%)
Frame = +3
Query: 7 PFFSLA-LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKD 65
P FSLA LA FL+ AAT ++ P L+A A+EFL+AHN ARAEVGVE L WSE L
Sbjct: 579 PLFSLASLATFLVLTHAATAPENPPPPLTAAAREFLEAHNQARAEVGVEALSWSEKLGNV 758
Query: 66 AGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWT-PSVAVQDWVNEKEFYSYRTNSCV 124
+ L RYQR++ GC+ AN + ++YG NQ G+ P V V++WV EK+FY N+CV
Sbjct: 759 SSLMVRYQRNKKGCEFANLTASRYGGNQLWAGVTEVAPRVVVEEWVKEKKFYVRENNTCV 938
Query: 125 APSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C YTQVVWR S +VGCAQ CV ++ LTICFYDPPGN+ GE PY
Sbjct: 939 G-KHECGVYTQVVWRNSTEVGCAQAVCVKEQASLTICFYDPPGNVIGEIPY 1088
>TC223139 weakly similar to UP|Q8LDA3 (Q8LDA3) Sts14, partial (59%)
Length = 634
Score = 142 bits (359), Expect = 6e-35
Identities = 69/124 (55%), Positives = 86/124 (68%), Gaps = 2/124 (1%)
Frame = +2
Query: 54 EPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDL--GGMGWTPSVAVQDWVN 111
EPL WSE +A ARYQR + GC+ AN + KYG+NQ L G TP +AV++WV
Sbjct: 2 EPLRWSEQVANVTSKLARYQRVKTGCQFANLTAGKYGANQLLARGSAAVTPRMAVEEWVK 181
Query: 112 EKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNING 171
+K+FY + NSC AP+ C YTQVVWRKS ++GCAQ TCV ++ LTICFY+PPGN G
Sbjct: 182 QKQFYFHADNSC-APNHRCGVYTQVVWRKSVELGCAQATCVKEQASLTICFYNPPGNYVG 358
Query: 172 ESPY 175
ESPY
Sbjct: 359 ESPY 370
>TC208378 similar to UP|PR1_MEDTR (Q40374) Pathogenesis-related protein PR-1
precursor, partial (84%)
Length = 779
Score = 114 bits (285), Expect = 2e-26
Identities = 71/173 (41%), Positives = 93/173 (53%), Gaps = 6/173 (3%)
Frame = +1
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAE-----AKEFLDAHNWARAEVGVEPLEWSEALA 63
F ALFLL AT +VVP + + A +FL N ARA + + PL W LA
Sbjct: 61 FLSCFALFLLLV--ATTYATVVPTTTQKPPRSFANQFLIPQNAARAVLRLRPLVWDSKLA 234
Query: 64 KDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNS 122
A +A +R+ C L + S+ YG N G G GW P+ AV WV E+++Y+Y NS
Sbjct: 235 HYAQWYANQRRN--DCALEH-SNGPYGENIFWGSGTGWKPAQAVSAWVEERQWYNYWHNS 405
Query: 123 CVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C A C HYTQ+VW ++++GCA + C K C YDPPGN GE PY
Sbjct: 406 C-ANGQMCGHYTQIVWSTTRKIGCASVVCSGGKGTFMTCNYDPPGNYYGERPY 561
>TC210258 weakly similar to UP|Q9LPM7 (Q9LPM7) F2J10.6 protein, partial (66%)
Length = 608
Score = 107 bits (267), Expect = 3e-24
Identities = 61/139 (43%), Positives = 79/139 (55%), Gaps = 1/139 (0%)
Frame = +1
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRL-GCKLANFSHNKYGSNQDLG 96
++FLD HN ARAEVGV PL W+ L A RY +R+ C L + S +G N G
Sbjct: 139 QDFLDVHNQARAEVGVGPLSWNHTLQAYA---QRYANERIPDCNLEH-SMGPFGENLAEG 306
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
S AV+ W+ EK +Y + +N+CV C HYTQ+VWR S +GCA+ C +
Sbjct: 307 YGEMKGSDAVKFWLTEKPYYDHYSNACVHDE--CLHYTQIVWRDSVHLGCARAKC-NNGW 477
Query: 157 ILTICFYDPPGNINGESPY 175
+ IC Y PPGNI GE PY
Sbjct: 478 VFVICSYSPPGNIEGERPY 534
>TC229998 weakly similar to PIR|T04232|T04232 pathogenesis-related protein
homolog F14M19.60 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (78%)
Length = 915
Score = 103 bits (257), Expect = 4e-23
Identities = 61/145 (42%), Positives = 85/145 (58%), Gaps = 4/145 (2%)
Frame = +1
Query: 35 AEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLAN-FSHN--KYGS 91
AE+ EFL HN RA PL W L + A +A ++ CKL + F + K G
Sbjct: 184 AESLEFLFRHNLVRAAKWELPLMWDFQLEQYARWWAGERK--ADCKLEHSFPEDGFKLGE 357
Query: 92 NQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLT 150
N G G WTPS AV+ W +E+++Y+Y TN+CV P C HYTQ+VW+ ++++GCA++
Sbjct: 358 NIYWGSGSAWTPSDAVRAWADEEKYYTYATNTCV-PGQMCGHYTQIVWKSTRRIGCARVV 534
Query: 151 CVVDKLILTICFYDPPGNINGESPY 175
C + +T C YDP GN GE PY
Sbjct: 535 CDDGDVFMT-CNYDPVGNYVGERPY 606
>TC227431 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (90%)
Length = 707
Score = 100 bits (250), Expect = 3e-22
Identities = 61/164 (37%), Positives = 87/164 (52%), Gaps = 1/164 (0%)
Frame = +2
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
++L LL ++ V ++++AHN AR++VGV + W A+A A +A
Sbjct: 74 ISLCLLCVLGLVIVGDHVAYAQDSPTDYVNAHNAARSQVGVPNIVWDNAVAAFAQNYANQ 253
Query: 73 QRDRLGCKLANFSHN-KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCW 131
++ CKL + + KYG N + AVQ WVNEK Y+Y +NSCV C
Sbjct: 254 RKG--DCKLVHSGGDGKYGENLAGSTGNLSGKDAVQLWVNEKSKYNYNSNSCVGGE--CL 421
Query: 132 HYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S ++GCA++ C + C Y PPGN G+ PY
Sbjct: 422 HYTQVVWRNSLRLGCAKVRCNNGGTFIG-CNYAPPGNYIGQRPY 550
>TC226269 homologue to UP|Q9XFB4 (Q9XFB4) PR1a precursor, complete
Length = 771
Score = 99.8 bits (247), Expect = 6e-22
Identities = 60/148 (40%), Positives = 84/148 (56%), Gaps = 9/148 (6%)
Frame = +3
Query: 37 AKEFLDAHNWARAEVG---------VEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHN 87
A+++++AHN ARAEVG V L W + +A A +A ++ C+L + S
Sbjct: 171 AEDYVNAHNAARAEVGSQSPRQTVIVPSLAWDDTVAAYAESYANQRKG--DCQLIH-SGG 341
Query: 88 KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCA 147
+YG N + + + AV+ WV+EK Y Y +NSCV C HYTQVVW S ++GCA
Sbjct: 342 EYGENIAMSTGELSGTDAVKMWVDEKSNYDYDSNSCVGGE--CLHYTQVVWANSVRLGCA 515
Query: 148 QLTCVVDKLILTICFYDPPGNINGESPY 175
++TC +T C YDPPGN GE PY
Sbjct: 516 KVTCDNGGTFIT-CNYDPPGNFVGERPY 596
>TC206040 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (90%)
Length = 772
Score = 99.4 bits (246), Expect = 8e-22
Identities = 60/164 (36%), Positives = 89/164 (53%), Gaps = 1/164 (0%)
Frame = +3
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
L L ++ + A DS ++++AHN AR+EVGV+ L W + +A A +A
Sbjct: 126 LGLVMIVSHVANAQDSPA--------DYVNAHNAARSEVGVQNLAWDDTVAAFAQNYANQ 281
Query: 73 QRDRLGCKLANFSHN-KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCW 131
++ C+L + +YG N + + + AV+ WV+EK Y Y +NSCV C
Sbjct: 282 RKG--DCQLIHSGGGGQYGENLAMSTGDLSGTDAVKLWVDEKSNYDYNSNSCVGGE--CL 449
Query: 132 HYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S ++GCA++ C +T C Y PPGN G+ PY
Sbjct: 450 HYTQVVWRDSVRLGCAKVACDNGGTFIT-CNYAPPGNYVGQRPY 578
>TC227432 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (75%)
Length = 448
Score = 75.9 bits (185), Expect = 9e-15
Identities = 48/135 (35%), Positives = 72/135 (52%), Gaps = 1/135 (0%)
Frame = +2
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
L L ++ AA DS +F++AHN AR++VGV + W + +A A +A
Sbjct: 77 LGLVIVGDHAAYAQDSPT--------DFVNAHNAARSQVGVPNIVWDDTVAAFAQNYANQ 232
Query: 73 QRDRLGCKLANFSHN-KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCW 131
++ CKL + + KYG N + + AV+ WV+EK Y Y +N+CV C
Sbjct: 233 RKG--DCKLVHSGGDGKYGENLAGSTGNLSGTNAVKLWVDEKSKYDYNSNTCVGGE--CR 400
Query: 132 HYTQVVWRKSKQVGC 146
HYTQVVW+ S ++GC
Sbjct: 401 HYTQVVWKNSVRLGC 445
>BG509716 similar to GP|4928711|gb|A PR1a precursor {Glycine max}, partial
(54%)
Length = 294
Score = 48.1 bits (113), Expect = 2e-06
Identities = 36/99 (36%), Positives = 48/99 (48%), Gaps = 2/99 (2%)
Frame = +3
Query: 72 YQRDRLG-CKLANFSHN-KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFP 129
Y R G C+L + +YG N + + + AV+ +EK Y Y +NSCV
Sbjct: 6 YANQRKGDCQLIHSGGGGQYGENLAMSTGDLSGTDAVRGGGDEKSNYDYNSNSCVGGE-- 179
Query: 130 CWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGN 168
C HYTQVVWR S + G A+ +T Y PPGN
Sbjct: 180 CLHYTQVVWRDSVRXGXAKXAWENGGPXIT-GNYAPPGN 293
>TC214232 weakly similar to UP|ATF5_HUMAN (Q9Y2D1) Cyclic-AMP-dependent
transcription factor ATF-5 (Activating transcription
factor 5) (Transcription factor ATFx), partial (8%)
Length = 1283
Score = 32.7 bits (73), Expect = 0.090
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +2
Query: 109 WVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC 146
W E+ F+ Y C PC HY+ + R SK VGC
Sbjct: 605 WTQERCFFEYHQKDCQV-LMPC*HYSSIY*R*SK*VGC 715
>TC209850 weakly similar to UP|Q947C2 (Q947C2) P450 monooxygenase, partial
(10%)
Length = 931
Score = 30.4 bits (67), Expect = 0.45
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = -1
Query: 56 LEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
LE+ L+K + L + + R+GCKL H+KYG
Sbjct: 151 LEFLNHLSKHSTLISVTRMVRIGCKLVESQHHKYG 47
>BU547798
Length = 687
Score = 30.4 bits (67), Expect = 0.45
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = -2
Query: 56 LEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
LE+ L+K + L + + R+GCKL H+KYG
Sbjct: 419 LEFLNHLSKHSTLISVTRMVRIGCKLVESQHHKYG 315
>TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (58%)
Length = 953
Score = 29.6 bits (65), Expect = 0.76
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = -1
Query: 90 GSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
GS ++LGG G PSVA + W+ S+R ++ SF C
Sbjct: 899 GSKKELGGGGGPPSVAFKKWM-----ISFRDSNQWLSSFTC 792
>TC208577 similar to UP|Q9SZ43 (Q9SZ43) SNF8 like protein, complete
Length = 778
Score = 27.3 bits (59), Expect = 3.8
Identities = 21/59 (35%), Positives = 28/59 (46%), Gaps = 3/59 (5%)
Frame = +1
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLD---AHNWARAEVGVEPLEWSEALAKDA 66
+ L LFLL E ++ SV P+L+ EFLD A + + L WS A DA
Sbjct: 256 VVLKLFLL--ERKKLVRSVPPELTKAPNEFLDLAQAQGFVPVDEVERRLSWSSGRAIDA 426
>TC220029
Length = 1096
Score = 26.6 bits (57), Expect = 6.4
Identities = 28/121 (23%), Positives = 47/121 (38%), Gaps = 11/121 (9%)
Frame = +1
Query: 10 SLALALFLLSAE-----AATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAK 64
S + + +LSAE ++ S + ++ A +E G P W+E +
Sbjct: 1 SRTVEMTILSAENLQMNRKSIRGSTFVTVQSDTSSDTSAATKVDSEGGSYP-SWNEKVVM 177
Query: 65 DAGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWV------NEKEFYSY 118
D L AR+ + CK ++ N G Q + V D++ N+ F SY
Sbjct: 178 DVPLHARFITVEVKCKTSSSGSNSVGVAQ----------IPVSDFIGGYMPENQLHFLSY 327
Query: 119 R 119
R
Sbjct: 328 R 330
>TC230139 UP|Q95RL3 (Q95RL3) LD21089p, partial (8%)
Length = 731
Score = 26.6 bits (57), Expect = 6.4
Identities = 19/68 (27%), Positives = 27/68 (38%), Gaps = 6/68 (8%)
Frame = +2
Query: 58 WSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWV------N 111
W+E L D L AR+ + CK ++ N G + V D+V N
Sbjct: 323 WNEKLVMDVPLHARFITVEVKCKTSSAGSNSIG----------VARIPVSDFVGGYVPEN 472
Query: 112 EKEFYSYR 119
+ F SYR
Sbjct: 473 QLHFLSYR 496
>TC218368 similar to UP|GPDA_CUPLA (P52425) Glycerol-3-phosphate
dehydrogenase [NAD+] , partial (92%)
Length = 1633
Score = 26.2 bits (56), Expect = 8.4
Identities = 14/39 (35%), Positives = 18/39 (45%)
Frame = -1
Query: 7 PFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHN 45
PF + A A FL V+ S PQLS + D +N
Sbjct: 892 PFLA*ASATFLFLPPKHVVIRSATPQLSKKVLSLTDGNN 776
>BI427454
Length = 427
Score = 26.2 bits (56), Expect = 8.4
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = +2
Query: 100 WTPSVAVQDWVNEKEFYS 117
W ++A+ DW+N+K YS
Sbjct: 335 WQRALALLDWINDKALYS 388
>CF922444
Length = 714
Score = 26.2 bits (56), Expect = 8.4
Identities = 14/49 (28%), Positives = 23/49 (46%)
Frame = -2
Query: 126 PSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESP 174
P P + T+ + +++ V C+V K +C P GNI+ SP
Sbjct: 527 PQLPIFP*TKYISKRTLGV------CIVKKTFTCVCSSTPNGNIHTSSP 399
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,579,506
Number of Sequences: 63676
Number of extensions: 149320
Number of successful extensions: 720
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 697
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 698
length of query: 175
length of database: 12,639,632
effective HSP length: 91
effective length of query: 84
effective length of database: 6,845,116
effective search space: 574989744
effective search space used: 574989744
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0323a.10


