

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.5
(210 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
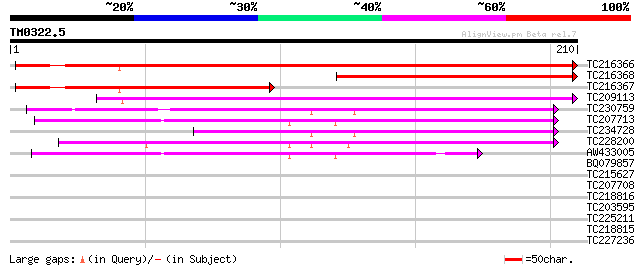
Score E
Sequences producing significant alignments: (bits) Value
TC216366 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane p... 317 2e-87
TC216368 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane p... 150 5e-37
TC216367 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane p... 124 4e-29
TC209113 similar to UP|Q6YZD2 (Q6YZD2) Coated vesicle membrane p... 119 1e-27
TC230759 similar to UP|Q6ZGK3 (Q6ZGK3) Emp24/gp25L/p24-like, par... 73 1e-13
TC207713 similar to UP|Q6IDL4 (Q6IDL4) At1g09580, partial (75%) 70 5e-13
TC234728 similar to UP|Q6ZGK3 (Q6ZGK3) Emp24/gp25L/p24-like, par... 60 7e-10
TC228200 53 9e-08
AW433005 similar to GP|21387055|gb| transmembrane protein-like p... 49 2e-06
BQ079857 similar to GP|22296377|dbj P0519E12.19 {Oryza sativa (j... 40 0.001
TC215627 similar to UP|Q9LNJ1 (Q9LNJ1) F6F3.12 protein, partial ... 31 0.36
TC207708 similar to UP|Q9LK35 (Q9LK35) Receptor-protein kinase-l... 28 2.3
TC218816 similar to UP|Q8GVB6 (Q8GVB6) ACC oxidase, partial (86%) 28 3.0
TC203595 similar to UP|Q8L7T3 (Q8L7T3) Ubiquitin-conjugating enz... 27 5.2
TC225211 homologue to UP|PRP2_SOYBN (P13993) Repetitive proline-... 27 6.7
TC218815 similar to UP|Q9ZUN4 (Q9ZUN4) 1-aminocyclopropane-1-car... 27 6.7
TC227236 weakly similar to UP|Q7PMV5 (Q7PMV5) ENSANGP00000017545... 27 8.8
>TC216366 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane protein-like
(AT3g22845/MWI23_22), partial (89%)
Length = 1173
Score = 317 bits (813), Expect = 2e-87
Identities = 154/209 (73%), Positives = 175/209 (83%), Gaps = 1/209 (0%)
Frame = +3
Query: 3 WFSKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHA-QSGDSVTGNFVVMDHDIFWSSD 61
W +C V FF + SLS+++NDVECV E+ GD+V+GNFVV+DHDIFW+SD
Sbjct: 201 WMVFMCLTVSFFA-----RIESLSVTVNDVECVYEYVLYEGDTVSGNFVVVDHDIFWNSD 365
Query: 62 HPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHI 121
HPGIDFTVT+ GN VH++KGTSGDKF FKAP +G+YKFCFHNP STPETVSFYIHVGHI
Sbjct: 366 HPGIDFTVTSAAGNTVHNIKGTSGDKFSFKAPTHGMYKFCFHNPYSTPETVSFYIHVGHI 545
Query: 122 PSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVL 181
PSEHDLAKDEHLDPINVKIAELREALES+ EQKYLKARD RHR+TNEST+ RV+FYTV
Sbjct: 546 PSEHDLAKDEHLDPINVKIAELREALESVTAEQKYLKARDARHRHTNESTRKRVVFYTVG 725
Query: 182 EYILLAVTSILQVVYIRRLFSKSFAYNRV 210
EY+LLA S LQV+YIRRLFSKS AYNRV
Sbjct: 726 EYLLLAAVSALQVIYIRRLFSKSVAYNRV 812
>TC216368 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane protein-like
(AT3g22845/MWI23_22), partial (42%)
Length = 570
Score = 150 bits (378), Expect = 5e-37
Identities = 74/89 (83%), Positives = 80/89 (89%)
Frame = +1
Query: 122 PSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVL 181
PSEHDLAKDEHLDPINVKIAELREALES+ EQKYLKARD RHR+TNEST+ RV+FYTV
Sbjct: 1 PSEHDLAKDEHLDPINVKIAELREALESVTAEQKYLKARDARHRHTNESTRKRVVFYTVG 180
Query: 182 EYILLAVTSILQVVYIRRLFSKSFAYNRV 210
EY+LLA S LQV+YIRRLFSKS AYNRV
Sbjct: 181 EYLLLAAVSALQVIYIRRLFSKSVAYNRV 267
>TC216367 similar to UP|Q9LIL4 (Q9LIL4) Coated vesicle membrane protein-like
(AT3g22845/MWI23_22), partial (36%)
Length = 447
Score = 124 bits (310), Expect = 4e-29
Identities = 58/97 (59%), Positives = 74/97 (75%), Gaps = 1/97 (1%)
Frame = +1
Query: 3 WFSKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHA-QSGDSVTGNFVVMDHDIFWSSD 61
W +C +V FF + SLS+++NDVECV E+ GD+V+GNFVV+DHDIFW+SD
Sbjct: 172 WTVLMCLMVSFFA-----RIESLSVTVNDVECVYEYVLYEGDTVSGNFVVVDHDIFWNSD 336
Query: 62 HPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIY 98
HPGIDFTVT+ GN VH++KGTSGDKF FKAP +G+Y
Sbjct: 337 HPGIDFTVTSAAGNTVHNIKGTSGDKFSFKAPTHGMY 447
>TC209113 similar to UP|Q6YZD2 (Q6YZD2) Coated vesicle membrane protein-like,
partial (88%)
Length = 857
Score = 119 bits (297), Expect = 1e-27
Identities = 64/179 (35%), Positives = 95/179 (52%), Gaps = 1/179 (0%)
Frame = +3
Query: 33 ECVSEHAQ-SGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFK 91
EC S + GD+V +FVV+ D W G+D V P G + + + +KF+F
Sbjct: 147 ECFSHDVKYEGDTVHVSFVVIKADSPWHYGDEGVDLVVKGPSGEQIQDFRDKTSEKFDFV 326
Query: 92 APQNGIYKFCFHNPMSTPETVSFYIHVGHIPSEHDLAKDEHLDPINVKIAELREALESII 151
A ++G++KFCF N ETV F +HVGH AKDEH P+ +I +L EAL +I
Sbjct: 327 AHKSGVHKFCFTNKSPYHETVDFDVHVGHFSYFEQHAKDEHFTPLLEQIGKLEEALYNIQ 506
Query: 152 TEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRLFSKSFAYNRV 210
EQ +L+A+ +R N++ R + + E L S LQV ++RLF + +RV
Sbjct: 507 FEQHWLEAQTDRQAIVNDAMSRRAVHKAIFESAALIGASALQVYLLQRLFERKLGTSRV 683
>TC230759 similar to UP|Q6ZGK3 (Q6ZGK3) Emp24/gp25L/p24-like, partial (75%)
Length = 1062
Score = 72.8 bits (177), Expect = 1e-13
Identities = 53/200 (26%), Positives = 100/200 (49%), Gaps = 3/200 (1%)
Frame = +1
Query: 7 ICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGID 66
+C + F++ L V+ L++ + +C+SE QS V ++ V+ + +
Sbjct: 169 LCFVAHFYVLPLAEAVW-LTLPTSGTKCLSEEIQSNVVVLADYYVVTQE----GGLQTLS 333
Query: 67 FTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE--TVSFYIHVGHIPSE 124
VT+P GN +H + + +F F ++G Y CF E TVS G +
Sbjct: 334 VKVTSPYGNNLHHNENATQGQFAFTTAESGNYVACFWMDGKHQEEATVSLDWKTGIYAKD 513
Query: 125 HD-LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEY 183
+ +AK E ++ + +++ +L A+E+I YLK ++ R R +E T RV +++++
Sbjct: 514 WESVAKKEKIEGVELELRKLEGAVEAIHGYLVYLKDKEARMREVSERTNGRVAWFSIMSL 693
Query: 184 ILLAVTSILQVVYIRRLFSK 203
+ + S+LQV Y++R F K
Sbjct: 694 SVCILVSVLQVWYLKRFFLK 753
>TC207713 similar to UP|Q6IDL4 (Q6IDL4) At1g09580, partial (75%)
Length = 859
Score = 70.5 bits (171), Expect = 5e-13
Identities = 50/200 (25%), Positives = 93/200 (46%), Gaps = 6/200 (3%)
Frame = +2
Query: 10 LVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTV 69
L++ T + L+I +C+SE Q+ V ++ V+ D+ I V
Sbjct: 44 LLLLLCFTTLTEAVWLTIPSKGTKCMSEEIQTHVVVLADYYVVADDV-QGHQLQTISAKV 220
Query: 70 TNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCF-----HNPMSTPETVSFYIHVG-HIPS 123
T+P GN +H + + +F F ++G Y CF H T+S G H
Sbjct: 221 TSPYGNNLHQNENVTHGQFAFTTTESGNYVACFWVNNKHQEGGGETTISLEWKTGIHAKD 400
Query: 124 EHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEY 183
+A+ E ++ + +++ +L A+E+I YLK R+ R +E+T RV ++++
Sbjct: 401 WDSVARKEKIEGVELELRKLEGAVEAIRYNLIYLKNREAEMREVSETTNARVAWFSIFSL 580
Query: 184 ILLAVTSILQVVYIRRLFSK 203
+ + S LQ+ +++R F K
Sbjct: 581 GICILVSGLQLWFLKRFFRK 640
>TC234728 similar to UP|Q6ZGK3 (Q6ZGK3) Emp24/gp25L/p24-like, partial (61%)
Length = 781
Score = 60.1 bits (144), Expect = 7e-10
Identities = 40/138 (28%), Positives = 71/138 (50%), Gaps = 3/138 (2%)
Frame = +2
Query: 69 VTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE--TVSFYIHVGHIPSEHD 126
VT+P GN + + +F F ++G Y CF E TVS G + +
Sbjct: 53 VTSPYGNNLXXNXXATQGQFAFTTAESGNYVACFWMDGKHQEEATVSLDWKTGIYAKDWE 232
Query: 127 -LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYIL 185
+AK E ++ + +++ +L A+E+I YLK ++ R R +E T RV +++++ +
Sbjct: 233 SVAKKEKIEGVELELRKLEGAVEAIHGYLVYLKDKEARMREVSERTNGRVAWFSIMSLSV 412
Query: 186 LAVTSILQVVYIRRLFSK 203
+ S+LQV Y++R F K
Sbjct: 413 CILVSVLQVWYLKRFFLK 466
>TC228200
Length = 1002
Score = 53.1 bits (126), Expect = 9e-08
Identities = 41/191 (21%), Positives = 83/191 (42%), Gaps = 6/191 (3%)
Frame = +2
Query: 19 IYNVFSLSISLNDVECVSEHAQSGDSVTGNF-VVMDHDIFWSSDHPGIDFTVTNPGGNAV 77
+ N + + +C+SE ++ G + VV + + D I VT+P N
Sbjct: 203 VANSMRFELQSGNTKCISEDIKTNAMSVGKYSVVNPREGYPLPDSHRIVVKVTSPHANMY 382
Query: 78 HSLKGTSGDKFEFKAPQNGIYKFCF--HNPMSTPE--TVSFYIHVGHIPSE-HDLAKDEH 132
H F F A + G Y CF + TP T+ F G + +AK
Sbjct: 383 HFGDHVDSGNFAFTASEAGDYSACFWVQDTRDTPSVVTIEFEWRTGVAAKDWSKVAKKGQ 562
Query: 133 LDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSIL 192
++ + ++ +L + + SI E YL+ R+ + N++T +++ ++ L ++ + L
Sbjct: 563 IEVMEFELKKLYDTVLSIHDEMFYLREREEEMQDLNKATNSKMFTFSFLSIVVCLSVAGL 742
Query: 193 QVVYIRRLFSK 203
Q+ +++ F +
Sbjct: 743 QLWHLKTFFER 775
>AW433005 similar to GP|21387055|gb| transmembrane protein-like protein
{Arabidopsis thaliana}, partial (27%)
Length = 508
Score = 48.9 bits (115), Expect = 2e-06
Identities = 38/173 (21%), Positives = 76/173 (42%), Gaps = 6/173 (3%)
Frame = +2
Query: 9 TLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFT 68
+L++ T + ++I +C+SE Q+ + ++ V D+ S I
Sbjct: 2 SLLLLLCFTTLTEALWVTIPSKGTKCMSEEIQTHVATLADYYVRADDV-QSHQLQTISAK 178
Query: 69 VTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCF-----HNPMSTPETVSFYIHVG-HIP 122
VT+P GN +H + + +F F + G Y CF H T+S + G H
Sbjct: 179 VTSPYGNNLHQNENVTHGQFAFTTTETGNYVACFRVDHKHQEGGGETTISLEWNTGIHAK 358
Query: 123 SEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRV 175
+A+++ +D + +++ + A+E++ Y+K N+ + TK R+
Sbjct: 359 DWDSVAREDEIDRVELELKKRDGAVEALRDNLMYMK---NKVAEMRDGTKPRM 508
>BQ079857 similar to GP|22296377|dbj P0519E12.19 {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 403
Score = 39.7 bits (91), Expect = 0.001
Identities = 26/89 (29%), Positives = 36/89 (40%), Gaps = 2/89 (2%)
Frame = +1
Query: 19 IYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVM--DHDIFWSSDHPGIDFTVTNPGGNA 76
I N + +C+SE +S GN+ V+ ++D F D I V +P GN
Sbjct: 10 IVNSMQFELKSGQSKCISEDLKSNVITVGNYYVLNPNNDGFPIPDSHKITVRVRSPYGND 189
Query: 77 VHSLKGTSGDKFEFKAPQNGIYKFCFHNP 105
H F F A + G Y CF P
Sbjct: 190 FHYGDSVHSGNFAFTAAEAGDYTACFSVP 276
>TC215627 similar to UP|Q9LNJ1 (Q9LNJ1) F6F3.12 protein, partial (4%)
Length = 697
Score = 31.2 bits (69), Expect = 0.36
Identities = 30/109 (27%), Positives = 41/109 (37%)
Frame = +1
Query: 26 SISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSG 85
S+ CV+E ++GD + + P D T NP ++ SL +
Sbjct: 199 SVPSTSKPCVNEKLENGDKLLSSL-------------PEPDPTDANPVKKSIVSLGKSPS 339
Query: 86 DKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHIPSEHDLAKDEHLD 134
K AP I KF +NP S V + SEHD K E D
Sbjct: 340 YKEVALAPPGTISKFQVYNPQS----------VISVSSEHDGGKHEEED 456
>TC207708 similar to UP|Q9LK35 (Q9LK35) Receptor-protein kinase-like protein,
partial (8%)
Length = 530
Score = 28.5 bits (62), Expect = 2.3
Identities = 8/29 (27%), Positives = 17/29 (58%)
Frame = +1
Query: 97 IYKFCFHNPMSTPETVSFYIHVGHIPSEH 125
++ +CF+NP P+++ F I++ H
Sbjct: 310 LFSYCFYNPFFPPQSIYFVIYLSDSDGHH 396
>TC218816 similar to UP|Q8GVB6 (Q8GVB6) ACC oxidase, partial (86%)
Length = 939
Score = 28.1 bits (61), Expect = 3.0
Identities = 12/36 (33%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = -2
Query: 98 YKFCFHNPMSTPETVSFYI-HVGHIPSEHDLAKDEH 132
++FCFH + P FYI H +P +H ++ H
Sbjct: 764 WRFCFHYCQA*PYAQHFYISHCSTLPLDHPCSQKRH 657
>TC203595 similar to UP|Q8L7T3 (Q8L7T3) Ubiquitin-conjugating enzyme
(At1g50490), partial (86%)
Length = 1115
Score = 27.3 bits (59), Expect = 5.2
Identities = 19/63 (30%), Positives = 23/63 (36%)
Frame = +2
Query: 41 SGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKF 100
SGDS F DH FW G TV + L + + FK P+
Sbjct: 542 SGDSGISAFPEEDHIFFWKGTITGSQDTVFE---GTEYKLSLSFPHDYPFKPPKVKFETT 712
Query: 101 CFH 103
CFH
Sbjct: 713 CFH 721
>TC225211 homologue to UP|PRP2_SOYBN (P13993) Repetitive proline-rich cell
wall protein 2 precursor, partial (62%)
Length = 718
Score = 26.9 bits (58), Expect = 6.7
Identities = 14/35 (40%), Positives = 17/35 (48%)
Frame = -3
Query: 8 CTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSG 42
CTLV+F LG VF L + C E +Q G
Sbjct: 503 CTLVVFLLGACTLGVFQLGV------CKQEASQLG 417
>TC218815 similar to UP|Q9ZUN4 (Q9ZUN4) 1-aminocyclopropane-1-carboxylate
oxidase, partial (38%)
Length = 666
Score = 26.9 bits (58), Expect = 6.7
Identities = 12/36 (33%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = -1
Query: 98 YKFCFHNPMSTPETVSFYI-HVGHIPSEHDLAKDEH 132
++FCFH + P FYI H +P +H ++ H
Sbjct: 198 WRFCFHYCQA*PYAQHFYISHCSTLPLDHPCSQKWH 91
>TC227236 weakly similar to UP|Q7PMV5 (Q7PMV5) ENSANGP00000017545 (Fragment),
partial (13%)
Length = 921
Score = 26.6 bits (57), Expect = 8.8
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +1
Query: 88 FEFKAPQNGIYKFCFHNPMSTPETVSFYIHV 118
F F++ + I K+ H P S T+S+ IHV
Sbjct: 589 FPFESTNSTICKYQLHPPSSMCVTISYSIHV 681
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,479,972
Number of Sequences: 63676
Number of extensions: 127822
Number of successful extensions: 698
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 690
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 696
length of query: 210
length of database: 12,639,632
effective HSP length: 93
effective length of query: 117
effective length of database: 6,717,764
effective search space: 785978388
effective search space used: 785978388
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0322.5


