

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
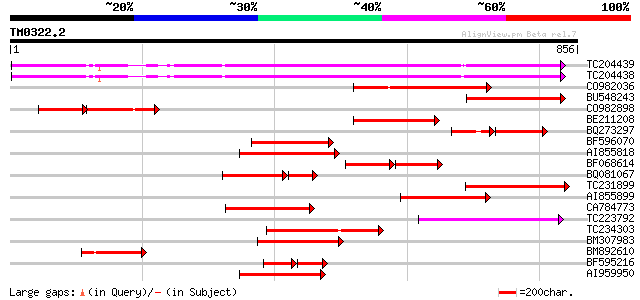
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 487 e-137
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 486 e-137
CO982036 219 5e-57
BU548243 197 2e-50
CO982898 114 2e-47
BE211208 167 2e-41
BQ273297 106 9e-39
BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vi... 157 1e-38
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 157 2e-38
BF068614 similar to GP|27817858|db OJ1081_B12.7 {Oryza sativa (j... 96 5e-38
BQ081067 weakly similar to GP|23495377|dbj orf490 {Oryza sativa ... 115 1e-37
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 153 4e-37
AI855899 similar to GP|2244960|emb| retrotransposon like protein... 150 2e-36
CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {V... 147 3e-35
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 140 2e-33
TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, parti... 140 3e-33
BM307983 135 6e-32
BM892610 133 4e-31
BF595216 73 4e-28
AI959950 121 1e-27
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 487 bits (1253), Expect = e-137
Identities = 288/842 (34%), Positives = 454/842 (53%), Gaps = 7/842 (0%)
Frame = +1
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKP--LTPYFTNLGI 61
++R+TWV +++KS+T+ F ++ + + +K +++D G EF+ T + T+ GI
Sbjct: 2341 FSRFTWVNFIREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGI 2520
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
H F+ T QNGIVERK+R + E +L + W A T Y+ NR V L
Sbjct: 2521 THEFSAAITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNR---VTL 2691
Query: 122 DNESPFTKLFQV----SFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKS 177
+P T L+++ VK IFG CY K+ S +FLGYST ++
Sbjct: 2692 RRGTP-TTLYEIWKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRA 2868
Query: 178 YKCLDPTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSS 237
Y+ +F + + S+ V + S
Sbjct: 2869 YR-------------------------VFNSRTRTVMESINVVVDDL------------S 2937
Query: 238 PAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSV 297
PA E S + + S A ++ P +SS+
Sbjct: 2938 PARKKD----VEEDVRTSGDNVADAAKSGENAENSDSATDESNINQPDKRSSTRIQKMHP 3105
Query: 298 SPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALH 357
++ + TR+++ + + +S+ EP + K AL D W +AMQ+E
Sbjct: 3106 KELIIGDPNRGVTTRSREVEIVSN---SCFVSKIEPKNVKEALTDEFWINAMQEELEQFK 3276
Query: 358 SNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPV 417
N+ W LVP P IG KW+F+ K N +G + R KARLVA+GY+Q+EG D+ ETF+PV
Sbjct: 3277 RNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPV 3456
Query: 418 VKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDRSLVCRLNK 476
+ ++R++L +A + ++Q+DV +AFLNG L EEVY+ QP GF + V RL K
Sbjct: 3457 ARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADPTHPDHVYRLKK 3636
Query: 477 ALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSL 536
ALYGLKQAPRAW+ERL L + G+ + +LF + + +YVDDI+ G S
Sbjct: 3637 ALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSN 3816
Query: 537 PVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKG 596
+L+H + ++ +EF + +G L YFLG+QV+ + D S+ L+QS+Y K+++++ M A
Sbjct: 3817 EMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMED-SIFLSQSRYAKNIVKKFGMENASH 3993
Query: 597 ISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEH 656
TP ++ KL+K A D SLYRS++G+L Y+T +RP+I+Y+V ++ + P H
Sbjct: 3994 KRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISH 4173
Query: 657 WAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPN 716
VKRIL+Y+ GT D+G+ CS P+ L+ +CDADWA DD +STSG +LG N
Sbjct: 4174 LTQVKRILKYVNGTSDYGIMYCHCS--NPM-LVGYCDADWAGSADDRKSTSGGCFYLGNN 4344
Query: 717 LVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVAL 776
L+SW SKKQ+ V+ S+ EAEY ++ ++L+W++ +L+E ++ + CDNMS + +
Sbjct: 4345 LISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINI 4524
Query: 777 THNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKL 836
+ NPV H+RTKH+++ ++R+ V +K + ++HV + +Q DI TKAL +FE LR KL
Sbjct: 4525 SKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKALDANQFEKLRGKL 4704
Query: 837 NV 838
+
Sbjct: 4705 GI 4710
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 486 bits (1252), Expect = e-137
Identities = 287/842 (34%), Positives = 450/842 (53%), Gaps = 7/842 (0%)
Frame = +1
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFK--PLTPYFTNLGI 61
++R+TWV +++KSDT+ F ++ + + +K +++D G EF+ T + T+ GI
Sbjct: 2344 FSRFTWVNFIREKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGI 2523
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
H F+ T QNGIVERK+R + E +L + W A T Y+ NR V L
Sbjct: 2524 THEFSAAITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNR---VTL 2694
Query: 122 DNESPFTKLFQV----SFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKS 177
+P T L+++ VK IFG CY K+ S +FLGYST ++
Sbjct: 2695 RRGTP-TTLYEIWKGRKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRA 2871
Query: 178 YKCLDPTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSS 237
Y+ +F + + S+ V + +
Sbjct: 2872 YR-------------------------VFNSRTRTVMESINVVVDDL------------T 2940
Query: 238 PAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSV 297
PA E S + + S A P ++ P + S
Sbjct: 2941 PARKKD----VEEDVRTSGDNVADTAKSAENAENSDSATDEPNINQPDKRPSIRIQKMHP 3108
Query: 298 SPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALH 357
++ + TR+++ + + +S+ EP + K AL D W +AMQ+E
Sbjct: 3109 KELIIGDPNRGVTTRSREIEIVSN---SCFVSKIEPKNVKEALTDEFWINAMQEELEQFK 3279
Query: 358 SNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPV 417
N+ W LVP P IG KW+F+ K N +G + R KARLVA+GY+Q+EG D+ ETF+PV
Sbjct: 3280 RNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIEGVDFDETFAPV 3459
Query: 418 VKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDRSLVCRLNK 476
+ ++R++L +A + ++Q+DV +AFLNG L EE Y+ QP GF + V RL K
Sbjct: 3460 ARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFVDPTHPDHVYRLKK 3639
Query: 477 ALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSL 536
ALYGLKQAPRAW+ERL L + G+ + +LF + + +YVDDI+ G S
Sbjct: 3640 ALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIYVDDIVFGGMSN 3819
Query: 537 PVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKG 596
+L+H + ++ +EF + +G L YFLG+QV+ + D S+ L+QSKY K+++++ M A
Sbjct: 3820 EMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMED-SIFLSQSKYAKNIVKKFGMENASH 3996
Query: 597 ISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEH 656
TP ++ KL+K A D SLYRS++G+L Y+T +RP+I+Y+V ++ + P H
Sbjct: 3997 KRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCARYQANPKISH 4176
Query: 657 WAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPN 716
VKRIL+Y+ GT D+G+ CS L+ +CDADWA DD +STSG +LG N
Sbjct: 4177 LNQVKRILKYVNGTSDYGIMYCHCS---DSMLVGYCDADWAGSADDRKSTSGGCFYLGTN 4347
Query: 717 LVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVAL 776
L+SW SKKQ+ V+ S+ EAEY ++ ++L+W++ +L+E ++ + CDNMS + +
Sbjct: 4348 LISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCDNMSAINI 4527
Query: 777 THNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKL 836
+ NPV H+RTKH+++ ++R+ V +K + ++HV + +Q DI TKAL +FE LR KL
Sbjct: 4528 SKNPVQHSRTKHIDIRHHYIRDLVDDKVITLEHVDTEEQIADIFTKALDANQFEKLRGKL 4707
Query: 837 NV 838
+
Sbjct: 4708 GI 4713
>CO982036
Length = 674
Score = 219 bits (557), Expect = 5e-57
Identities = 112/209 (53%), Positives = 150/209 (71%)
Frame = -2
Query: 519 SVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQ 578
+VY+LVYVD IIITG+S ++Q+L +KLN+ F LK LG LDYF+ ++V+ + D LL +
Sbjct: 655 TVYLLVYVD-IIITGSSCTLIQNLTSKLNSSFPLKLLGKLDYFVEIEVKSMPD--LLFSL 485
Query: 579 SKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEI 638
I ++ R A+ IS+PM + KL+K ++ + P+ YRS+VGALQY T+ RPEI
Sbjct: 484 RTSIFEIFCRKPR*QAQPISSPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTVIRPEI 305
Query: 639 SYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAA 698
S++VNKVCQF+S PL+ HW VKRILRYLKG++ +GL LKP QPL + FCDADWA+
Sbjct: 304 SFAVNKVCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAISSQPLPIRGFCDADWAS 125
Query: 699 DPDDHRSTSGSLVFLGPNLVSWCSKKQSL 727
DD RSTSG+ VFLGPNL+SW KQ +
Sbjct: 124 AVDDKRSTSGAAVFLGPNLISWWXXKQQV 38
>BU548243
Length = 599
Score = 197 bits (501), Expect = 2e-50
Identities = 95/149 (63%), Positives = 115/149 (76%)
Frame = -1
Query: 690 AFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLW 749
A CDA WA+D DDHRST GS +FLGPNL+SW S+KQ + A+SSTEAEYR++A T AEL W
Sbjct: 596 ALCDAGWASDVDDHRSTLGSAIFLGPNLISWWSRKQQVTAQSSTEAEYRSIAQTSAELTW 417
Query: 750 VQSLLQELHIPHKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQH 809
+Q+LL EL IP P+ILCDN S VA+ HN V H+RTKHME+D+FFV EKVL+K L I H
Sbjct: 416 IQALLMELQIPFTPPVILCDNKSAVAIAHNLVFHSRTKHMEIDVFFVHEKVLSKQLQIFH 237
Query: 810 VPSLDQRDDILTKALSPLRFEDLRTKLNV 838
+P+LDQ ILTK LS RF L++KL V
Sbjct: 236 IPALDQWAGILTKPLSSARFTFLKSKLTV 150
>CO982898
Length = 816
Score = 114 bits (285), Expect(2) = 2e-47
Identities = 54/109 (49%), Positives = 73/109 (66%)
Frame = -1
Query: 117 PTVVLDNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHK 176
PTV ++ P+T+LF D F R+FGCAC+P +RPYN K + S EC+FLG T+HK
Sbjct: 597 PTVSINFAVPYTQLFNQMLDYSFPRVFGCACFPLIRPYNIQKFNFKSHECIFLGNYTSHK 418
Query: 177 SYKCLDPTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIP 225
+YKCL P G+L +SKDV+FNE FPYKELF SS + + +++P
Sbjct: 417 NYKCLAPAGKL-VSKDVIFNEI*FPYKELFLPDSSTCIPVPQPIPATVP 274
Score = 94.4 bits (233), Expect(2) = 2e-47
Identities = 43/74 (58%), Positives = 57/74 (76%)
Frame = -3
Query: 44 DGGGEFKPLTPYFTNLGIVHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWD 103
D GGEF PLT + + GI+H+ PHT +QN +VERK RHIV+ GL+LL+HA + L++W
Sbjct: 814 DWGGEF*PLTEFLND*GIIHKLIFPHTQYQN-VVERKQRHIVKFGLSLLSHASLPLKFWG 638
Query: 104 HAFLTVVYLINRLP 117
HAF+T V+LINRLP
Sbjct: 637 HAFITPVFLINRLP 596
>BE211208
Length = 413
Score = 167 bits (423), Expect = 2e-41
Identities = 82/129 (63%), Positives = 105/129 (80%)
Frame = +2
Query: 520 VYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQS 579
VY+LVYVDDIIITG S ++Q L+ LN+ FSLKQLG LDYFLG++V H S+LLTQS
Sbjct: 26 VYLLVYVDDIIITGRSNYLIQSLVHHLNSNFSLKQLGQLDYFLGIEVHHTPTGSVLLTQS 205
Query: 580 KYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEIS 639
KYI DLL + MA AK IS+PM++N +L+K+G + ++DP++YRS+VGALQY TITRPEIS
Sbjct: 206 KYICDLLHKTDMAEAKPISSPMVTNLRLSKNGDDLLSDPTMYRSVVGALQYPTITRPEIS 385
Query: 640 YSVNKVCQF 648
++ NKVCQF
Sbjct: 386 FAANKVCQF 412
>BQ273297
Length = 420
Score = 106 bits (265), Expect(2) = 9e-39
Identities = 50/78 (64%), Positives = 62/78 (79%)
Frame = +2
Query: 734 EAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVALTHNPVLHTRTKHMELDI 793
E E+R++ VA++ +QSLL EL + H TPLILCDN STV+L HN +LH+RTKHMELD+
Sbjct: 185 ETEHRSMTLIVAKVT*IQSLLSELQVAHSTPLILCDNTSTVSLAHNLILHSRTKHMELDL 364
Query: 794 FFVREKVLNKSLIIQHVP 811
FFVREKVLNK L + HVP
Sbjct: 365 FFVREKVLNKKLDVVHVP 418
Score = 73.2 bits (178), Expect(2) = 9e-39
Identities = 37/66 (56%), Positives = 46/66 (69%)
Frame = +1
Query: 667 LKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQS 726
LKGT+ +GLH + SL P+ L A+C +DWA+DPD RS FLGPNL+ SKKQS
Sbjct: 1 LKGTISWGLHFQATSLQHPIPLHAYCGSDWASDPDYRRS------FLGPNLIC*WSKKQS 162
Query: 727 LVARSS 732
+VARSS
Sbjct: 163 VVARSS 180
>BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vinifera},
partial (34%)
Length = 407
Score = 157 bits (398), Expect = 1e-38
Identities = 73/126 (57%), Positives = 96/126 (75%), Gaps = 1/126 (0%)
Frame = -2
Query: 365 VPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVR 424
VPLP + +GC+WV+ +K G V+R KARLVAKGY+QV G DY +TFSPV K TVR
Sbjct: 406 VPLPPGKTPVGCRWVYTVKVGPTGEVDRLKARLVAKGYTQVYGIDYCDTFSPVAKLTTVR 227
Query: 425 IILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK-DRSLVCRLNKALYGLKQ 483
+ L++A W +HQLD+ NAFL+G LEE++YM QPPGF A+ + LVC+L+++LYGLKQ
Sbjct: 226 LFLAMAAICHWPLHQLDIKNAFLHGDLEEDIYMEQPPGFVAQGEYGLVCKLHRSLYGLKQ 47
Query: 484 APRAWF 489
+PRAWF
Sbjct: 46 SPRAWF 29
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 157 bits (397), Expect = 2e-38
Identities = 78/152 (51%), Positives = 105/152 (68%), Gaps = 1/152 (0%)
Frame = -3
Query: 348 AMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEG 407
AMQ+E N N W LV P + IG KWVFR K + G + R KARLVAKGY+Q EG
Sbjct: 458 AMQEELNQFERNNVWKLVEKPENYPVIGTKWVFRNKLDEHGIIIRNKARLVAKGYNQEEG 279
Query: 408 FDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD 467
DY+ET++PV + +R++L+ + ++Q+DV +AFLNG ++EEVY+ QPPGFE D
Sbjct: 278 IDYEETYAPVARLEVIRMLLAYVSIMNFKLYQMDVKSAFLNGLIQEEVYVEQPPGFEIPD 99
Query: 468 R-SLVCRLNKALYGLKQAPRAWFERLKAALLK 498
+ + V +L KALYGLKQAPRAW+ER+ LL+
Sbjct: 98 KPTHVYKLQKALYGLKQAPRAWYERISNFLLE 3
>BF068614 similar to GP|27817858|db OJ1081_B12.7 {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 413
Score = 96.3 bits (238), Expect(2) = 5e-38
Identities = 45/71 (63%), Positives = 58/71 (81%)
Frame = +1
Query: 583 KDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSV 642
+DLL + MA A+ IS+PM+S+ KL+K G++ DP+LY S+VGALQY T+TRPE SYSV
Sbjct: 199 RDLLPKTKMAEAQPISSPMVSSCKLSKSGSDLFQDPTLYGSVVGALQYATLTRPEFSYSV 378
Query: 643 NKVCQFLSQPL 653
NKVCQF+SQPL
Sbjct: 379 NKVCQFMSQPL 411
Score = 81.3 bits (199), Expect(2) = 5e-38
Identities = 38/75 (50%), Positives = 57/75 (75%)
Frame = +2
Query: 507 NPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQV 566
+PSLF +V++LVYVDDI ITGN + ++Q LI++LN++F+LKQLG LDYFLG++V
Sbjct: 2 DPSLFVYKQQQYTVFLLVYVDDITITGNCVLLIQQLISQLNSQFALKQLGLLDYFLGIEV 181
Query: 567 QHLHDKSLLLTQSKY 581
++L DK + + K+
Sbjct: 182 KYLPDKGISCPRLKW 226
>BQ081067 weakly similar to GP|23495377|dbj orf490 {Oryza sativa (japonica
cultivar-group)}, partial (18%)
Length = 430
Score = 115 bits (288), Expect(2) = 1e-37
Identities = 57/98 (58%), Positives = 71/98 (72%)
Frame = +1
Query: 322 LVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFR 381
L PTLL + AEP++ K AL P W AMQ +F+AL N+T TL LPS + AI CKWVFR
Sbjct: 1 LHPTLLFTTAEPSTVKQALISPPWRQAMQADFDALMENKTLTLTSLPSGKAAIDCKWVFR 180
Query: 382 IKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVK 419
IKEN G++NRY++RLVAKG+ G DY ETFSPV++
Sbjct: 181 IKENLYGTLNRYRSRLVAKGFHLKFGCDYSETFSPVIE 294
Score = 60.8 bits (146), Expect(2) = 1e-37
Identities = 26/44 (59%), Positives = 36/44 (81%)
Frame = +3
Query: 421 ITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFE 464
+T+R+IL +A++ W + Q+D+NNAFL+G L EEVYMVQ PGFE
Sbjct: 297 VTIRLILFIALTNHWPLQQVDINNAFLHGLLTEEVYMVQLPGFE 428
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment),
partial (30%)
Length = 687
Score = 153 bits (386), Expect = 4e-37
Identities = 76/159 (47%), Positives = 106/159 (65%), Gaps = 1/159 (0%)
Frame = +2
Query: 688 LIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAEL 747
L +CDADWA P D RSTSG VF+G NLVSW SKKQ++VARSS EAEYR++A EL
Sbjct: 17 LSGYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCEL 196
Query: 748 LWVQSLLQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLI 806
+W++ LQEL + + + CDN + + + NPV H RTKH+E+D F+REK+L+K ++
Sbjct: 197 MWIKQFLQELRFCEELQMKLYCDNQAALHIASNPVFHERTKHIEIDCHFIREKLLSKEIV 376
Query: 807 IQHVPSLDQRDDILTKALSPLRFEDLRTKLNVTDKFTLA 845
+ + S DQ DILTK+L + + + +KL D + A
Sbjct: 377 TEFIGSNDQPVDILTKSLRGPKIQIVCSKLGAYDLYAPA 493
>AI855899 similar to GP|2244960|emb| retrotransposon like protein
{Arabidopsis thaliana}, partial (18%)
Length = 418
Score = 150 bits (380), Expect = 2e-36
Identities = 74/136 (54%), Positives = 97/136 (70%)
Frame = +1
Query: 590 SMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFL 649
+M GISTPM+S+ KL+K G+ + + YR IVGALQYVT+TRP I+Y+VNKV +F+
Sbjct: 10 NMLDCNGISTPMVSSYKLSKFGSELLPNAHQYRDIVGALQYVTLTRPNIAYNVNKVSEFM 189
Query: 650 SQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGS 709
S PL+ + VKRILRYL GTV GL L+P + +SL A+ D DW +DP + RSTSGS
Sbjct: 190 SSPLQSY*LTVKRILRYLSGTVTQGLLLQPAHMDAKISLRAYNDLDWGSDPAEMRSTSGS 369
Query: 710 LVFLGPNLVSWCSKKQ 725
+F G NL++W SKKQ
Sbjct: 370 CIFSGSNLIAWSSKKQ 417
>CA784773 weakly similar to GP|27901698|gb gag-pol polyprotein {Vitis
vinifera}, partial (34%)
Length = 409
Score = 147 bits (370), Expect = 3e-35
Identities = 69/134 (51%), Positives = 92/134 (68%)
Frame = +3
Query: 327 LLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENA 386
L S P++ + AL P W AM DE AL +N TW LVPLP + +GC+WV+ +K
Sbjct: 3 LSSLTVPSTIREALDHPGWRQAMVDEMQALENNGTWELVPLPPGKTTVGCRWVYTVKVGP 182
Query: 387 DGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAF 446
+G V+R KARLVAKGY+QV G +Y +TFSPV TVR+ L++A + W +HQLD+ NAF
Sbjct: 183 NGKVDRLKARLVAKGYTQVYGIEYCDTFSPVFFLTTVRLFLAMAAIRHWPLHQLDIKNAF 362
Query: 447 LNGKLEEEVYMVQP 460
L+G LEE++YM QP
Sbjct: 363 LHGDLEEDIYMEQP 404
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 140 bits (353), Expect = 2e-33
Identities = 70/221 (31%), Positives = 131/221 (58%), Gaps = 1/221 (0%)
Frame = +1
Query: 617 DPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLH 676
D + +R ++G+L+Y+ +RP I ++V+ + +F+ +P H A KR+LR +KGT+ G+
Sbjct: 10 DVTEFRRLIGSLRYLCNSRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVL 189
Query: 677 LKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAE 736
+ L+ + D+DW DP+ +ST G L V+ SKKQ ++A S+ EAE
Sbjct: 190 FPFKAKSGKPDLLGYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAE 369
Query: 737 YRNLANTVAELLWVQSLLQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFF 795
Y + + +W+ +LL+EL + + P+ +L DN S + L +P LH R+KH+EL +
Sbjct: 370 YVAASLGACQAVWMMNLLEELKLRERKPVNLLIDNKSAINLAKHPTLHGRSKHIELRFHY 549
Query: 796 VREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKL 836
+R++V ++ +++ + +Q D++TK + RF+ + ++L
Sbjct: 550 IRDQVSKGNVTVEYCKAEEQLADLMTKPIQVSRFKQICSEL 672
>TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, partial (10%)
Length = 558
Score = 140 bits (352), Expect = 3e-33
Identities = 84/180 (46%), Positives = 114/180 (62%), Gaps = 3/180 (1%)
Frame = +1
Query: 388 GSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFL 447
G+++++KARLVAK Y+QV G DY TFSPV K V ++ S+AV W + LD NAFL
Sbjct: 28 GTIDQFKARLVAKSYTQVYGQDYTGTFSPVAKMAYVHLLWSMAVVCHWPLF*LDAKNAFL 207
Query: 448 NGKLEEEVYMVQPPGFEAKDRS--LVCRLNKALYGLKQAPRAW-FERLKAALLKFGFTAS 504
+G LEEEVYM QP GF A+ S +VC+L ++ YGLKQ+PRAW F AA+ + +
Sbjct: 208 HGYLEEEVYMEQPLGFVAQGESSNMVCQLCRSFYGLKQSPRAWPFLYCGAAI---WYDSH 378
Query: 505 CCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGV 564
+ S+F +S +Y++VYVDDI ITG+ + L L +F K LG L YFLG+
Sbjct: 379 EADHSVFYCHSPQGCIYLIVYVDDIGITGSDQHGIT*LK*XLCCQFQTKDLGKLRYFLGI 558
>BM307983
Length = 406
Score = 135 bits (341), Expect = 6e-32
Identities = 70/133 (52%), Positives = 90/133 (67%), Gaps = 2/133 (1%)
Frame = +2
Query: 374 IGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSK 433
+GC+W++ +K AD +++RYKARLVAKGY Q G DY+ETF+ K I ++
Sbjct: 2 VGCRWIYTVKY*ADDTLDRYKARLVAKGYIQTYGIDYEETFAQWQK*IQSGSSSP*QQAQ 181
Query: 434 -GWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD-RSLVCRLNKALYGLKQAPRAWFER 491
GW +HQ DV NAFL+G LEEEVYM PPG+ A + + VCRL KALYGLKQ+PRAWF R
Sbjct: 182 FGWEMHQFDVKNAFLHGSLEEEVYMEIPPGYGASNGGNKVCRLKKALYGLKQSPRAWFGR 361
Query: 492 LKAALLKFGFTAS 504
A+L G+ S
Sbjct: 362 FTQAMLSLGYKQS 400
>BM892610
Length = 421
Score = 133 bits (334), Expect = 4e-31
Identities = 60/98 (61%), Positives = 73/98 (74%)
Frame = -3
Query: 109 VVYLINRLPTVVLDNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVF 168
+VYLINRLPT L + P T +F + FLR FGC+CYP+LRPYN HKL S EC+F
Sbjct: 419 IVYLINRLPTTSLKFQVPCT-VFNKLPNYNFLRNFGCSCYPFLRPYNKHKLDFRSHECLF 243
Query: 169 LGYSTTHKSYKCLDPTGRLYISKDVLFNEFRFPYKELF 206
LGYST HK YKCL P G+LY+SKDV+FNE ++PY +LF
Sbjct: 242 LGYSTPHKGYKCLSPNGKLYVSKDVIFNELKYPYSDLF 129
>BF595216
Length = 421
Score = 72.8 bits (177), Expect(2) = 4e-28
Identities = 32/45 (71%), Positives = 39/45 (86%)
Frame = +3
Query: 435 WHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVCRLNKALY 479
W I QLDVNNAFLNG L+EEVYM QPPGF++ +SLVC+L+KA+Y
Sbjct: 285 WPIQQLDVNNAFLNGILQEEVYMQQPPGFDSTTKSLVCKLHKAIY 419
Score = 71.2 bits (173), Expect(2) = 4e-28
Identities = 33/50 (66%), Positives = 44/50 (88%)
Frame = +1
Query: 383 KENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVS 432
KEN+DG+VN+YKA+LVAKG+ Q G DY ETFS V+KPIT+R++L+LAV+
Sbjct: 124 KENSDGTVNKYKAKLVAKGFHQQYGTDYIETFSLVIKPITMRLLLTLAVT 273
Score = 29.3 bits (64), Expect = 7.7
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +2
Query: 3 AYTRYTWVFPLKKKSDTYT 21
A+TRYTW+F L+ KS+ T
Sbjct: 77 AFTRYTWIFLLRHKSEKKT 133
>AI959950
Length = 466
Score = 121 bits (303), Expect = 1e-27
Identities = 63/130 (48%), Positives = 84/130 (64%), Gaps = 1/130 (0%)
Frame = -1
Query: 348 AMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEG 407
AMQ+E + N LV LP +K +G KW+F K + DG V RYKARLVAKGYSQ EG
Sbjct: 391 AMQEELDQFQKNNV*KLVKLPKRKKVVGVKWIFCNKLDEDGKVVRYKARLVAKGYSQQEG 212
Query: 408 FDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD 467
DY +TF+ V + + I+LS A ++Q+DV +AFLNG +++EVY+ QPPGFE +
Sbjct: 211 IDYPKTFALVARLEVICILLSFATYSNMKLYQMDVKSAFLNGLIQKEVYVEQPPGFENET 32
Query: 468 -RSLVCRLNK 476
V +LNK
Sbjct: 31 LHQHVFKLNK 2
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,466,304
Number of Sequences: 63676
Number of extensions: 752175
Number of successful extensions: 14411
Number of sequences better than 10.0: 1428
Number of HSP's better than 10.0 without gapping: 8314
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11398
length of query: 856
length of database: 12,639,632
effective HSP length: 105
effective length of query: 751
effective length of database: 5,953,652
effective search space: 4471192652
effective search space used: 4471192652
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0322.2


