

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
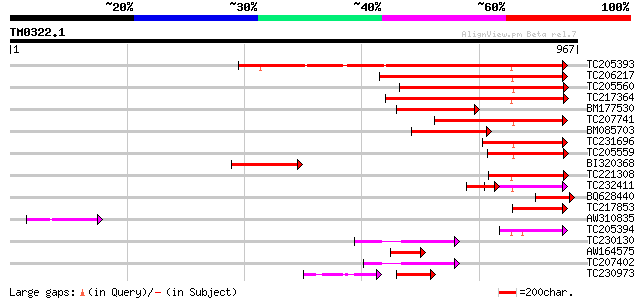
Query= TM0322.1
(967 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC205393 UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase, partial... 501 e-142
TC206217 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-tra... 419 e-117
TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transpor... 407 e-113
TC217364 plasma membrane Ca2+-ATPase 354 1e-97
BM177530 homologue to GP|16508164|g type IIB calcium ATPase {Med... 270 3e-72
TC207741 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, ... 257 2e-68
BM085703 homologue to SP|Q9LIK7|ACA Potential calcium-transporti... 245 8e-65
TC231696 weakly similar to UP|ACAA_ARATH (Q9SZR1) Potential calc... 176 4e-44
TC205559 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transpor... 171 1e-42
BI320368 similar to SP|Q9LU41|ACA Potential calcium-transporting... 127 3e-29
TC221308 similar to UP|Q9FVE7 (Q9FVE7) Plasma membrane Ca2+-ATPa... 112 6e-25
TC232411 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, ... 110 4e-24
BQ628440 similar to GP|16508164|gb type IIB calcium ATPase {Medi... 108 1e-23
TC217853 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-AT... 103 5e-22
AW310835 similar to GP|16508162|g type IIB calcium ATPase {Medic... 88 2e-17
TC205394 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-AT... 88 2e-17
TC230130 similar to UP|AHM7_ARATH (Q9SH30) Potential copper-tran... 85 1e-16
AW164575 similar to GP|8843813|dbj Ca2+-transporting ATPase-like... 79 1e-14
TC207402 similar to UP|Q941L1 (Q941L1) Copper-transporting P-typ... 78 2e-14
TC230973 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPa... 55 4e-14
>TC205393 UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase, partial (55%)
Length = 2009
Score = 501 bits (1290), Expect = e-142
Identities = 281/572 (49%), Positives = 377/572 (65%), Gaps = 12/572 (2%)
Frame = +2
Query: 391 KTGTLTLNQMRVTK--FWLGLENVVENFSNAMAPTVLE----LFHQGVGLNTTGSVYKPS 444
KTGTLT N M V K F + + V N ++++ + E L + + NT G V +
Sbjct: 2 KTGTLTTNHMTVVKTCFCMNSKEVSNNNASSLCSELPEPAVKLLLESIFNNTGGEVVV-N 178
Query: 445 AESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNN 504
+ EI G+PTE A+L + +S LG D KQ K++ VE FNS KK+ V V
Sbjct: 179 QNGKREILGTPTEAAILEFGLS-LGGDFQGEKQACKLVKVEPFNSTKKKMSVVVELPGGG 355
Query: 505 TVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEER-SKIEKIIQGMAASSLRCIAFAYMEI 563
+ H KGA+E++LA C ++SNG LDEE S ++ I A+ +LR + AY+E+
Sbjct: 356 -LRAHCKGASEIILAACDKVLNSNGEVVPLDEESTSHLKATINQFASEALRTLCLAYVEL 532
Query: 564 SEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIF 623
G P + G T +G++G+KDP RP VK++V + AG+ ++M+TGDNI
Sbjct: 533 ENGFS------PEDPIPVSGYTCIGVIGIKDPVRPGVKESVAM*RSAGITVRMVTGDNIN 694
Query: 624 TAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCL 683
TAKAIA ECGIL + G+ +EG EFR +++E +E + KI+VMARSSP+DK +V+ L
Sbjct: 695 TAKAIARECGILTDD---GIAIEGPEFREKSQKELLELIPKIQVMARSSPLDKHTLVKHL 865
Query: 684 KKK-GHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRW 742
+ G VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +W
Sbjct: 866 RTTFGEVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKW 1045
Query: 743 GRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALAT 802
GR VY NIQKF+QFQLTVNV AL++NF +A +G PLT VQLLWVN+IMDTLGALALAT
Sbjct: 1046GRSVYINIQKFVQFQLTVNVVALIVNFTSACLTGTAPLTAVQLLWVNMIMDTLGALALAT 1225
Query: 803 ERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEV 858
E P +LM++ P+GR I+ +MWRN+L Q+LYQ V+ Q GKSIF + S V
Sbjct: 1226EPPNDDLMKRSPVGRKGNFISNVMWRNILGQSLYQFMVIWFLQSRGKSIFLLEGPNSDLV 1405
Query: 859 KNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFA 918
NTLIFN+FV CQVFNE NSR MEK+NVF+GIL N++F+G++ T+ Q+++VE L FA
Sbjct: 1406LNTLIFNSFVFCQVFNEINSREMEKINVFKGILDNYVFVGVISATVFFQIIIVEYLGTFA 1585
Query: 919 DTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
+T L QW C+ + + PIA K PV
Sbjct: 1586NTTPLTLSQWFFCLLVGFMGMPIAARLKKIPV 1681
>TC206217 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-transporting
ATPase 9, plasma membrane-type (Ca(2+)-ATPase isoform
9) , partial (32%)
Length = 1383
Score = 419 bits (1078), Expect = e-117
Identities = 217/329 (65%), Positives = 255/329 (76%), Gaps = 8/329 (2%)
Frame = +1
Query: 631 ECGIL-DLNDAGGV-VVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGH 688
ECGIL + DA ++EG +FR +E+ER + KI VM RSSP DKLL+VQ L+K G
Sbjct: 4 ECGILASIEDAVEPNIIEGKKFRELSEKEREDIAKKITVMGRSSPNDKLLLVQALRKGGE 183
Query: 689 VVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYN 748
VVAVTGDGTNDAPAL EADIGLSMGIQGTEVAKESSDI+ILDDNF SV V+RWGR VY
Sbjct: 184 VVAVTGDGTNDAPALHEADIGLSMGIQGTEVAKESSDIIILDDNFASVVKVVRWGRSVYA 363
Query: 749 NIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKE 808
NIQKFIQFQLTVNVAALVIN +AA++SGDVPL VQLLWVNLIMDTLGALALATE PT
Sbjct: 364 NIQKFIQFQLTVNVAALVINVVAAITSGDVPLNAVQLLWVNLIMDTLGALALATEPPTDR 543
Query: 809 LMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSK------EVKNTL 862
LM + P+GR E LIT IMWRNL+ QA+YQIAVLLV F G+SI +VKNTL
Sbjct: 544 LMHRSPVGRRESLITNIMWRNLIVQAVYQIAVLLVLNFCGESILPKQNTRADAFQVKNTL 723
Query: 863 IFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTER 922
IFN FVLCQ+FNEFN+R +++NVF G+ KN LF+GIVG+T +LQ++++E L KF T R
Sbjct: 724 IFNAFVLCQIFNEFNARKPDEMNVFRGVTKNKLFVGIVGVTFILQIIIIEFLGKFTSTVR 903
Query: 923 LNWEQWGICIGIAAVSWPIAWLTKLTPVP 951
L+W+ W +GI VSWP+A + K PVP
Sbjct: 904 LDWKLWLASLGIGFVSWPLAIVGKFIPVP 990
>TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(27%)
Length = 1356
Score = 407 bits (1046), Expect = e-113
Identities = 202/294 (68%), Positives = 240/294 (80%), Gaps = 7/294 (2%)
Frame = +1
Query: 666 RVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSD 725
+VM RSSP DKLL+VQ L++KGHVVAVTGDGTNDAPAL EADIGL+MGIQGTEVAKESSD
Sbjct: 7 QVMGRSSPNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEADIGLAMGIQGTEVAKESSD 186
Query: 726 IVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQL 785
I+ILDDNF SV V+RWGR VY NIQKFIQFQLTVNVAALVIN +AAVSSGDVPL VQL
Sbjct: 187 IIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTVNVAALVINVVAAVSSGDVPLNAVQL 366
Query: 786 LWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQ 845
LWVNLIMDTLGALALATE PT LM + P+GR EPLIT IMWRNLL QA+YQ++VLLV
Sbjct: 367 LWVNLIMDTLGALALATEPPTDHLMDRTPVGRREPLITNIMWRNLLIQAMYQVSVLLVLN 546
Query: 846 FYGKSIFNVSKE-------VKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLG 898
F G SI +S + VKNTLIFN FVLCQ+FNEFN+R ++ N+F+G+ +N+LF+G
Sbjct: 547 FRGISILGLSHDRKDHAIKVKNTLIFNAFVLCQIFNEFNARKPDEFNIFKGVTRNYLFMG 726
Query: 899 IVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPS 952
I+G+T+VLQ++++ L KF T RLNW+QW I + I + WP+A + KL PVP+
Sbjct: 727 IIGLTVVLQIVIILFLGKFTTTVRLNWKQWLISVVIGLIGWPLAVIGKLIPVPT 888
>TC217364 plasma membrane Ca2+-ATPase
Length = 1375
Score = 354 bits (908), Expect = 1e-97
Identities = 179/316 (56%), Positives = 236/316 (74%), Gaps = 5/316 (1%)
Frame = +3
Query: 642 GVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTNDA 700
G+ +EG EFR +EEE ++ + KI+VMARSSPMDK +V+ L+ VV+VTGDGTNDA
Sbjct: 9 GIAIEGPEFREKSEEELLDIIPKIQVMARSSPMDKHTLVKHLRTTFQEVVSVTGDGTNDA 188
Query: 701 PALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTV 760
PAL EADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +WGR VY NIQKF+QFQLTV
Sbjct: 189 PALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTV 368
Query: 761 NVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEP 820
NV AL++NF +A +G+ PLT VQLLWVN+IMDTLGALALATE P ELM++ P+GR
Sbjct: 369 NVVALIVNFSSACLTGNAPLTAVQLLWVNMIMDTLGALALATEPPNDELMKRPPVGRKGN 548
Query: 821 LITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEF 876
I+ +MWRN+L Q++YQ V+ Q GK F++ S + NTLIFN+FV CQVFNE
Sbjct: 549 FISNVMWRNILGQSIYQFVVIWFLQTRGKVTFHLDGPDSDLILNTLIFNSFVFCQVFNEI 728
Query: 877 NSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAA 936
+SR ME++NVFEGILKN++F+ ++ T+V Q+++VE L FA+T L+ +QW +
Sbjct: 729 SSRDMERINVFEGILKNYVFVAVLTSTVVFQIIIVEFLGTFANTSPLSLKQWFGSVLFGV 908
Query: 937 VSWPIAWLTKLTPVPS 952
+ PIA K+ PV S
Sbjct: 909 LGMPIAAALKMIPVGS 956
>BM177530 homologue to GP|16508164|g type IIB calcium ATPase {Medicago
truncatula}, partial (13%)
Length = 422
Score = 270 bits (689), Expect = 3e-72
Identities = 137/140 (97%), Positives = 140/140 (99%)
Frame = +2
Query: 661 KVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVA 720
KV+KIRVMARSSP+DKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVA
Sbjct: 2 KVEKIRVMARSSPLDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVA 181
Query: 721 KESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPL 780
KESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINF+AAVSSGDVPL
Sbjct: 182 KESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFVAAVSSGDVPL 361
Query: 781 TTVQLLWVNLIMDTLGALAL 800
TTVQLLWVNLIMDTLGALAL
Sbjct: 362 TTVQLLWVNLIMDTLGALAL 421
>TC207741 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, partial
(24%)
Length = 947
Score = 257 bits (656), Expect = 2e-68
Identities = 125/230 (54%), Positives = 164/230 (70%), Gaps = 4/230 (1%)
Frame = +3
Query: 725 DIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQ 784
D++I+DDNF ++ V RWGR +Y NIQKF+QFQLTVN+ AL+INF++A +G PLT VQ
Sbjct: 3 DVIIMDDNFTTIVNVARWGRAIYINIQKFVQFQLTVNIVALIINFVSACITGSAPLTAVQ 182
Query: 785 LLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVF 844
LLWVNLIMDTLGALALATE P LM + P+GRT ITK MWRN+ Q+LYQ+ VL V
Sbjct: 183 LLWVNLIMDTLGALALATEPPNDGLMLRPPVGRTTNFITKPMWRNIFGQSLYQLIVLAVL 362
Query: 845 QFYGKSIFNVSKE----VKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIV 900
F GK + ++ V NTLIFN+FV CQVFNE NSR +EK+N+F+G+ ++ +F ++
Sbjct: 363 TFDGKRLLRINGPDATIVLNTLIFNSFVFCQVFNEINSREIEKINIFKGMFESWIFFTVI 542
Query: 901 GITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
T+V QVL+VE L FA T L+W+ W + + I A S PI+ + K PV
Sbjct: 543 FSTVVFQVLIVEFLGTFASTVPLSWQFWVLSVVIGAFSMPISVILKCIPV 692
>BM085703 homologue to SP|Q9LIK7|ACA Potential calcium-transporting ATPase 13
plasma membrane-type (EC 3.6.3.8) (Ca(2+)-ATPase
isoform, partial (12%)
Length = 411
Score = 245 bits (625), Expect = 8e-65
Identities = 123/136 (90%), Positives = 128/136 (93%)
Frame = +3
Query: 686 KGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRC 745
KGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNF SV TVLRWGRC
Sbjct: 3 KGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFASVVTVLRWGRC 182
Query: 746 VYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERP 805
VYNNIQKFIQFQLTVNVAAL INF+AAVS+G VPLT VQLLWVNLIMDTLGALALATE+P
Sbjct: 183 VYNNIQKFIQFQLTVNVAALAINFVAAVSAGKVPLTAVQLLWVNLIMDTLGALALATEKP 362
Query: 806 TKELMQKKPIGRTEPL 821
T ELM K P+GRT+PL
Sbjct: 363 TMELMHKPPVGRTKPL 410
>TC231696 weakly similar to UP|ACAA_ARATH (Q9SZR1) Potential
calcium-transporting ATPase 10, plasma membrane-type
(Ca(2+)-ATPase isoform 10) , partial (15%)
Length = 824
Score = 176 bits (446), Expect = 4e-44
Identities = 83/153 (54%), Positives = 113/153 (73%), Gaps = 7/153 (4%)
Frame = +1
Query: 806 TKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKE-------V 858
T LM + P G+ EPL++ IMWRNLL QA+YQ++VLL+ F G S+ + E V
Sbjct: 1 TDSLMDQSPKGQREPLVSNIMWRNLLIQAMYQVSVLLILNFRGVSLLALRDEPNRPAIKV 180
Query: 859 KNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFA 918
KN+LIFN FVLCQVFNEFN+R +K N+F+G+ +N+LF+GIVGIT+VLQ++++E L KF
Sbjct: 181 KNSLIFNAFVLCQVFNEFNARKPDKFNIFKGVTRNYLFMGIVGITVVLQIVIIEYLGKFT 360
Query: 919 DTERLNWEQWGICIGIAAVSWPIAWLTKLTPVP 951
T +LNW+QW I + IA +SWP+A + KL PVP
Sbjct: 361 KTAKLNWKQWLISVIIAFISWPLAVVGKLIPVP 459
>TC205559 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(14%)
Length = 855
Score = 171 bits (434), Expect = 1e-42
Identities = 79/144 (54%), Positives = 107/144 (73%), Gaps = 7/144 (4%)
Frame = +3
Query: 816 GRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKE-------VKNTLIFNTFV 868
GR EPLIT IMWRNLL QA+YQ++VLLV F G SI +S + VKNTLIFN FV
Sbjct: 3 GRREPLITNIMWRNLLIQAMYQVSVLLVLNFRGISILGLSHDRKAHAIKVKNTLIFNAFV 182
Query: 869 LCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQW 928
LCQ+FNEFN+R ++ N+F+G+ +N+LF+GI+G+T+VLQ++++E L KF T RLNW+ W
Sbjct: 183 LCQIFNEFNARKPDEFNIFKGVTRNYLFMGIIGLTVVLQIVIIEFLGKFTSTVRLNWKHW 362
Query: 929 GICIGIAAVSWPIAWLTKLTPVPS 952
I + I + WP+A + KL PVP+
Sbjct: 363 LISVVIGLIGWPLAVIGKLIPVPT 434
>BI320368 similar to SP|Q9LU41|ACA Potential calcium-transporting ATPase 9
plasma membrane-type (EC 3.6.3.8) (Ca(2+)-ATPase isoform
9), partial (10%)
Length = 376
Score = 127 bits (318), Expect = 3e-29
Identities = 67/123 (54%), Positives = 86/123 (69%), Gaps = 1/123 (0%)
Frame = +3
Query: 378 CETMGSATVICTDKTGTLTLNQMRVTKFWLGLENV-VENFSNAMAPTVLELFHQGVGLNT 436
CETMGSAT IC+DKTGTLTLNQM V + ++G V + S+ + P L L ++G+ NT
Sbjct: 3 CETMGSATTICSDKTGTLTLNQMTVVEAYVGSTKVNPPDDSSKLHPKALSLINEGIAQNT 182
Query: 437 TGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGV 496
TG+V+ P E E+SGSPTEKA+L WAV LGM+ D ++ VLHV FNSEKKR GV
Sbjct: 183 TGNVFVPKDGGETEVSGSPTEKAIL*WAVK-LGMNFDVIRSNSTVLHVFPFNSEKKRGGV 359
Query: 497 AVR 499
A++
Sbjct: 360 ALK 368
>TC221308 similar to UP|Q9FVE7 (Q9FVE7) Plasma membrane Ca2+-ATPase, partial
(14%)
Length = 790
Score = 112 bits (281), Expect = 6e-25
Identities = 58/140 (41%), Positives = 86/140 (61%), Gaps = 4/140 (2%)
Frame = +2
Query: 817 RTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQV 872
R E I+ +MWRN+ ++ Q GK F++ S + NTLIFN+FV CQV
Sbjct: 2 RKENFISNVMWRNIXXXSIXXXVXXWFLQTRGKVXFHLDGPDSDLILNTLIFNSFVFCQV 181
Query: 873 FNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICI 932
FNE +SR ME++NVF+GILKN++F+ ++ T+V Q+++VE L FA+T L+ +QW +
Sbjct: 182 FNEISSRDMERVNVFQGILKNYVFVAVLTCTVVFQIIIVEFLGTFANTSPLSLKQWFGSV 361
Query: 933 GIAAVSWPIAWLTKLTPVPS 952
+ PIA K+ PV S
Sbjct: 362 LFGVLGMPIAAALKMIPVGS 421
>TC232411 similar to UP|Q8W0V0 (Q8W0V0) Type IIB calcium ATPase, partial
(19%)
Length = 802
Score = 110 bits (274), Expect = 4e-24
Identities = 57/144 (39%), Positives = 85/144 (58%), Gaps = 4/144 (2%)
Frame = +2
Query: 811 QKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVS----KEVKNTLIFNT 866
+K P + L + L Q++YQ+ +L + F GK + +S ++ NTLIFN+
Sbjct: 137 EKTPSCKGSKLYHQTYVEEYLFQSIYQLIILGILNFDGKRLLGLSGSDATKILNTLIFNS 316
Query: 867 FVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWE 926
FV CQVFNE NSR ++K+N+F G+ + +F+ I+ T QV++VE L FA T LNW+
Sbjct: 317 FVFCQVFNEINSRDIDKINIFRGMFDSWIFMAIIFATAAFQVVIVEFLGTFASTVPLNWQ 496
Query: 927 QWGICIGIAAVSWPIAWLTKLTPV 950
W + + I A S PIA + K PV
Sbjct: 497 FWLLSVVIGAFSMPIAAILKCIPV 568
Score = 74.3 bits (181), Expect = 2e-13
Identities = 36/56 (64%), Positives = 42/56 (74%)
Frame = +3
Query: 779 PLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQA 834
PLT VQLLWVNLIMDTLGALALATE P L+++ P+ R ITK MWRN+ +A
Sbjct: 42 PLTAVQLLWVNLIMDTLGALALATEPPNDGLLKRPPVARGANFITKPMWRNIFFKA 209
>BQ628440 similar to GP|16508164|gb type IIB calcium ATPase {Medicago
truncatula}, partial (5%)
Length = 446
Score = 108 bits (270), Expect = 1e-23
Identities = 52/67 (77%), Positives = 57/67 (84%), Gaps = 1/67 (1%)
Frame = +3
Query: 898 GIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFT 957
G GIT+VLQVLMVELLRKFADTERL WEQWGICI IAAVSWPIAW+TKL PV + FF+
Sbjct: 27 GTRGITLVLQVLMVELLRKFADTERLTWEQWGICIVIAAVSWPIAWITKLVPVSDRTFFS 206
Query: 958 -NAKWVK 963
+ KWVK
Sbjct: 207 HHVKWVK 227
>TC217853 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase,
partial (10%)
Length = 806
Score = 103 bits (256), Expect = 5e-22
Identities = 49/93 (52%), Positives = 64/93 (68%)
Frame = +2
Query: 858 VKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKF 917
V NTLIFNTFV CQVFNE NSR MEK+NVF+GIL N++F+G++ T+ Q+++VE L F
Sbjct: 71 VLNTLIFNTFVFCQVFNEINSREMEKINVFKGILDNYVFVGVISATVFFQIIIVEYLGTF 250
Query: 918 ADTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
A+T L QW C+ + + PIA K PV
Sbjct: 251 ANTTPLTLAQWFFCLLVGFLGMPIAARLKKIPV 349
>AW310835 similar to GP|16508162|g type IIB calcium ATPase {Medicago
truncatula}, partial (12%)
Length = 385
Score = 88.2 bits (217), Expect = 2e-17
Identities = 46/129 (35%), Positives = 78/129 (59%)
Frame = -1
Query: 29 LSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKI 88
++++V+ + Y + G VEG+ + L G+ DT R++++G N Y P K
Sbjct: 385 IASVVRGHDYNYYKKIGQVEGIIEKLSASADDGVGQVSIDT--RQDIYGVNRYTEKPSKS 212
Query: 89 FLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNF 148
FL FV EA +D T++IL+ CA +S+ G+ G +G Y+G I L++FLVV+V+A+S++
Sbjct: 211 FLMFVWEARHDLTLMILMVCAIVSIAIGLPTEGWPKGVYDGLGIILSIFLVVIVTAISDY 32
Query: 149 RQDRQFDKL 157
+Q + F L
Sbjct: 31 QQSKPFRDL 5
>TC205394 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase,
partial (12%)
Length = 782
Score = 88.2 bits (217), Expect = 2e-17
Identities = 56/158 (35%), Positives = 74/158 (46%), Gaps = 43/158 (27%)
Frame = +1
Query: 836 YQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQV------------------- 872
YQ V+ Q G SIF + S V NT IFN+FV CQV
Sbjct: 1 YQXMVIXFLQSRGXSIFLLEGPNSXXVXNTXIFNSFVXCQVTFLKS*QFF*EYLLLNS*C 180
Query: 873 --------------------FNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVE 912
FNE NSR MEK+NVF+GIL N++F+G++ T+ Q+++VE
Sbjct: 181 MYAICIEVSYKMWVDLVLQVFNEINSREMEKINVFKGILDNYVFVGVISATVFFQIIIVE 360
Query: 913 LLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
L FA+T L QW C+ + + PIA K PV
Sbjct: 361 YLGTFANTTPLTLSQWFFCLLVGFMGMPIAARLKKIPV 474
>TC230130 similar to UP|AHM7_ARATH (Q9SH30) Potential copper-transporting
ATPase 3 , partial (43%)
Length = 1409
Score = 85.1 bits (209), Expect = 1e-16
Identities = 53/180 (29%), Positives = 88/180 (48%)
Frame = +1
Query: 588 GIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEG 647
G++ + DP +P ++ + K + M+TGDN TA +IA E GI +
Sbjct: 655 GVLAVSDPLKPGAQEVISILKSMKIKSIMVTGDNFGTASSIAREVGIEN----------- 801
Query: 648 VEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEAD 707
V+A + P K V+ L+ G+ VA+ GDG ND+PAL AD
Sbjct: 802 -------------------VIAEAKPDQKAEKVKDLQASGYTVAMVGDGINDSPALVAAD 924
Query: 708 IGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVI 767
+G+++G GT++A E++DIV++ N V T + R ++ I+ + L N+ + I
Sbjct: 925 VGMAIG-AGTDIAIEAADIVLMKSNLEDVITAIDLSRKTFSRIRLNYFWALGYNLLGIPI 1101
Score = 31.6 bits (70), Expect = 1.8
Identities = 21/91 (23%), Positives = 41/91 (44%)
Frame = +1
Query: 339 VTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLN 398
++++V+A P L LA + + +++ A E+ I DKTGTLT+
Sbjct: 121 ISVMVIACPCALGLATPTAVMVGTGVGASQGVLIKGGQALESAHKVDCIVFDKTGTLTVG 300
Query: 399 QMRVTKFWLGLENVVENFSNAMAPTVLELFH 429
+ + + L + V++ F +A + H
Sbjct: 301 KPVIVRTELLTKMVLQEFYELVAAAEVNSEH 393
>AW164575 similar to GP|8843813|dbj Ca2+-transporting ATPase-like protein
{Arabidopsis thaliana}, partial (5%)
Length = 445
Score = 79.0 bits (193), Expect = 1e-14
Identities = 39/59 (66%), Positives = 46/59 (77%)
Frame = +3
Query: 650 FRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADI 708
FR ++ +R E D+I VM RSSP DKLL+VQ L++KGHVVAVTGDGTNDAPAL E I
Sbjct: 6 FRGLSDAQRDEIADRISVMGRSSPNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEVCI 182
>TC207402 similar to UP|Q941L1 (Q941L1) Copper-transporting P-type ATPase,
partial (19%)
Length = 721
Score = 78.2 bits (191), Expect = 2e-14
Identities = 50/164 (30%), Positives = 80/164 (48%)
Frame = +1
Query: 604 VETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVD 663
+E + GV M+TGDN TA+A+A E GI D
Sbjct: 13 IEGLQKMGVTPVMVTGDNWRTARAVAKEVGIQD--------------------------- 111
Query: 664 KIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKES 723
V A P K +V+ +K G + A+ GDG ND+PAL AD+G+++G GT++A E+
Sbjct: 112 ---VRAEVMPAGKADVVRSFQKDGSIAAMVGDGINDSPALAAADVGMAIG-AGTDIAIEA 279
Query: 724 SDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVI 767
++ V++ +N V T + R ++ I+ F + NV A+ +
Sbjct: 280 AEYVLMRNNLEDVITAIDLSRKTFSRIRLNYVFAMAYNVVAIPV 411
>TC230973 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPase, partial
(22%)
Length = 654
Score = 54.7 bits (130), Expect(2) = 4e-14
Identities = 28/67 (41%), Positives = 44/67 (64%)
Frame = +3
Query: 660 EKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEV 719
E ++K A P K +V+ L+++ H+ +TGDG NDAPALK+ADIG+++ T+
Sbjct: 456 ELIEKADGFAGVFPEHKYEIVKRLQERKHICGMTGDGVNDAPALKKADIGIAVA-DATDA 632
Query: 720 AKESSDI 726
A+ +SDI
Sbjct: 633 ARSASDI 653
Score = 42.4 bits (98), Expect(2) = 4e-14
Identities = 34/134 (25%), Positives = 56/134 (41%)
Frame = +2
Query: 501 ETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAY 560
+++ H KGA E +L +C+ ++ R ++ I A LR + A
Sbjct: 35 DSDGNWHRSSKGAPEQILNLCN----------CKEDVRKRVHGTIDKFAERGLRSLGVAR 184
Query: 561 MEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGD 620
E+ E G P Q +G++ L DP R + + + GV++KMITGD
Sbjct: 185 QEVPEKNKD-SPGAPWQ--------FVGLLPLFDPPRHDSAETITRALNLGVNVKMITGD 337
Query: 621 NIFTAKAIATECGI 634
+ AK G+
Sbjct: 338 QLAIAKETGRRLGM 379
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,010,221
Number of Sequences: 63676
Number of extensions: 453738
Number of successful extensions: 2276
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 2241
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2259
length of query: 967
length of database: 12,639,632
effective HSP length: 106
effective length of query: 861
effective length of database: 5,889,976
effective search space: 5071269336
effective search space used: 5071269336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0322.1


