

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0321.6
(497 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
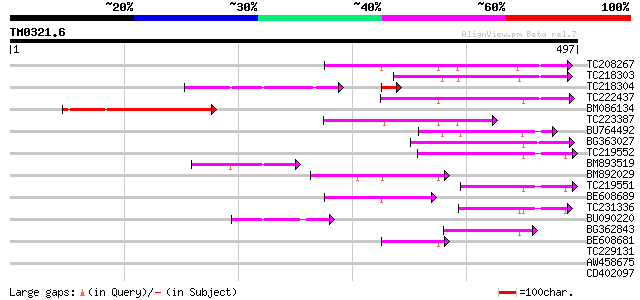
Score E
Sequences producing significant alignments: (bits) Value
TC208267 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, p... 116 2e-26
TC218303 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, p... 107 2e-23
TC218304 similar to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, par... 99 2e-23
TC222437 weakly similar to UP|O23045 (O23045) YUP8H12.16 protein... 106 2e-23
BM086134 99 3e-21
TC223387 89 6e-18
BU764492 78 1e-14
BG363027 71 1e-12
TC219552 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX... 70 2e-12
BM893519 similar to GP|22475197|gb homeodomain protein GhHOX2 {G... 68 1e-11
BM892029 68 1e-11
TC219551 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX... 61 1e-09
BE608689 similar to GP|3925363|gb| homeodomain protein {Malus x ... 59 7e-09
TC231336 similar to UP|Q9ZTA8 (Q9ZTA8) Homeodomain protein (Frag... 57 1e-08
BU090220 similar to GP|22475197|gb| homeodomain protein GhHOX2 {... 55 6e-08
BG362843 similar to GP|8885530|dbj| homeodomain transcription fa... 53 3e-07
BE608681 weakly similar to GP|3925363|gb| homeodomain protein {M... 49 4e-06
TC229131 similar to UP|Q9ZU11 (Q9ZU11) F5F19.21 protein (At1g521... 39 0.004
AW458675 similar to GP|1881536|gb|A A20 {Arabidopsis thaliana}, ... 39 0.007
CD402097 similar to GP|18481701|gb| OCL5 protein {Sorghum bicolo... 36 0.045
>TC208267 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (31%)
Length = 1049
Score = 116 bits (291), Expect = 2e-26
Identities = 77/236 (32%), Positives = 128/236 (53%), Gaps = 19/236 (8%)
Frame = +2
Query: 277 GRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNTI----LDGIVVIA 332
GRKS++ L+ RMV SF ++ S ++++ ++ GIV+ A
Sbjct: 47 GRKSMMKLAERMVISFCAGVSASTAHTWTTLSGTGADDVRVMTRKSADDPGRPPGIVLSA 226
Query: 333 VSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--INESNHVSILQTSSII 390
+S WLPV PK +F FL+DE + EWD L+ G +++ AHI+ + N VS+L+ +S
Sbjct: 227 ATSFWLPVPPKRVFDFLRDENSRNEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSAN 406
Query: 391 G--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRD----- 443
+M+I QES + G +V+YAP+D + +++NG D +LPSGF I D
Sbjct: 407 SSQSNMLILQESCTNSTGSFVIYAPVDIVAMNVVLNGGDPDYVALLPSGFAILPDGTTSH 586
Query: 444 ------GREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDAL 493
G G GSLLT+ FQ+LVD ++++ ++ ++++ ++RIK +L
Sbjct: 587 NGSGGIGETGPSGSLLTVAFQILVDSV-PTAKLSLGSVATVNNLIACTVERIKASL 751
>TC218303 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (32%)
Length = 1230
Score = 107 bits (266), Expect = 2e-23
Identities = 63/170 (37%), Positives = 101/170 (59%), Gaps = 13/170 (7%)
Frame = +2
Query: 337 WLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISINES--NHVSILQTSSIIGE-- 392
WLPV+PK +F FL+DE ++EWD L+ G +++ AHI+ N VS+L+ +S
Sbjct: 206 WLPVSPKRVFEFLRDENSRSEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSANSSQS 385
Query: 393 HMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDGR------- 445
+M+I QES D G +V+YAP+D + +++NG D +LPSGF I DG
Sbjct: 386 NMLILQESCADSTGSFVIYAPVDIVAMNVVLNGGDPDYVALLPSGFAILPDGTTAHGGGI 565
Query: 446 --EGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDAL 493
G+ GSLLT+ FQ+LVD ++++ ++ ++++ ++RIK AL
Sbjct: 566 GDTGHGGSLLTVAFQILVDSV-PTAKLSLGSVATVNNLIACTVERIKAAL 712
>TC218304 similar to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (26%)
Length = 575
Score = 99.4 bits (246), Expect(2) = 2e-23
Identities = 57/139 (41%), Positives = 82/139 (58%)
Frame = +1
Query: 154 PSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIRELPNGCCMVTWV 213
P+RE +F+RYC+Q G W +VDVSLD+ S ++ PSGC I+E+PNG V WV
Sbjct: 1 PTREIYFVRYCKQHGDGTWAVVDVSLDNLRPSPSARCRRR-PSGCLIQEMPNGYSKVIWV 177
Query: 214 EQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVK 273
E VEV + L + +V + A+GAKRW+ L R E+ A + + + + G +
Sbjct: 178 EHVEVDDR-GVHNLYKQLVSSGHAFGAKRWIANLDRQCERLASAMATNIPTVD--VGVIT 348
Query: 274 LLDGRKSVINLSNRMVGSF 292
DGRKS++ L+ RMV SF
Sbjct: 349 NPDGRKSMLKLAERMVISF 405
Score = 28.1 bits (61), Expect(2) = 2e-23
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +2
Query: 327 GIVVIAVSSIWLPVTPK 343
GIV+ A +S WLPV+PK
Sbjct: 521 GIVLSAATSFWLPVSPK 571
>TC222437 weakly similar to UP|O23045 (O23045) YUP8H12.16 protein, partial
(16%)
Length = 719
Score = 106 bits (265), Expect = 2e-23
Identities = 61/192 (31%), Positives = 109/192 (56%), Gaps = 22/192 (11%)
Frame = +1
Query: 326 DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--INESNHVSI 383
+G+V+ A ++IWLP P +F F KDE ++ +WD L+ G +++ A+I+ ++ N +S+
Sbjct: 43 NGVVLSAATTIWLPTPPHAVFNFFKDENKRPQWDVLSNGNAVQEVANIANGLHPGNSISV 222
Query: 384 LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRD 443
L+ + ++M+I QES ID G +VVY P+D + ++G D S +LP+GFTI D
Sbjct: 223 LRAFNNSTQNMLILQESCIDSYGSFVVYCPVDLPSINLAMSGEDPSYIPLLPNGFTILPD 402
Query: 444 GREGNE--------------------GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCM 483
G+ E GSL+T+ FQ+LV S+++ +++ ++++
Sbjct: 403 GQPDQEGDDGASTSSNNANRNIVRSGGSLVTIAFQILVSSL-PSAKLNMESVTTVNNLIG 579
Query: 484 LMLQRIKDALHC 495
+Q+IK +L C
Sbjct: 580 STVQQIKSSLSC 615
>BM086134
Length = 428
Score = 99.4 bits (246), Expect = 3e-21
Identities = 51/136 (37%), Positives = 90/136 (65%), Gaps = 1/136 (0%)
Frame = +1
Query: 47 EPLWVKSSNDVVGFVLVREIYAKKFPR-ITNSENLYVCEESSKDSLLVMASGMELVKVFL 105
EPLW+ ++ D +L + Y + FPR I + + CE +S+++ +V+ + + LV++ +
Sbjct: 22 EPLWL-TTLDGTSTMLNEDEYIRSFPRGIGPKPSGFKCE-ASRETAVVIMNHVNLVEILM 195
Query: 106 DSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQ 165
D ++W+ + IVS+A T++V G N++GAL++M E+ + +PLVP+RE +F+RYC+
Sbjct: 196 DVNQWSTVFSGIVSRAMTLEVLSTGVAGNYNGALQVMTAELQLPTPLVPTRESYFVRYCK 375
Query: 166 QVEAGEWLIVDVSLDS 181
Q G W +VDVSLD+
Sbjct: 376 QHADGTWAVVDVSLDN 423
>TC223387
Length = 569
Score = 88.6 bits (218), Expect = 6e-18
Identities = 55/160 (34%), Positives = 90/160 (55%), Gaps = 8/160 (5%)
Frame = +2
Query: 276 DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNTILDG----IVVI 331
+GRKS++ L+ RMV S+ + S ++++ ++T G IV+
Sbjct: 77 EGRKSMMKLAERMVMSYCTGVGASTAHAWTTLSATGCDDVRVMTRKSTDEPGRPPGIVLS 256
Query: 332 AVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--INESNHVSILQTSSI 389
A +S WLPV PK +F FL+D+ + EWD L+ G +++ AHI+ + N VS+L+ +S
Sbjct: 257 AATSFWLPVPPKRVFHFLRDQNSRNEWDILSNGGLVQELAHIANGRDPGNCVSLLRVNSA 436
Query: 390 IG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCD 427
+M+I QES D G YVVYAP+D + ++++G D
Sbjct: 437 NSSQSNMLILQESCTDSTGSYVVYAPVDIVAMNVVLSGGD 556
>BU764492
Length = 428
Score = 77.8 bits (190), Expect = 1e-14
Identities = 54/146 (36%), Positives = 77/146 (51%), Gaps = 24/146 (16%)
Frame = +3
Query: 359 DFLAGGRPMRQKAHISINES--NHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPI 414
D L+ G PM++ AHI+ + N VS+L+ S+I M+I QE+ ID G VVYAP+
Sbjct: 3 DILSNGGPMQEMAHIAKGQDHGNAVSLLRASAINSNQSSMLILQETCIDAAGSLVVYAPV 182
Query: 415 DKLDAYMIVNGCDSSKKLILPSGFTISRDGREGN--------------------EGSLLT 454
D Y+++NG DS+ +LPSGF I DG GSLLT
Sbjct: 183 DIPVLYVVMNGGDSAYVALLPSGFAIVPDGPGSRGPPNGPTSTTNGGDNGVTRVSGSLLT 362
Query: 455 MCFQVLVDEEGSSSQVRKKTLESIDS 480
+ FQ+LV +S K T+ES+++
Sbjct: 363 VAFQILV----NSLPTAKLTVESVET 428
>BG363027
Length = 492
Score = 71.2 bits (173), Expect = 1e-12
Identities = 47/159 (29%), Positives = 79/159 (49%), Gaps = 15/159 (9%)
Frame = +2
Query: 352 EFRKAEWDFLAGGRPMRQKAHISINESN-HVSILQTSSIIGEHMVIFQESSIDDLGGYVV 410
+F +WD ++ G ++ A+++ + +V +Q + I Q+S VV
Sbjct: 17 KFSLPQWDIMSSGGSVQSVANLAKGKDRGNVVNIQKIQSKDNSVWILQDSCTSAYESMVV 196
Query: 411 YAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDGREGNE--------------GSLLTMC 456
YAP++ ++ GCDSS ILPSGF+I DG EG GSL TM
Sbjct: 197 YAPVEFAGIQSVLTGCDSSNLAILPSGFSILPDGIEGRPLVISSRQEEKYTEGGSLFTMA 376
Query: 457 FQVLVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
FQ+LV+ + ++ +++ES+++ L+ IK +L C
Sbjct: 377 FQILVN-PSPTVKLTTESVESVNNLVSCTLRNIKTSLQC 490
>TC219552 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX1, partial
(20%)
Length = 804
Score = 70.1 bits (170), Expect = 2e-12
Identities = 50/158 (31%), Positives = 76/158 (47%), Gaps = 18/158 (11%)
Frame = +1
Query: 358 WDFLAGGRPMRQKAHISINESNHVSI-LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDK 416
WD ++ G ++ A+++ + ++ +QT + I Q+S + V YA +D
Sbjct: 40 WDIMSSGGTVQSIANLAKGQDRGNAVAIQTIKSKENSVWILQDSYTNPYESMVAYASVDI 219
Query: 417 LDAYMIVNGCDSSKKLILPSGFTISRDGREGNE--------------GSLLTMCFQVLVD 462
++ GCDSS ILPSGF+I DG E GSL TM FQ+L
Sbjct: 220 TGIQSVMTGCDSSNLAILPSGFSIIPDGLESRPLVISSRQEEKNTEGGSLFTMAFQILT- 396
Query: 463 EEGSSSQVRKKTLESIDSSCMLM---LQRIKDALHCSD 497
++S K TLES+DS L+ L+ I+ +L C D
Sbjct: 397 ---NASPTAKLTLESVDSVNTLVSCTLRNIRTSLQCED 501
>BM893519 similar to GP|22475197|gb homeodomain protein GhHOX2 {Gossypium
hirsutum}, partial (13%)
Length = 421
Score = 67.8 bits (164), Expect = 1e-11
Identities = 41/99 (41%), Positives = 57/99 (57%), Gaps = 3/99 (3%)
Frame = +3
Query: 160 FLRYCQQ-VEAGEWLIVDVSLDSFGNCASRSVAK--KFPSGCRIRELPNGCCMVTWVEQV 216
FLRYCQQ G W IVD +DSF S + + SGC I+++PNG VTWVE
Sbjct: 3 FLRYCQQNAMEGTWAIVDFPVDSFHQNFHPSYPRYCRRSSGCVIQDMPNGYSRVTWVEHA 182
Query: 217 EVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKA 255
+V + + + + V + +A+GA+RWL LQR E+ A
Sbjct: 183 KVEEK-PVHQIFCNYVYSGMAFGAQRWLGVLQRQCERVA 296
>BM892029
Length = 421
Score = 67.8 bits (164), Expect = 1e-11
Identities = 37/129 (28%), Positives = 74/129 (56%), Gaps = 7/129 (5%)
Frame = +2
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQN--LLDPSNQLTTSIKLIAHR 321
ST L G + +G++S++ L+ RMV +F +++ L S +++ H+
Sbjct: 32 STRDLGGVIPSPEGKRSMMKLAQRMVTNFCASISSSAGHRWTTLSGSGMNEVGVRVTVHK 211
Query: 322 NTIL---DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--IN 376
++ +G+V+ A ++IWLP+ P+ +F F KDE ++ +WD L+ G +++ AHI+ +
Sbjct: 212 SSDPGQPNGVVLSAATTIWLPIPPQTVFNFFKDEKKRPQWDVLSNGNAVQEVAHIANGSH 391
Query: 377 ESNHVSILQ 385
N +S+L+
Sbjct: 392 PGNCISVLR 418
>TC219551 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX1, partial
(15%)
Length = 671
Score = 61.2 bits (147), Expect = 1e-09
Identities = 42/119 (35%), Positives = 56/119 (46%), Gaps = 17/119 (14%)
Frame = +1
Query: 396 IFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDGREGNE------ 449
I +S VVYA +D ++ GCDSS ILP GF+I DG E
Sbjct: 13 ILXDSXXXXXESMVVYASVDITXTQSVMTGCDSSNLAILPXGFSIIPDGLESRPLVISSR 192
Query: 450 --------GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLM---LQRIKDALHCSD 497
GSL TM FQ+L ++S K T+ES+DS L+ L+ I+ +L C D
Sbjct: 193 QEEKNTEGGSLFTMAFQILT----NASPAAKLTMESVDSVNTLVSCTLRNIRTSLQCED 357
>BE608689 similar to GP|3925363|gb| homeodomain protein {Malus x domestica},
partial (18%)
Length = 413
Score = 58.5 bits (140), Expect = 7e-09
Identities = 33/103 (32%), Positives = 57/103 (55%), Gaps = 5/103 (4%)
Frame = +2
Query: 277 GRKSVINLSNRMVGSFFKVLNTQGIQNLLDPS-NQLTTSIKLIAHRNTI----LDGIVVI 331
GR+S++ L+ RM +F + + L +K++ +N GIV+
Sbjct: 92 GRRSMLKLAQRMTSNFCSGVCASSARKWDSLHIGTLGDDMKVMTRKNVDDPGEPPGIVLS 271
Query: 332 AVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
A +S+W+PV+ + LF FL+DE ++EWD L+ G PM++ HI+
Sbjct: 272 AATSVWVPVSRQRLFDFLRDERLRSEWDILSNGGPMQEMVHIA 400
>TC231336 similar to UP|Q9ZTA8 (Q9ZTA8) Homeodomain protein (Fragment),
partial (19%)
Length = 869
Score = 57.4 bits (137), Expect = 1e-08
Identities = 44/119 (36%), Positives = 65/119 (53%), Gaps = 19/119 (15%)
Frame = +3
Query: 394 MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDGR-------- 445
M+I QE+ +D VVYAP+D ++++G DS+ +LPSGF I DG
Sbjct: 93 MLILQETWMDASCSVVVYAPVDVQSLNVVMSGGDSAYVALLPSGFAILPDGHCNDNGCNG 272
Query: 446 ------EGNE--GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLM---LQRIKDAL 493
GN+ GSLLT+ FQ+LV +S K T+ES+D+ L+ +Q+IK +L
Sbjct: 273 TLQKGGGGNDGGGSLLTVGFQILV----NSLPTAKLTVESVDTVNNLISCTIQKIKASL 437
>BU090220 similar to GP|22475197|gb| homeodomain protein GhHOX2 {Gossypium
hirsutum}, partial (11%)
Length = 267
Score = 55.5 bits (132), Expect = 6e-08
Identities = 33/90 (36%), Positives = 51/90 (56%)
Frame = +2
Query: 195 PSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKK 254
PSGC I+++PNG VTWVE +V + + + + V + +A+GA+RWL LQR E+
Sbjct: 11 PSGCVIQDMPNGYSRVTWVEHAKVEEK-PVHQIFCNYVYSGMAFGAQRWLGVLQRQCERV 187
Query: 255 ACSEFSYTASTEGLEGEVKLLDGRKSVINL 284
A S A G + + D RK+++ L
Sbjct: 188 A----SLMARNISDLGVIPIADARKNLMKL 265
>BG362843 similar to GP|8885530|dbj| homeodomain transcription factor-like
{Arabidopsis thaliana}, partial (10%)
Length = 466
Score = 53.1 bits (126), Expect = 3e-07
Identities = 35/99 (35%), Positives = 51/99 (51%), Gaps = 17/99 (17%)
Frame = +1
Query: 381 VSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTI 440
V+ LQ + M+I QE+ +D VVYAP+D ++++G DS+ +LPSGF I
Sbjct: 154 VTYLQAVNANDSSMLILQETWMDASCSVVVYAPVDVQSLNVVMSGGDSAYVALLPSGFAI 333
Query: 441 SRDGR-----------------EGNEGSLLTMCFQVLVD 462
DG G+ GSLLT+ FQ+LV+
Sbjct: 334 LPDGHCNDNGCNGSLQKGRGSDYGSGGSLLTVGFQILVN 450
>BE608681 weakly similar to GP|3925363|gb| homeodomain protein {Malus x
domestica}, partial (19%)
Length = 475
Score = 49.3 bits (116), Expect = 4e-06
Identities = 27/61 (44%), Positives = 34/61 (55%), Gaps = 2/61 (3%)
Frame = +2
Query: 327 GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--INESNHVSIL 384
GIV+ A +S+W+PVT L L DE ++EWD L G PM HI N VSIL
Sbjct: 245 GIVLTAATSVWVPVTRMRLLNSLLDEHLRSEWDILFNGGPMHDMVHIVR*QGHGNCVSIL 424
Query: 385 Q 385
+
Sbjct: 425 R 427
>TC229131 similar to UP|Q9ZU11 (Q9ZU11) F5F19.21 protein (At1g52150/F5F19_21)
(Homeodomain-leucine zipper protein), partial (49%)
Length = 1253
Score = 39.3 bits (90), Expect = 0.004
Identities = 26/120 (21%), Positives = 59/120 (48%), Gaps = 6/120 (5%)
Frame = +1
Query: 135 HSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKF 194
+ G + L++ +++ + L P+R+++ LRY +E G +I + SL + N S + F
Sbjct: 85 NGGTIELLYMQLYAPTTLAPARDFWLLRYTSVLEDGSLVICERSLKNTQNGPSMPPVQHF 264
Query: 195 ------PSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQ 248
PSG IR G ++ V+ +++ + + + ++R + ++ K + L+
Sbjct: 265 VRAEMLPSGYLIRPCEGGGSIIHIVDHMDL-EPWSVPEVLRPLYESSTVLAQKTTMAALR 441
>AW458675 similar to GP|1881536|gb|A A20 {Arabidopsis thaliana}, partial
(11%)
Length = 411
Score = 38.5 bits (88), Expect = 0.007
Identities = 27/82 (32%), Positives = 45/82 (53%), Gaps = 5/82 (6%)
Frame = +2
Query: 276 DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNTILD-----GIVV 330
+GRKS++ L+ RMV SF + N P +++++ R ++ D GIV+
Sbjct: 11 EGRKSMMKLAERMVLSFSTGVGAS-TANAWTPLPLDLENVRVMT-RKSVDDPGRPSGIVL 184
Query: 331 IAVSSIWLPVTPKNLFLFLKDE 352
A +S+WLPV + +F FL+ E
Sbjct: 185 SAATSLWLPVPARRVFDFLRSE 250
>CD402097 similar to GP|18481701|gb| OCL5 protein {Sorghum bicolor}, partial
(9%)
Length = 665
Score = 35.8 bits (81), Expect = 0.045
Identities = 24/71 (33%), Positives = 40/71 (55%), Gaps = 10/71 (14%)
Frame = -3
Query: 433 ILPSGFTISRDG----------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSC 482
+LPSGF I DG G+ GSLLT+ FQ+LVD ++++ ++ +++S
Sbjct: 642 LLPSGFAILPDGPPALNGGPKHEVGSGGSLLTVGFQILVD-SAPTAKLSLGSVATVNSLI 466
Query: 483 MLMLQRIKDAL 493
++RIK A+
Sbjct: 465 KCTVERIKVAV 433
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,191,610
Number of Sequences: 63676
Number of extensions: 286705
Number of successful extensions: 1231
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1199
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1205
length of query: 497
length of database: 12,639,632
effective HSP length: 101
effective length of query: 396
effective length of database: 6,208,356
effective search space: 2458508976
effective search space used: 2458508976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0321.6


